Please be patient as the page loads
|
NUPL2_MOUSE
|
||||||
| SwissProt Accessions | Q8CIC2, Q8BJX3, Q8BVP7, Q924S0 | Gene names | Nupl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoporin-like 2 (NLP-1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
NUPL2_MOUSE
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q8CIC2, Q8BJX3, Q8BVP7, Q924S0 | Gene names | Nupl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoporin-like 2 (NLP-1). | |||||
|
NUPL2_HUMAN
|
||||||
| θ value | 1.42253e-146 (rank : 2) | NC score | 0.976554 (rank : 2) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O15504, Q49AE7, Q9BS49 | Gene names | NUPL2, CG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoporin-like 2 (NLP-1) (hCG1) (NUP42 homolog). | |||||
|
MUC1_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 3) | NC score | 0.043025 (rank : 20) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
SC24C_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 4) | NC score | 0.071843 (rank : 6) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 662 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P53992, Q8WV25 | Gene names | SEC24C, KIAA0079 | |||
|
Domain Architecture |

|
|||||
| Description | Protein transport protein Sec24C (SEC24-related protein C). | |||||
|
NU153_HUMAN
|
||||||
| θ value | 0.279714 (rank : 5) | NC score | 0.078564 (rank : 5) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P49790 | Gene names | NUP153 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup153 (Nucleoporin Nup153) (153 kDa nucleoporin). | |||||
|
PAR12_HUMAN
|
||||||
| θ value | 0.365318 (rank : 6) | NC score | 0.093345 (rank : 4) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9H0J9, Q9H610, Q9NP36, Q9NTI3 | Gene names | PARP12, ZC3HDC1 | |||
|
Domain Architecture |
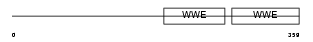
|
|||||
| Description | Poly [ADP-ribose] polymerase 12 (EC 2.4.2.30) (PARP-12) (Zinc finger CCCH domain-containing protein 1). | |||||
|
PAR12_MOUSE
|
||||||
| θ value | 0.365318 (rank : 7) | NC score | 0.094507 (rank : 3) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BZ20, Q80VL6, Q8K333, Q8R3U2 | Gene names | Parp12, Zc3hdc1 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 12 (EC 2.4.2.30) (PARP-12) (Zinc finger CCCH domain-containing protein 1). | |||||
|
TROP_HUMAN
|
||||||
| θ value | 0.47712 (rank : 8) | NC score | 0.031953 (rank : 34) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q12816, Q9NU89, Q9UPN8 | Gene names | TRO, KIAA1114, MAGED3 | |||
|
Domain Architecture |

|
|||||
| Description | Trophinin (MAGE-D3 antigen). | |||||
|
HRBL_MOUSE
|
||||||
| θ value | 1.06291 (rank : 9) | NC score | 0.040429 (rank : 23) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q80WC7, Q8BKS5, Q99J67 | Gene names | Hrbl, Rabr | |||
|
Domain Architecture |
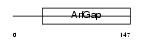
|
|||||
| Description | HIV-1 Rev-binding protein-like protein (Rev/Rex activation domain- binding protein related) (RAB-R). | |||||
|
DHX57_HUMAN
|
||||||
| θ value | 1.38821 (rank : 10) | NC score | 0.050164 (rank : 18) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6P158, Q53SI4, Q6P9G1, Q7Z6H3, Q8NG17, Q96M33 | Gene names | DHX57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative ATP-dependent RNA helicase DHX57 (EC 3.6.1.-) (DEAH box protein 57). | |||||
|
FBX42_HUMAN
|
||||||
| θ value | 1.81305 (rank : 11) | NC score | 0.031211 (rank : 35) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6P3S6, Q5TEU8, Q86XI0, Q8N3N4, Q8N5F8, Q9BRM0, Q9P2L4 | Gene names | FBXO42, FBX42, KIAA1332 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 42. | |||||
|
NDF4_MOUSE
|
||||||
| θ value | 1.81305 (rank : 12) | NC score | 0.021380 (rank : 42) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O09105 | Gene names | Neurod4, Ath3, Atoh3 | |||
|
Domain Architecture |
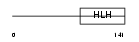
|
|||||
| Description | Neurogenic differentiation factor 4 (NeuroD4) (Atonal protein homolog 3) (Helix-loop-helix protein mATH-3) (MATH3). | |||||
|
TNR6A_HUMAN
|
||||||
| θ value | 1.81305 (rank : 13) | NC score | 0.040014 (rank : 25) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8NDV7, O15408, Q658L5, Q6NVB5, Q8NEZ0, Q8TBT8, Q8TCR0, Q9NV59, Q9P268 | Gene names | TNRC6A, CAGH26, KIAA1460, TNRC6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trinucleotide repeat-containing gene 6A protein (CAG repeat protein 26) (Glycine-tryptophan protein of 182 kDa) (GW182 autoantigen) (Protein GW1) (EMSY interactor protein). | |||||
|
SET1A_HUMAN
|
||||||
| θ value | 2.36792 (rank : 14) | NC score | 0.026257 (rank : 37) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O15047, Q6PIF3, Q8TAJ6 | Gene names | SETD1A, KIAA0339, SET1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET1 (EC 2.1.1.43) (Set1/Ash2 histone methyltransferase complex subunit SET1) (SET-domain-containing protein 1A). | |||||
|
TPO_MOUSE
|
||||||
| θ value | 2.36792 (rank : 15) | NC score | 0.034929 (rank : 29) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P40226 | Gene names | Thpo | |||
|
Domain Architecture |

|
|||||
| Description | Thrombopoietin precursor (Megakaryocyte colony-stimulating factor) (Myeloproliferative leukemia virus oncogene ligand) (C-mpl ligand) (ML) (Megakaryocyte growth and development factor) (MGDF). | |||||
|
CXXC6_HUMAN
|
||||||
| θ value | 3.0926 (rank : 16) | NC score | 0.036511 (rank : 28) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NFU7, Q5VUP7, Q7Z6B6, Q8TCR1, Q9C0I7 | Gene names | CXXC6, KIAA1676, LCX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CXXC-type zinc finger protein 6 (Leukemia-associated protein with a CXXC domain). | |||||
|
MKRN2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 17) | NC score | 0.068850 (rank : 11) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9H000, Q8N391, Q96BD4, Q9BUY2, Q9NRY1 | Gene names | MKRN2, RNF62 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-2 (RING finger protein 62). | |||||
|
MKRN3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 18) | NC score | 0.069467 (rank : 10) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q60764, Q7TQE4 | Gene names | Mkrn3, Zfp127, Znf127 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-3 (Zinc finger protein 127). | |||||
|
NAB2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 19) | NC score | 0.042012 (rank : 22) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15742, O76006, Q14797 | Gene names | NAB2, MADER | |||
|
Domain Architecture |

|
|||||
| Description | NGFI-A-binding protein 2 (EGR-1-binding protein 2) (Melanoma- associated delayed early response protein) (Protein MADER). | |||||
|
NAB2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 20) | NC score | 0.042299 (rank : 21) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q61127 | Gene names | Nab2 | |||
|
Domain Architecture |

|
|||||
| Description | NGFI-A-binding protein 2 (EGR-1-binding protein 2). | |||||
|
ZC3H6_HUMAN
|
||||||
| θ value | 3.0926 (rank : 21) | NC score | 0.070410 (rank : 7) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P61129 | Gene names | ZC3H6, KIAA2035, ZC3HDC6 | |||
|
Domain Architecture |
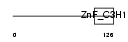
|
|||||
| Description | Zinc finger CCCH domain-containing protein 6. | |||||
|
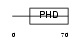
AF10_MOUSE
|
||||||
| θ value | 4.03905 (rank : 22) | NC score | 0.017561 (rank : 43) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 182 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O54826 | Gene names | Mllt10, Af10 | |||
|
Domain Architecture |
|
|||||
| Description | Protein AF-10. | |||||
|
CS007_HUMAN
|
||||||
| θ value | 4.03905 (rank : 23) | NC score | 0.069511 (rank : 9) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UPT8, Q9Y420 | Gene names | C19orf7, KIAA1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7. | |||||
|
CS007_MOUSE
|
||||||
| θ value | 4.03905 (rank : 24) | NC score | 0.070397 (rank : 8) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 464 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q6ZPZ3 | Gene names | Kiaa1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7 homolog. | |||||
|
MCM3A_MOUSE
|
||||||
| θ value | 4.03905 (rank : 25) | NC score | 0.029155 (rank : 36) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9WUU9 | Gene names | Mcm3ap, Ganp, Map80 | |||
|
Domain Architecture |

|
|||||
| Description | 80 kDa MCM3-associated protein (GANP protein). | |||||
|
MKRN1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 26) | NC score | 0.067310 (rank : 13) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9UHC7, Q6GSF1, Q9H0G0, Q9UEZ7 | Gene names | MKRN1, RNF61 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-1 (RING finger protein 61). | |||||
|
MKRN1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 27) | NC score | 0.067633 (rank : 12) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9QXP6 | Gene names | Mkrn1 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-1. | |||||
|
SCYL2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 28) | NC score | 0.013107 (rank : 48) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 603 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CFE4, Q3UT57, Q3UWU9, Q8K0M4 | Gene names | Scyl2, Cvak104, D10Ertd802e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SCY1-like protein 2 (Coated vesicle-associated kinase of 104 kDa). | |||||
|
SMG1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 29) | NC score | 0.022037 (rank : 40) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96Q15, O43305, Q13284, Q8NFX2, Q96QV0, Q96RW3 | Gene names | SMG1, ATX, KIAA0421, LIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SMG1 (EC 2.7.11.1) (SMG-1) (hSMG-1) (Lambda/iota protein kinase C-interacting protein) (Lambda-interacting protein) (61E3.4). | |||||
|
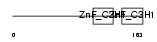
TTP_HUMAN
|
||||||
| θ value | 4.03905 (rank : 30) | NC score | 0.039433 (rank : 27) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P26651 | Gene names | ZFP36, G0S24, TIS11A, TTP | |||
|
Domain Architecture |
|
|||||
| Description | Tristetraproline (TTP) (Zinc finger protein 36 homolog) (Zfp-36) (TIS11A protein) (TIS11) (Growth factor-inducible nuclear protein NUP475) (G0/G1 switch regulatory protein 24). | |||||
|
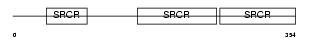
CD6_HUMAN
|
||||||
| θ value | 5.27518 (rank : 31) | NC score | 0.013646 (rank : 47) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P30203, Q9UMF2, Q9Y4K7, Q9Y4K8, Q9Y4K9, Q9Y4L0 | Gene names | CD6 | |||
|
Domain Architecture |
|
|||||
| Description | T-cell differentiation antigen CD6 precursor (T12) (TP120). | |||||
|
CDSN_MOUSE
|
||||||
| θ value | 5.27518 (rank : 32) | NC score | 0.031985 (rank : 33) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7TPC1 | Gene names | Cdsn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Corneodesmosin precursor. | |||||
|
DAB2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 33) | NC score | 0.021540 (rank : 41) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P98082, Q13598, Q9BTY0, Q9UK04 | Gene names | DAB2, DOC2 | |||
|
Domain Architecture |

|
|||||
| Description | Disabled homolog 2 (Differentially expressed protein 2) (DOC-2). | |||||
|
DHX57_MOUSE
|
||||||
| θ value | 5.27518 (rank : 34) | NC score | 0.034358 (rank : 31) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6P5D3, Q3TS93, Q6NZK4, Q6P1B4, Q8BI63, Q8BIA2, Q8R360 | Gene names | Dhx57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative ATP-dependent RNA helicase DHX57 (EC 3.6.1.-) (DEAH box protein 57). | |||||
|
HNF3A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 35) | NC score | 0.009422 (rank : 50) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P55317, Q9H2A0 | Gene names | FOXA1, HNF3A, TCF3A | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 3-alpha (HNF-3A) (Forkhead box protein A1). | |||||
|
PDXL2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 36) | NC score | 0.034464 (rank : 30) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8CAE9, Q8CFW3 | Gene names | Podxl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Podocalyxin-like protein 2 precursor (Endoglycan). | |||||
|
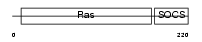
RB40B_MOUSE
|
||||||
| θ value | 5.27518 (rank : 37) | NC score | 0.007767 (rank : 51) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 265 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VHP8 | Gene names | Rab40b | |||
|
Domain Architecture |
|
|||||
| Description | Ras-related protein Rab-40B (SOCS box-containing protein RAR). | |||||
|
CRSP7_HUMAN
|
||||||
| θ value | 6.88961 (rank : 38) | NC score | 0.022420 (rank : 39) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O95402 | Gene names | CRSP7, ARC70 | |||
|
Domain Architecture |

|
|||||
| Description | CRSP complex subunit 7 (Cofactor required for Sp1 transcriptional activation subunit 7) (Transcriptional coactivator CRSP70) (Activator- recruited cofactor 70 kDa component) (ARC70). | |||||
|
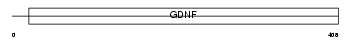
GFRA1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 39) | NC score | 0.013926 (rank : 46) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P56159, O15507, O43912 | Gene names | GFRA1, GDNFRA, RETL1, TRNR1 | |||
|
Domain Architecture |
|
|||||
| Description | GDNF family receptor alpha-1 precursor (GFR-alpha-1) (GDNF receptor alpha) (GDNFR-alpha) (TGF-beta-related neurotrophic factor receptor 1) (RET ligand 1). | |||||
|
LRC4B_HUMAN
|
||||||
| θ value | 6.88961 (rank : 40) | NC score | 0.004189 (rank : 54) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NT99, Q3ZCQ4, Q58F20 | Gene names | LRRC4B, LRIG4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 4B precursor. | |||||
|
MBD5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 41) | NC score | 0.022732 (rank : 38) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9P267, Q9NUV6 | Gene names | MBD5, KIAA1461 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 5 (Methyl-CpG-binding protein MBD5). | |||||
|
MKRN4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 42) | NC score | 0.066853 (rank : 14) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q13434 | Gene names | MKRN4, RNF64, ZNF127L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Makorin-4 (Zinc finger protein 127-Xp) (ZNF127-Xp) (RING finger protein 64). | |||||
|
MUC1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 43) | NC score | 0.055844 (rank : 17) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q02496 | Gene names | Muc1, Muc-1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (Polymorphic epithelial mucin) (PEMT) (Episialin). | |||||
|
PDXL2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 44) | NC score | 0.032734 (rank : 32) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NZ53, Q6UVY4, Q8WUV6 | Gene names | PODXL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Podocalyxin-like protein 2 precursor (Endoglycan). | |||||
|
PO121_MOUSE
|
||||||
| θ value | 6.88961 (rank : 45) | NC score | 0.062361 (rank : 15) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8K3Z9, Q7TSH5 | Gene names | Pom121, Nup121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear envelope pore membrane protein POM 121 (Pore membrane protein of 121 kDa). | |||||
|
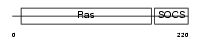
RB40B_HUMAN
|
||||||
| θ value | 6.88961 (rank : 46) | NC score | 0.007163 (rank : 53) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q12829, Q8WVG3 | Gene names | RAB40B, SEC4L | |||
|
Domain Architecture |
|
|||||
| Description | Ras-related protein Rab-40B (SOCS box-containing protein RAR) (Rar protein). | |||||
|
ZC3H6_MOUSE
|
||||||
| θ value | 6.88961 (rank : 47) | NC score | 0.059820 (rank : 16) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8BYK8, Q80UK3, Q8C1H1, Q9D604 | Gene names | Zc3h6, Zc3hdc6 | |||
|
Domain Architecture |
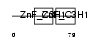
|
|||||
| Description | Zinc finger CCCH domain-containing protein 6. | |||||
|
CO5A2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 48) | NC score | 0.015601 (rank : 45) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 760 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P05997 | Gene names | COL5A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(V) chain precursor. | |||||
|
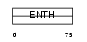
EPN3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 49) | NC score | 0.012794 (rank : 49) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q91W69, Q9CV55 | Gene names | Epn3 | |||
|
Domain Architecture |
|
|||||
| Description | Epsin-3 (EPS-15-interacting protein 3). | |||||
|
SOX15_HUMAN
|
||||||
| θ value | 8.99809 (rank : 50) | NC score | 0.007322 (rank : 52) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O60248, P35717, Q9Y6W7 | Gene names | SOX15, SOX12, SOX20, SOX26, SOX27 | |||
|
Domain Architecture |

|
|||||
| Description | SOX-15 protein (SOX-20 protein) (SOX-12 protein). | |||||
|
SUHW1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 51) | NC score | 0.003165 (rank : 55) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 648 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P59817 | Gene names | SUHW1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppressor of hairy wing homolog 1 (3'OY11.1). | |||||
|
TTC15_MOUSE
|
||||||
| θ value | 8.99809 (rank : 52) | NC score | 0.017154 (rank : 44) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8K2L8, Q8BX27 | Gene names | Ttc15 | |||
|
Domain Architecture |

|
|||||
| Description | Tetratricopeptide repeat protein 15 (TPR repeat protein 15). | |||||
|
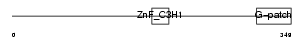
ZGPAT_HUMAN
|
||||||
| θ value | 8.99809 (rank : 53) | NC score | 0.040379 (rank : 24) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8N5A5, Q5JWI9, Q8NC55, Q96JI0, Q96JU4, Q9H401 | Gene names | ZGPAT, KIAA1847, ZC3H9, ZC3HDC9 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger CCCH-type with G patch domain protein (Zinc finger CCCH domain-containing protein 9). | |||||
|
ZGPAT_MOUSE
|
||||||
| θ value | 8.99809 (rank : 54) | NC score | 0.039473 (rank : 26) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8VDM1, Q8BWW2 | Gene names | Zgpat | |||
|
Domain Architecture |
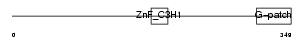
|
|||||
| Description | Zinc finger CCCH-type with G patch domain protein. | |||||
|
MKRN2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 55) | NC score | 0.050106 (rank : 19) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9ERV1, Q9D0L9 | Gene names | Mkrn2 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-2. | |||||
|
NUPL2_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q8CIC2, Q8BJX3, Q8BVP7, Q924S0 | Gene names | Nupl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoporin-like 2 (NLP-1). | |||||
|
NUPL2_HUMAN
|
||||||
| NC score | 0.976554 (rank : 2) | θ value | 1.42253e-146 (rank : 2) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O15504, Q49AE7, Q9BS49 | Gene names | NUPL2, CG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoporin-like 2 (NLP-1) (hCG1) (NUP42 homolog). | |||||
|
PAR12_MOUSE
|
||||||
| NC score | 0.094507 (rank : 3) | θ value | 0.365318 (rank : 7) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BZ20, Q80VL6, Q8K333, Q8R3U2 | Gene names | Parp12, Zc3hdc1 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 12 (EC 2.4.2.30) (PARP-12) (Zinc finger CCCH domain-containing protein 1). | |||||
|
PAR12_HUMAN
|
||||||
| NC score | 0.093345 (rank : 4) | θ value | 0.365318 (rank : 6) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9H0J9, Q9H610, Q9NP36, Q9NTI3 | Gene names | PARP12, ZC3HDC1 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 12 (EC 2.4.2.30) (PARP-12) (Zinc finger CCCH domain-containing protein 1). | |||||
|
NU153_HUMAN
|
||||||
| NC score | 0.078564 (rank : 5) | θ value | 0.279714 (rank : 5) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P49790 | Gene names | NUP153 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup153 (Nucleoporin Nup153) (153 kDa nucleoporin). | |||||
|
SC24C_HUMAN
|
||||||
| NC score | 0.071843 (rank : 6) | θ value | 0.0330416 (rank : 4) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 662 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P53992, Q8WV25 | Gene names | SEC24C, KIAA0079 | |||
|
Domain Architecture |

|
|||||
| Description | Protein transport protein Sec24C (SEC24-related protein C). | |||||
|
ZC3H6_HUMAN
|
||||||
| NC score | 0.070410 (rank : 7) | θ value | 3.0926 (rank : 21) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P61129 | Gene names | ZC3H6, KIAA2035, ZC3HDC6 | |||
|
Domain Architecture |
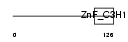
|
|||||
| Description | Zinc finger CCCH domain-containing protein 6. | |||||
|
CS007_MOUSE
|
||||||
| NC score | 0.070397 (rank : 8) | θ value | 4.03905 (rank : 24) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 464 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q6ZPZ3 | Gene names | Kiaa1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7 homolog. | |||||
|
CS007_HUMAN
|
||||||
| NC score | 0.069511 (rank : 9) | θ value | 4.03905 (rank : 23) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UPT8, Q9Y420 | Gene names | C19orf7, KIAA1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7. | |||||
|
MKRN3_MOUSE
|
||||||
| NC score | 0.069467 (rank : 10) | θ value | 3.0926 (rank : 18) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q60764, Q7TQE4 | Gene names | Mkrn3, Zfp127, Znf127 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-3 (Zinc finger protein 127). | |||||
|
MKRN2_HUMAN
|
||||||
| NC score | 0.068850 (rank : 11) | θ value | 3.0926 (rank : 17) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9H000, Q8N391, Q96BD4, Q9BUY2, Q9NRY1 | Gene names | MKRN2, RNF62 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-2 (RING finger protein 62). | |||||
|
MKRN1_MOUSE
|
||||||
| NC score | 0.067633 (rank : 12) | θ value | 4.03905 (rank : 27) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9QXP6 | Gene names | Mkrn1 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-1. | |||||
|
MKRN1_HUMAN
|
||||||
| NC score | 0.067310 (rank : 13) | θ value | 4.03905 (rank : 26) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9UHC7, Q6GSF1, Q9H0G0, Q9UEZ7 | Gene names | MKRN1, RNF61 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-1 (RING finger protein 61). | |||||
|
MKRN4_HUMAN
|
||||||
| NC score | 0.066853 (rank : 14) | θ value | 6.88961 (rank : 42) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q13434 | Gene names | MKRN4, RNF64, ZNF127L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Makorin-4 (Zinc finger protein 127-Xp) (ZNF127-Xp) (RING finger protein 64). | |||||
|
PO121_MOUSE
|
||||||
| NC score | 0.062361 (rank : 15) | θ value | 6.88961 (rank : 45) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8K3Z9, Q7TSH5 | Gene names | Pom121, Nup121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear envelope pore membrane protein POM 121 (Pore membrane protein of 121 kDa). | |||||
|
ZC3H6_MOUSE
|
||||||
| NC score | 0.059820 (rank : 16) | θ value | 6.88961 (rank : 47) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8BYK8, Q80UK3, Q8C1H1, Q9D604 | Gene names | Zc3h6, Zc3hdc6 | |||
|
Domain Architecture |
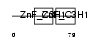
|
|||||
| Description | Zinc finger CCCH domain-containing protein 6. | |||||
|
MUC1_MOUSE
|
||||||
| NC score | 0.055844 (rank : 17) | θ value | 6.88961 (rank : 43) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q02496 | Gene names | Muc1, Muc-1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (Polymorphic epithelial mucin) (PEMT) (Episialin). | |||||
|
DHX57_HUMAN
|
||||||
| NC score | 0.050164 (rank : 18) | θ value | 1.38821 (rank : 10) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6P158, Q53SI4, Q6P9G1, Q7Z6H3, Q8NG17, Q96M33 | Gene names | DHX57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative ATP-dependent RNA helicase DHX57 (EC 3.6.1.-) (DEAH box protein 57). | |||||
|
MKRN2_MOUSE
|
||||||
| NC score | 0.050106 (rank : 19) | θ value | θ > 10 (rank : 55) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9ERV1, Q9D0L9 | Gene names | Mkrn2 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-2. | |||||
|
MUC1_HUMAN
|
||||||
| NC score | 0.043025 (rank : 20) | θ value | 0.00509761 (rank : 3) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
NAB2_MOUSE
|
||||||
| NC score | 0.042299 (rank : 21) | θ value | 3.0926 (rank : 20) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q61127 | Gene names | Nab2 | |||
|
Domain Architecture |

|
|||||
| Description | NGFI-A-binding protein 2 (EGR-1-binding protein 2). | |||||
|
NAB2_HUMAN
|
||||||
| NC score | 0.042012 (rank : 22) | θ value | 3.0926 (rank : 19) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15742, O76006, Q14797 | Gene names | NAB2, MADER | |||
|
Domain Architecture |

|
|||||
| Description | NGFI-A-binding protein 2 (EGR-1-binding protein 2) (Melanoma- associated delayed early response protein) (Protein MADER). | |||||
|
HRBL_MOUSE
|
||||||
| NC score | 0.040429 (rank : 23) | θ value | 1.06291 (rank : 9) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q80WC7, Q8BKS5, Q99J67 | Gene names | Hrbl, Rabr | |||
|
Domain Architecture |

|
|||||
| Description | HIV-1 Rev-binding protein-like protein (Rev/Rex activation domain- binding protein related) (RAB-R). | |||||
|
ZGPAT_HUMAN
|
||||||
| NC score | 0.040379 (rank : 24) | θ value | 8.99809 (rank : 53) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8N5A5, Q5JWI9, Q8NC55, Q96JI0, Q96JU4, Q9H401 | Gene names | ZGPAT, KIAA1847, ZC3H9, ZC3HDC9 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger CCCH-type with G patch domain protein (Zinc finger CCCH domain-containing protein 9). | |||||
|
TNR6A_HUMAN
|
||||||
| NC score | 0.040014 (rank : 25) | θ value | 1.81305 (rank : 13) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8NDV7, O15408, Q658L5, Q6NVB5, Q8NEZ0, Q8TBT8, Q8TCR0, Q9NV59, Q9P268 | Gene names | TNRC6A, CAGH26, KIAA1460, TNRC6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trinucleotide repeat-containing gene 6A protein (CAG repeat protein 26) (Glycine-tryptophan protein of 182 kDa) (GW182 autoantigen) (Protein GW1) (EMSY interactor protein). | |||||
|
ZGPAT_MOUSE
|
||||||
| NC score | 0.039473 (rank : 26) | θ value | 8.99809 (rank : 54) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8VDM1, Q8BWW2 | Gene names | Zgpat | |||
|
Domain Architecture |
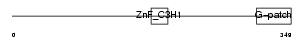
|
|||||
| Description | Zinc finger CCCH-type with G patch domain protein. | |||||
|
TTP_HUMAN
|
||||||
| NC score | 0.039433 (rank : 27) | θ value | 4.03905 (rank : 30) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P26651 | Gene names | ZFP36, G0S24, TIS11A, TTP | |||
|
Domain Architecture |

|
|||||
| Description | Tristetraproline (TTP) (Zinc finger protein 36 homolog) (Zfp-36) (TIS11A protein) (TIS11) (Growth factor-inducible nuclear protein NUP475) (G0/G1 switch regulatory protein 24). | |||||
|
CXXC6_HUMAN
|
||||||
| NC score | 0.036511 (rank : 28) | θ value | 3.0926 (rank : 16) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NFU7, Q5VUP7, Q7Z6B6, Q8TCR1, Q9C0I7 | Gene names | CXXC6, KIAA1676, LCX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CXXC-type zinc finger protein 6 (Leukemia-associated protein with a CXXC domain). | |||||
|
TPO_MOUSE
|
||||||
| NC score | 0.034929 (rank : 29) | θ value | 2.36792 (rank : 15) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P40226 | Gene names | Thpo | |||
|
Domain Architecture |

|
|||||
| Description | Thrombopoietin precursor (Megakaryocyte colony-stimulating factor) (Myeloproliferative leukemia virus oncogene ligand) (C-mpl ligand) (ML) (Megakaryocyte growth and development factor) (MGDF). | |||||
|
PDXL2_MOUSE
|
||||||
| NC score | 0.034464 (rank : 30) | θ value | 5.27518 (rank : 36) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8CAE9, Q8CFW3 | Gene names | Podxl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Podocalyxin-like protein 2 precursor (Endoglycan). | |||||
|
DHX57_MOUSE
|
||||||
| NC score | 0.034358 (rank : 31) | θ value | 5.27518 (rank : 34) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6P5D3, Q3TS93, Q6NZK4, Q6P1B4, Q8BI63, Q8BIA2, Q8R360 | Gene names | Dhx57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative ATP-dependent RNA helicase DHX57 (EC 3.6.1.-) (DEAH box protein 57). | |||||
|
PDXL2_HUMAN
|
||||||
| NC score | 0.032734 (rank : 32) | θ value | 6.88961 (rank : 44) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NZ53, Q6UVY4, Q8WUV6 | Gene names | PODXL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Podocalyxin-like protein 2 precursor (Endoglycan). | |||||
|
CDSN_MOUSE
|
||||||
| NC score | 0.031985 (rank : 33) | θ value | 5.27518 (rank : 32) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7TPC1 | Gene names | Cdsn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Corneodesmosin precursor. | |||||
|
TROP_HUMAN
|
||||||
| NC score | 0.031953 (rank : 34) | θ value | 0.47712 (rank : 8) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q12816, Q9NU89, Q9UPN8 | Gene names | TRO, KIAA1114, MAGED3 | |||
|
Domain Architecture |

|
|||||
| Description | Trophinin (MAGE-D3 antigen). | |||||
|
FBX42_HUMAN
|
||||||
| NC score | 0.031211 (rank : 35) | θ value | 1.81305 (rank : 11) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6P3S6, Q5TEU8, Q86XI0, Q8N3N4, Q8N5F8, Q9BRM0, Q9P2L4 | Gene names | FBXO42, FBX42, KIAA1332 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 42. | |||||
|
MCM3A_MOUSE
|
||||||
| NC score | 0.029155 (rank : 36) | θ value | 4.03905 (rank : 25) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9WUU9 | Gene names | Mcm3ap, Ganp, Map80 | |||
|
Domain Architecture |

|
|||||
| Description | 80 kDa MCM3-associated protein (GANP protein). | |||||
|
SET1A_HUMAN
|
||||||
| NC score | 0.026257 (rank : 37) | θ value | 2.36792 (rank : 14) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O15047, Q6PIF3, Q8TAJ6 | Gene names | SETD1A, KIAA0339, SET1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET1 (EC 2.1.1.43) (Set1/Ash2 histone methyltransferase complex subunit SET1) (SET-domain-containing protein 1A). | |||||
|
MBD5_HUMAN
|
||||||
| NC score | 0.022732 (rank : 38) | θ value | 6.88961 (rank : 41) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9P267, Q9NUV6 | Gene names | MBD5, KIAA1461 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 5 (Methyl-CpG-binding protein MBD5). | |||||
|
CRSP7_HUMAN
|
||||||
| NC score | 0.022420 (rank : 39) | θ value | 6.88961 (rank : 38) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O95402 | Gene names | CRSP7, ARC70 | |||
|
Domain Architecture |

|
|||||
| Description | CRSP complex subunit 7 (Cofactor required for Sp1 transcriptional activation subunit 7) (Transcriptional coactivator CRSP70) (Activator- recruited cofactor 70 kDa component) (ARC70). | |||||
|
SMG1_HUMAN
|
||||||
| NC score | 0.022037 (rank : 40) | θ value | 4.03905 (rank : 29) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96Q15, O43305, Q13284, Q8NFX2, Q96QV0, Q96RW3 | Gene names | SMG1, ATX, KIAA0421, LIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SMG1 (EC 2.7.11.1) (SMG-1) (hSMG-1) (Lambda/iota protein kinase C-interacting protein) (Lambda-interacting protein) (61E3.4). | |||||
|
DAB2_HUMAN
|
||||||
| NC score | 0.021540 (rank : 41) | θ value | 5.27518 (rank : 33) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P98082, Q13598, Q9BTY0, Q9UK04 | Gene names | DAB2, DOC2 | |||
|
Domain Architecture |

|
|||||
| Description | Disabled homolog 2 (Differentially expressed protein 2) (DOC-2). | |||||
|
NDF4_MOUSE
|
||||||
| NC score | 0.021380 (rank : 42) | θ value | 1.81305 (rank : 12) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O09105 | Gene names | Neurod4, Ath3, Atoh3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 4 (NeuroD4) (Atonal protein homolog 3) (Helix-loop-helix protein mATH-3) (MATH3). | |||||
|
AF10_MOUSE
|
||||||
| NC score | 0.017561 (rank : 43) | θ value | 4.03905 (rank : 22) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 182 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O54826 | Gene names | Mllt10, Af10 | |||
|
Domain Architecture |

|
|||||
| Description | Protein AF-10. | |||||
|
TTC15_MOUSE
|
||||||
| NC score | 0.017154 (rank : 44) | θ value | 8.99809 (rank : 52) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8K2L8, Q8BX27 | Gene names | Ttc15 | |||
|
Domain Architecture |

|
|||||
| Description | Tetratricopeptide repeat protein 15 (TPR repeat protein 15). | |||||
|
CO5A2_HUMAN
|
||||||
| NC score | 0.015601 (rank : 45) | θ value | 8.99809 (rank : 48) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 760 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P05997 | Gene names | COL5A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(V) chain precursor. | |||||
|
GFRA1_HUMAN
|
||||||
| NC score | 0.013926 (rank : 46) | θ value | 6.88961 (rank : 39) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P56159, O15507, O43912 | Gene names | GFRA1, GDNFRA, RETL1, TRNR1 | |||
|
Domain Architecture |

|
|||||
| Description | GDNF family receptor alpha-1 precursor (GFR-alpha-1) (GDNF receptor alpha) (GDNFR-alpha) (TGF-beta-related neurotrophic factor receptor 1) (RET ligand 1). | |||||
|
CD6_HUMAN
|
||||||
| NC score | 0.013646 (rank : 47) | θ value | 5.27518 (rank : 31) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P30203, Q9UMF2, Q9Y4K7, Q9Y4K8, Q9Y4K9, Q9Y4L0 | Gene names | CD6 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell differentiation antigen CD6 precursor (T12) (TP120). | |||||
|
SCYL2_MOUSE
|
||||||
| NC score | 0.013107 (rank : 48) | θ value | 4.03905 (rank : 28) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 603 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CFE4, Q3UT57, Q3UWU9, Q8K0M4 | Gene names | Scyl2, Cvak104, D10Ertd802e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SCY1-like protein 2 (Coated vesicle-associated kinase of 104 kDa). | |||||
|
EPN3_MOUSE
|
||||||
| NC score | 0.012794 (rank : 49) | θ value | 8.99809 (rank : 49) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q91W69, Q9CV55 | Gene names | Epn3 | |||
|
Domain Architecture |

|
|||||
| Description | Epsin-3 (EPS-15-interacting protein 3). | |||||
|
HNF3A_HUMAN
|
||||||
| NC score | 0.009422 (rank : 50) | θ value | 5.27518 (rank : 35) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P55317, Q9H2A0 | Gene names | FOXA1, HNF3A, TCF3A | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 3-alpha (HNF-3A) (Forkhead box protein A1). | |||||
|
RB40B_MOUSE
|
||||||
| NC score | 0.007767 (rank : 51) | θ value | 5.27518 (rank : 37) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 265 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VHP8 | Gene names | Rab40b | |||
|
Domain Architecture |

|
|||||
| Description | Ras-related protein Rab-40B (SOCS box-containing protein RAR). | |||||
|
SOX15_HUMAN
|
||||||
| NC score | 0.007322 (rank : 52) | θ value | 8.99809 (rank : 50) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O60248, P35717, Q9Y6W7 | Gene names | SOX15, SOX12, SOX20, SOX26, SOX27 | |||
|
Domain Architecture |

|
|||||
| Description | SOX-15 protein (SOX-20 protein) (SOX-12 protein). | |||||
|
RB40B_HUMAN
|
||||||
| NC score | 0.007163 (rank : 53) | θ value | 6.88961 (rank : 46) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q12829, Q8WVG3 | Gene names | RAB40B, SEC4L | |||
|
Domain Architecture |

|
|||||
| Description | Ras-related protein Rab-40B (SOCS box-containing protein RAR) (Rar protein). | |||||
|
LRC4B_HUMAN
|
||||||
| NC score | 0.004189 (rank : 54) | θ value | 6.88961 (rank : 40) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NT99, Q3ZCQ4, Q58F20 | Gene names | LRRC4B, LRIG4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 4B precursor. | |||||
|
SUHW1_HUMAN
|
||||||
| NC score | 0.003165 (rank : 55) | θ value | 8.99809 (rank : 51) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 648 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P59817 | Gene names | SUHW1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppressor of hairy wing homolog 1 (3'OY11.1). | |||||