Please be patient as the page loads
|
LOXL2_HUMAN
|
||||||
| SwissProt Accessions | Q9Y4K0, Q9BW70, Q9Y5Y8 | Gene names | LOXL2 | |||
|
Domain Architecture |

|
|||||
| Description | Lysyl oxidase homolog 2 precursor (EC 1.4.3.-) (Lysyl oxidase-like protein 2) (Lysyl oxidase-related protein 2) (Lysyl oxidase-related protein WS9-14). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
LOXL2_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9Y4K0, Q9BW70, Q9Y5Y8 | Gene names | LOXL2 | |||
|
Domain Architecture |

|
|||||
| Description | Lysyl oxidase homolog 2 precursor (EC 1.4.3.-) (Lysyl oxidase-like protein 2) (Lysyl oxidase-related protein 2) (Lysyl oxidase-related protein WS9-14). | |||||
|
LOXL2_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999206 (rank : 2) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P58022, Q8BRE6, Q8BS86, Q8C4K0, Q9JJ39 | Gene names | Loxl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lysyl oxidase homolog 2 precursor (EC 1.4.3.-) (Lysyl oxidase-like protein 2). | |||||
|
LOXL3_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.995369 (rank : 3) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P58215, Q96RS1 | Gene names | LOXL3, LOXL | |||
|
Domain Architecture |

|
|||||
| Description | Lysyl oxidase homolog 3 precursor (EC 1.4.3.-) (Lysyl oxidase-like protein 3). | |||||
|
LOXL3_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.994615 (rank : 5) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9Z175, Q9JJ39 | Gene names | Loxl3, Lor2 | |||
|
Domain Architecture |

|
|||||
| Description | Lysyl oxidase homolog 3 precursor (EC 1.4.3.-) (Lysyl oxidase-like protein 3) (Lysyl oxidase-related protein 2). | |||||
|
LOXL4_HUMAN
|
||||||
| θ value | 0 (rank : 5) | NC score | 0.994586 (rank : 6) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q96JB6, Q96DY1, Q96PC0, Q9H6T5 | Gene names | LOXL4, LOXC | |||
|
Domain Architecture |

|
|||||
| Description | Lysyl oxidase homolog 4 precursor (EC 1.4.3.-) (Lysyl oxidase-like protein 4) (Lysyl oxidase-related protein C). | |||||
|
LOXL4_MOUSE
|
||||||
| θ value | 0 (rank : 6) | NC score | 0.994863 (rank : 4) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q924C6 | Gene names | Loxl4, Loxc | |||
|
Domain Architecture |

|
|||||
| Description | Lysyl oxidase homolog 4 precursor (EC 1.4.3.-) (Lysyl oxidase-like protein 4) (Lysyl oxidase-related protein C). | |||||
|
DMBT1_HUMAN
|
||||||
| θ value | 1.96316e-71 (rank : 7) | NC score | 0.669619 (rank : 19) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UGM3, Q59EX0, Q5JR26, Q6MZN4, Q96DU4, Q9UGM2, Q9UJ57, Q9UKJ4, Q9Y211, Q9Y4V9 | Gene names | DMBT1, GP340 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Deleted in malignant brain tumors 1 protein precursor (Glycoprotein 340) (Gp-340) (Surfactant pulmonary-associated D-binding protein). | |||||
|
DMBT1_MOUSE
|
||||||
| θ value | 2.84138e-62 (rank : 8) | NC score | 0.610958 (rank : 21) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q60997, Q80YC6, Q9JMJ9 | Gene names | Dmbt1, Crpd | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Deleted in malignant brain tumors 1 protein precursor (CRP-ductin) (Vomeroglandin). | |||||
|
LYOX_MOUSE
|
||||||
| θ value | 1.23536e-57 (rank : 9) | NC score | 0.733753 (rank : 16) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P28301 | Gene names | Lox, Rrg | |||
|
Domain Architecture |
|
|||||
| Description | Protein-lysine 6-oxidase precursor (EC 1.4.3.13) (Lysyl oxidase) (RAS excision protein). | |||||
|
LYOX_HUMAN
|
||||||
| θ value | 2.10721e-57 (rank : 10) | NC score | 0.736739 (rank : 15) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P28300, Q5FWF0 | Gene names | LOX | |||
|
Domain Architecture |
|
|||||
| Description | Protein-lysine 6-oxidase precursor (EC 1.4.3.13) (Lysyl oxidase). | |||||
|
C163A_MOUSE
|
||||||
| θ value | 6.13105e-57 (rank : 11) | NC score | 0.822708 (rank : 9) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q2VLH6, Q2VLH5, Q99MX8 | Gene names | Cd163, M130 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Scavenger receptor cysteine-rich type 1 protein M130 precursor (CD163 antigen) [Contains: Soluble CD163 (sCD163)]. | |||||
|
LOXL1_MOUSE
|
||||||
| θ value | 3.974e-56 (rank : 12) | NC score | 0.716424 (rank : 18) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P97873, Q91ZY4, Q99KX3 | Gene names | Loxl1, Lox2, Loxl | |||
|
Domain Architecture |
|
|||||
| Description | Lysyl oxidase homolog 1 precursor (EC 1.4.3.-) (Lysyl oxidase-like protein 1) (Lysyl oxidase 2). | |||||
|
LOXL1_HUMAN
|
||||||
| θ value | 5.19021e-56 (rank : 13) | NC score | 0.718739 (rank : 17) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q08397, Q96BW7 | Gene names | LOXL1, LOXL | |||
|
Domain Architecture |

|
|||||
| Description | Lysyl oxidase homolog 1 precursor (EC 1.4.3.-) (Lysyl oxidase-like protein 1) (LOL). | |||||
|
C163A_HUMAN
|
||||||
| θ value | 4.39379e-55 (rank : 14) | NC score | 0.822181 (rank : 10) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q86VB7, Q07898, Q07899, Q07900, Q07901, Q2VLH7 | Gene names | CD163, M130 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Scavenger receptor cysteine-rich type 1 protein M130 precursor (CD163 antigen) (Hemoglobin scavenger receptor) [Contains: Soluble CD163 (sCD163)]. | |||||
|
NETR_MOUSE
|
||||||
| θ value | 5.73848e-55 (rank : 15) | NC score | 0.401148 (rank : 26) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O08762 | Gene names | Prss12, Bssp3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurotrypsin precursor (EC 3.4.21.-) (Motopsin) (Brain-specific serine protease 3) (BSSP-3). | |||||
|
NETR_HUMAN
|
||||||
| θ value | 1.66964e-54 (rank : 16) | NC score | 0.409015 (rank : 25) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P56730, Q9UP16 | Gene names | PRSS12 | |||
|
Domain Architecture |

|
|||||
| Description | Neurotrypsin precursor (EC 3.4.21.-) (Motopsin) (Leydin). | |||||
|
C163B_HUMAN
|
||||||
| θ value | 1.08223e-53 (rank : 17) | NC score | 0.818968 (rank : 11) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9NR16, Q6UWC2 | Gene names | CD163L1, CD163B, M160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Scavenger receptor cysteine-rich type 1 protein M160 precursor (CD163 antigen-like 1) (CD163b antigen). | |||||
|
CD5L_HUMAN
|
||||||
| θ value | 1.51715e-39 (rank : 18) | NC score | 0.840067 (rank : 7) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O43866, Q6UX63 | Gene names | CD5L | |||
|
Domain Architecture |
|
|||||
| Description | CD5 antigen-like precursor (SP-alpha) (CT-2) (IgM-associated peptide). | |||||
|
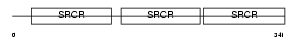
CD5L_MOUSE
|
||||||
| θ value | 1.42001e-37 (rank : 19) | NC score | 0.836471 (rank : 8) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9QWK4, O35300, O35301, Q505P6, Q91W05 | Gene names | Cd5l, Aim, Api6 | |||
|
Domain Architecture |
|
|||||
| Description | CD5 antigen-like precursor (SP-alpha) (CT-2) (Apoptosis inhibitory 6) (Apoptosis inhibitor expressed by macrophages). | |||||
|
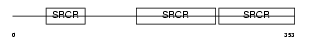
CD6_MOUSE
|
||||||
| θ value | 4.42448e-31 (rank : 20) | NC score | 0.796129 (rank : 13) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q61003, Q60679, Q61004 | Gene names | Cd6 | |||
|
Domain Architecture |
|
|||||
| Description | T-cell differentiation antigen CD6 precursor. | |||||
|
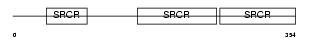
CD6_HUMAN
|
||||||
| θ value | 9.8567e-31 (rank : 21) | NC score | 0.802122 (rank : 12) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P30203, Q9UMF2, Q9Y4K7, Q9Y4K8, Q9Y4K9, Q9Y4L0 | Gene names | CD6 | |||
|
Domain Architecture |
|
|||||
| Description | T-cell differentiation antigen CD6 precursor (T12) (TP120). | |||||
|
MSRE_MOUSE
|
||||||
| θ value | 4.73814e-25 (rank : 22) | NC score | 0.622179 (rank : 20) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P30204, Q923G0, Q9QZ56 | Gene names | Msr1, Scvr | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage scavenger receptor types I and II (Macrophage acetylated LDL receptor I and II) (Scavenger receptor type A) (SR-A). | |||||
|
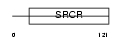
LG3BP_HUMAN
|
||||||
| θ value | 1.05554e-24 (rank : 23) | NC score | 0.784644 (rank : 14) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q08380 | Gene names | LGALS3BP, M2BP | |||
|
Domain Architecture |
|
|||||
| Description | Galectin-3-binding protein precursor (Lectin galactoside-binding soluble 3-binding protein) (Mac-2-binding protein) (Mac-2 BP) (MAC2BP) (Tumor-associated antigen 90K). | |||||
|
MARCO_HUMAN
|
||||||
| θ value | 4.43474e-23 (rank : 24) | NC score | 0.490000 (rank : 23) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UEW3, Q9Y5S3 | Gene names | MARCO, SCARA2 | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage receptor MARCO (Macrophage receptor with collagenous structure) (Scavenger receptor class A member 2). | |||||
|
MSRE_HUMAN
|
||||||
| θ value | 5.79196e-23 (rank : 25) | NC score | 0.568370 (rank : 22) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P21757, P21759, Q45F10 | Gene names | MSR1, SCARA1 | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage scavenger receptor types I and II (Macrophage acetylated LDL receptor I and II) (Scavenger receptor class A member 1) (CD204 antigen). | |||||
|
MARCO_MOUSE
|
||||||
| θ value | 7.56453e-23 (rank : 26) | NC score | 0.391268 (rank : 27) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q60754 | Gene names | Marco | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage receptor MARCO (Macrophage receptor with collagenous structure). | |||||
|
CFAI_MOUSE
|
||||||
| θ value | 0.00035302 (rank : 27) | NC score | 0.116250 (rank : 29) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q61129, Q9WU07 | Gene names | Cfi, If | |||
|
Domain Architecture |

|
|||||
| Description | Complement factor I precursor (EC 3.4.21.45) (C3B/C4B inactivator) [Contains: Complement factor I heavy chain; Complement factor I light chain]. | |||||
|
CFAI_HUMAN
|
||||||
| θ value | 0.00102713 (rank : 28) | NC score | 0.115302 (rank : 30) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P05156, O60442 | Gene names | CFI, IF | |||
|
Domain Architecture |

|
|||||
| Description | Complement factor I precursor (EC 3.4.21.45) (C3B/C4B inactivator) [Contains: Complement factor I heavy chain; Complement factor I light chain]. | |||||
|
ENTK_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 29) | NC score | 0.099133 (rank : 31) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 338 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P97435 | Gene names | Prss7, Entk | |||
|
Domain Architecture |

|
|||||
| Description | Enteropeptidase (EC 3.4.21.9) (Enterokinase) [Contains: Enteropeptidase non-catalytic heavy chain; Enteropeptidase catalytic light chain]. | |||||
|
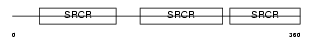
CD5_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 30) | NC score | 0.413634 (rank : 24) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P13379 | Gene names | Cd5, Ly-1 | |||
|
Domain Architecture |
|
|||||
| Description | T-cell surface glycoprotein CD5 precursor (Lymphocyte antigen Ly-1) (Lyt-1). | |||||
|
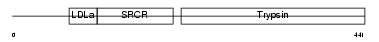
TMPS3_HUMAN
|
||||||
| θ value | 0.163984 (rank : 31) | NC score | 0.079884 (rank : 35) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P57727, Q6ZMC3 | Gene names | TMPRSS3, ECHOS1, TADG12 | |||
|
Domain Architecture |
|
|||||
| Description | Transmembrane protease, serine 3 (EC 3.4.21.-) (Serine protease TADG- 12) (Tumor-associated differentially-expressed gene 12 protein). | |||||
|
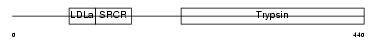
TMPS3_MOUSE
|
||||||
| θ value | 0.21417 (rank : 32) | NC score | 0.077876 (rank : 36) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8K1T0, Q8VDE0 | Gene names | Tmprss3 | |||
|
Domain Architecture |
|
|||||
| Description | Transmembrane protease, serine 3 (EC 3.4.21.-). | |||||
|
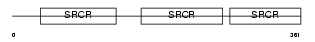
CD5_HUMAN
|
||||||
| θ value | 0.47712 (rank : 33) | NC score | 0.385444 (rank : 28) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P06127 | Gene names | CD5, LEU1 | |||
|
Domain Architecture |
|
|||||
| Description | T-cell surface glycoprotein CD5 precursor (Lymphocyte antigen T1/Leu- 1). | |||||
|
ENTK_HUMAN
|
||||||
| θ value | 0.813845 (rank : 34) | NC score | 0.083397 (rank : 34) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P98073 | Gene names | PRSS7, ENTK | |||
|
Domain Architecture |

|
|||||
| Description | Enteropeptidase precursor (EC 3.4.21.9) (Enterokinase) [Contains: Enteropeptidase non-catalytic heavy chain; Enteropeptidase catalytic light chain]. | |||||
|
LN28B_HUMAN
|
||||||
| θ value | 1.06291 (rank : 35) | NC score | 0.029219 (rank : 45) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6ZN17 | Gene names | LIN28B, CSDD2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lin-28 homolog B. | |||||
|
TMPSD_MOUSE
|
||||||
| θ value | 1.38821 (rank : 36) | NC score | 0.069287 (rank : 37) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 334 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q5U405, Q8CFE0, Q91VQ8 | Gene names | Tmprss13, Msp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protease, serine 13 (EC 3.4.21.-) (Mosaic serine protease) (Membrane-type mosaic serine protease). | |||||
|
ITA2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 37) | NC score | 0.004987 (rank : 54) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q62469, Q62163 | Gene names | Itga2 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-2 precursor (Platelet membrane glycoprotein Ia) (GPIa) (Collagen receptor) (VLA-2 alpha chain) (CD49b antigen). | |||||
|
NRX1A_MOUSE
|
||||||
| θ value | 1.81305 (rank : 38) | NC score | 0.010958 (rank : 51) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9CS84, O88722, Q80Y87, Q8CHE6 | Gene names | Nrxn1, Kiaa0578 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurexin-1-alpha precursor (Neurexin I-alpha). | |||||
|
NRX1A_HUMAN
|
||||||
| θ value | 3.0926 (rank : 39) | NC score | 0.010302 (rank : 52) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9ULB1, O60323, Q53TJ9, Q53TQ1, Q9C079, Q9C080, Q9C081, Q9H3M2, Q9UDM6 | Gene names | NRXN1, KIAA0578 | |||
|
Domain Architecture |

|
|||||
| Description | Neurexin-1-alpha precursor (Neurexin I-alpha). | |||||
|
NRX1B_HUMAN
|
||||||
| θ value | 3.0926 (rank : 40) | NC score | 0.015623 (rank : 48) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P58400 | Gene names | NRXN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurexin-1-beta precursor (Neurexin I-beta). | |||||
|
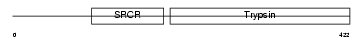
TMPS4_MOUSE
|
||||||
| θ value | 3.0926 (rank : 41) | NC score | 0.059917 (rank : 38) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8VCA5 | Gene names | Tmprss4, Cap2 | |||
|
Domain Architecture |
|
|||||
| Description | Transmembrane protease, serine 4 (EC 3.4.21.-) (Channel-activating protease 2) (mCAP2). | |||||
|
UBP20_HUMAN
|
||||||
| θ value | 3.0926 (rank : 42) | NC score | 0.008920 (rank : 53) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y2K6, Q96LG5, Q9UQN8, Q9UQP0 | Gene names | USP20, KIAA1003, LSFR3A | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 20 (EC 3.1.2.15) (Ubiquitin thioesterase 20) (Ubiquitin-specific-processing protease 20) (Deubiquitinating enzyme 20). | |||||
|
LN28A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 43) | NC score | 0.027915 (rank : 46) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H9Z2 | Gene names | LIN28, CSDD1, LIN28A, ZCCHC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lin-28 homolog A (Zinc finger CCHC domain-containing protein 1). | |||||
|
MIRO2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 44) | NC score | 0.004545 (rank : 55) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8JZN7 | Gene names | Rhot2, Arht2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitochondrial Rho GTPase 2 (EC 3.6.5.-) (MIRO-2) (Ras homolog gene family member T2). | |||||
|
TRMU_MOUSE
|
||||||
| θ value | 5.27518 (rank : 45) | NC score | 0.013527 (rank : 49) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 5 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9DAT5 | Gene names | Trmu, Trmt1 | |||
|
Domain Architecture |
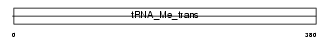
|
|||||
| Description | tRNA (5-methylaminomethyl-2-thiouridylate)-methyltransferase (EC 2.1.1.61). | |||||
|
LN28A_MOUSE
|
||||||
| θ value | 6.88961 (rank : 46) | NC score | 0.024119 (rank : 47) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8K3Y3, Q6NV62 | Gene names | Lin28, Lin28a, Tex17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lin-28 homolog A (Testis-expressed protein 17). | |||||
|
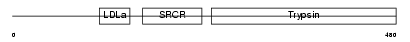
TMPS2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 47) | NC score | 0.050792 (rank : 43) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 252 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O15393, Q9BXX1 | Gene names | TMPRSS2, PRSS10 | |||
|
Domain Architecture |
|
|||||
| Description | Transmembrane protease, serine 2 precursor (EC 3.4.21.-) [Contains: Transmembrane protease, serine 2 non-catalytic chain; Transmembrane protease, serine 2 catalytic chain]. | |||||
|
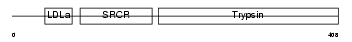
TMPS4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 48) | NC score | 0.049304 (rank : 44) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9NRS4, Q5XKQ6, Q6UX37, Q9NZA5 | Gene names | TMPRSS4, TMPRSS3 | |||
|
Domain Architecture |
|
|||||
| Description | Transmembrane protease, serine 4 (EC 3.4.21.-) (Membrane-type serine protease 2) (MT-SP2). | |||||
|
TRMU_HUMAN
|
||||||
| θ value | 8.99809 (rank : 49) | NC score | 0.011696 (rank : 50) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 3 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O75648, Q5W9C8 | Gene names | TRMU, MTU1, TRMT1 | |||
|
Domain Architecture |
|
|||||
| Description | tRNA (5-methylaminomethyl-2-thiouridylate)-methyltransferase (EC 2.1.1.61) (Mitochondrial tRNA-specific 2-thiouridylase 1) (MTO2 homolog). | |||||
|
CUZD1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 50) | NC score | 0.084189 (rank : 33) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q86UP6, Q7Z660, Q7Z661, Q86SG1, Q86UP5, Q9HAR7 | Gene names | CUZD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CUB and zona pellucida-like domain-containing protein 1 precursor (Transmembrane protein UO-44). | |||||
|
CUZD1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 51) | NC score | 0.087029 (rank : 32) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P70412, Q9CTZ7 | Gene names | Cuzd1, Itmap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CUB and zona pellucida-like domain-containing protein 1 precursor (Integral membrane-associated protein 1). | |||||
|
ST14_HUMAN
|
||||||
| θ value | θ > 10 (rank : 52) | NC score | 0.052127 (rank : 41) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9Y5Y6, Q9BS01, Q9H3S0, Q9HB36, Q9HCA3 | Gene names | ST14, PRSS14, SNC19, TADG15 | |||
|
Domain Architecture |

|
|||||
| Description | Suppressor of tumorigenicity protein 14 (EC 3.4.21.-) (Serine protease 14) (Matriptase) (Membrane-type serine protease 1) (MT-SP1) (Prostamin) (Serine protease TADG-15) (Tumor-associated differentially-expressed gene 15 protein). | |||||
|
ST14_MOUSE
|
||||||
| θ value | θ > 10 (rank : 53) | NC score | 0.053838 (rank : 39) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P56677 | Gene names | St14, Prss14 | |||
|
Domain Architecture |

|
|||||
| Description | Suppressor of tumorigenicity protein 14 (EC 3.4.21.-) (Serine protease 14) (Epithin). | |||||
|
TMPS5_MOUSE
|
||||||
| θ value | θ > 10 (rank : 54) | NC score | 0.052229 (rank : 40) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9ER04, Q9ER02, Q9ER03 | Gene names | Tmprss5 | |||
|
Domain Architecture |
|
|||||
| Description | Transmembrane protease, serine 5 (EC 3.4.21.-) (Spinesin). | |||||
|
TMPSD_HUMAN
|
||||||
| θ value | θ > 10 (rank : 55) | NC score | 0.051983 (rank : 42) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9BYE2, Q86YM4, Q96JY8, Q9BYE1 | Gene names | TMPRSS13, MSP, TMPRSS11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protease, serine 13 (EC 3.4.21.-) (Mosaic serine protease) (Membrane-type mosaic serine protease). | |||||
|
LOXL2_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9Y4K0, Q9BW70, Q9Y5Y8 | Gene names | LOXL2 | |||
|
Domain Architecture |

|
|||||
| Description | Lysyl oxidase homolog 2 precursor (EC 1.4.3.-) (Lysyl oxidase-like protein 2) (Lysyl oxidase-related protein 2) (Lysyl oxidase-related protein WS9-14). | |||||
|
LOXL2_MOUSE
|
||||||
| NC score | 0.999206 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P58022, Q8BRE6, Q8BS86, Q8C4K0, Q9JJ39 | Gene names | Loxl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lysyl oxidase homolog 2 precursor (EC 1.4.3.-) (Lysyl oxidase-like protein 2). | |||||
|
LOXL3_HUMAN
|
||||||
| NC score | 0.995369 (rank : 3) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P58215, Q96RS1 | Gene names | LOXL3, LOXL | |||
|
Domain Architecture |

|
|||||
| Description | Lysyl oxidase homolog 3 precursor (EC 1.4.3.-) (Lysyl oxidase-like protein 3). | |||||
|
LOXL4_MOUSE
|
||||||
| NC score | 0.994863 (rank : 4) | θ value | 0 (rank : 6) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q924C6 | Gene names | Loxl4, Loxc | |||
|
Domain Architecture |

|
|||||
| Description | Lysyl oxidase homolog 4 precursor (EC 1.4.3.-) (Lysyl oxidase-like protein 4) (Lysyl oxidase-related protein C). | |||||
|
LOXL3_MOUSE
|
||||||
| NC score | 0.994615 (rank : 5) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9Z175, Q9JJ39 | Gene names | Loxl3, Lor2 | |||
|
Domain Architecture |

|
|||||
| Description | Lysyl oxidase homolog 3 precursor (EC 1.4.3.-) (Lysyl oxidase-like protein 3) (Lysyl oxidase-related protein 2). | |||||
|
LOXL4_HUMAN
|
||||||
| NC score | 0.994586 (rank : 6) | θ value | 0 (rank : 5) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q96JB6, Q96DY1, Q96PC0, Q9H6T5 | Gene names | LOXL4, LOXC | |||
|
Domain Architecture |

|
|||||
| Description | Lysyl oxidase homolog 4 precursor (EC 1.4.3.-) (Lysyl oxidase-like protein 4) (Lysyl oxidase-related protein C). | |||||
|
CD5L_HUMAN
|
||||||
| NC score | 0.840067 (rank : 7) | θ value | 1.51715e-39 (rank : 18) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O43866, Q6UX63 | Gene names | CD5L | |||
|
Domain Architecture |

|
|||||
| Description | CD5 antigen-like precursor (SP-alpha) (CT-2) (IgM-associated peptide). | |||||
|
CD5L_MOUSE
|
||||||
| NC score | 0.836471 (rank : 8) | θ value | 1.42001e-37 (rank : 19) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9QWK4, O35300, O35301, Q505P6, Q91W05 | Gene names | Cd5l, Aim, Api6 | |||
|
Domain Architecture |

|
|||||
| Description | CD5 antigen-like precursor (SP-alpha) (CT-2) (Apoptosis inhibitory 6) (Apoptosis inhibitor expressed by macrophages). | |||||
|
C163A_MOUSE
|
||||||
| NC score | 0.822708 (rank : 9) | θ value | 6.13105e-57 (rank : 11) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q2VLH6, Q2VLH5, Q99MX8 | Gene names | Cd163, M130 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Scavenger receptor cysteine-rich type 1 protein M130 precursor (CD163 antigen) [Contains: Soluble CD163 (sCD163)]. | |||||
|
C163A_HUMAN
|
||||||
| NC score | 0.822181 (rank : 10) | θ value | 4.39379e-55 (rank : 14) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q86VB7, Q07898, Q07899, Q07900, Q07901, Q2VLH7 | Gene names | CD163, M130 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Scavenger receptor cysteine-rich type 1 protein M130 precursor (CD163 antigen) (Hemoglobin scavenger receptor) [Contains: Soluble CD163 (sCD163)]. | |||||
|
C163B_HUMAN
|
||||||
| NC score | 0.818968 (rank : 11) | θ value | 1.08223e-53 (rank : 17) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9NR16, Q6UWC2 | Gene names | CD163L1, CD163B, M160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Scavenger receptor cysteine-rich type 1 protein M160 precursor (CD163 antigen-like 1) (CD163b antigen). | |||||
|
CD6_HUMAN
|
||||||
| NC score | 0.802122 (rank : 12) | θ value | 9.8567e-31 (rank : 21) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P30203, Q9UMF2, Q9Y4K7, Q9Y4K8, Q9Y4K9, Q9Y4L0 | Gene names | CD6 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell differentiation antigen CD6 precursor (T12) (TP120). | |||||
|
CD6_MOUSE
|
||||||
| NC score | 0.796129 (rank : 13) | θ value | 4.42448e-31 (rank : 20) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q61003, Q60679, Q61004 | Gene names | Cd6 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell differentiation antigen CD6 precursor. | |||||
|
LG3BP_HUMAN
|
||||||
| NC score | 0.784644 (rank : 14) | θ value | 1.05554e-24 (rank : 23) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q08380 | Gene names | LGALS3BP, M2BP | |||
|
Domain Architecture |

|
|||||
| Description | Galectin-3-binding protein precursor (Lectin galactoside-binding soluble 3-binding protein) (Mac-2-binding protein) (Mac-2 BP) (MAC2BP) (Tumor-associated antigen 90K). | |||||
|
LYOX_HUMAN
|
||||||
| NC score | 0.736739 (rank : 15) | θ value | 2.10721e-57 (rank : 10) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P28300, Q5FWF0 | Gene names | LOX | |||
|
Domain Architecture |

|
|||||
| Description | Protein-lysine 6-oxidase precursor (EC 1.4.3.13) (Lysyl oxidase). | |||||
|
LYOX_MOUSE
|
||||||
| NC score | 0.733753 (rank : 16) | θ value | 1.23536e-57 (rank : 9) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P28301 | Gene names | Lox, Rrg | |||
|
Domain Architecture |

|
|||||
| Description | Protein-lysine 6-oxidase precursor (EC 1.4.3.13) (Lysyl oxidase) (RAS excision protein). | |||||
|
LOXL1_HUMAN
|
||||||
| NC score | 0.718739 (rank : 17) | θ value | 5.19021e-56 (rank : 13) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q08397, Q96BW7 | Gene names | LOXL1, LOXL | |||
|
Domain Architecture |

|
|||||
| Description | Lysyl oxidase homolog 1 precursor (EC 1.4.3.-) (Lysyl oxidase-like protein 1) (LOL). | |||||
|
LOXL1_MOUSE
|
||||||
| NC score | 0.716424 (rank : 18) | θ value | 3.974e-56 (rank : 12) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P97873, Q91ZY4, Q99KX3 | Gene names | Loxl1, Lox2, Loxl | |||
|
Domain Architecture |
|
|||||
| Description | Lysyl oxidase homolog 1 precursor (EC 1.4.3.-) (Lysyl oxidase-like protein 1) (Lysyl oxidase 2). | |||||
|
DMBT1_HUMAN
|
||||||
| NC score | 0.669619 (rank : 19) | θ value | 1.96316e-71 (rank : 7) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UGM3, Q59EX0, Q5JR26, Q6MZN4, Q96DU4, Q9UGM2, Q9UJ57, Q9UKJ4, Q9Y211, Q9Y4V9 | Gene names | DMBT1, GP340 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Deleted in malignant brain tumors 1 protein precursor (Glycoprotein 340) (Gp-340) (Surfactant pulmonary-associated D-binding protein). | |||||
|
MSRE_MOUSE
|
||||||
| NC score | 0.622179 (rank : 20) | θ value | 4.73814e-25 (rank : 22) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P30204, Q923G0, Q9QZ56 | Gene names | Msr1, Scvr | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage scavenger receptor types I and II (Macrophage acetylated LDL receptor I and II) (Scavenger receptor type A) (SR-A). | |||||
|
DMBT1_MOUSE
|
||||||
| NC score | 0.610958 (rank : 21) | θ value | 2.84138e-62 (rank : 8) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q60997, Q80YC6, Q9JMJ9 | Gene names | Dmbt1, Crpd | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Deleted in malignant brain tumors 1 protein precursor (CRP-ductin) (Vomeroglandin). | |||||
|
MSRE_HUMAN
|
||||||
| NC score | 0.568370 (rank : 22) | θ value | 5.79196e-23 (rank : 25) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P21757, P21759, Q45F10 | Gene names | MSR1, SCARA1 | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage scavenger receptor types I and II (Macrophage acetylated LDL receptor I and II) (Scavenger receptor class A member 1) (CD204 antigen). | |||||
|
MARCO_HUMAN
|
||||||
| NC score | 0.490000 (rank : 23) | θ value | 4.43474e-23 (rank : 24) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UEW3, Q9Y5S3 | Gene names | MARCO, SCARA2 | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage receptor MARCO (Macrophage receptor with collagenous structure) (Scavenger receptor class A member 2). | |||||
|
CD5_MOUSE
|
||||||
| NC score | 0.413634 (rank : 24) | θ value | 0.0961366 (rank : 30) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P13379 | Gene names | Cd5, Ly-1 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell surface glycoprotein CD5 precursor (Lymphocyte antigen Ly-1) (Lyt-1). | |||||
|
NETR_HUMAN
|
||||||
| NC score | 0.409015 (rank : 25) | θ value | 1.66964e-54 (rank : 16) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P56730, Q9UP16 | Gene names | PRSS12 | |||
|
Domain Architecture |

|
|||||
| Description | Neurotrypsin precursor (EC 3.4.21.-) (Motopsin) (Leydin). | |||||
|
NETR_MOUSE
|
||||||
| NC score | 0.401148 (rank : 26) | θ value | 5.73848e-55 (rank : 15) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O08762 | Gene names | Prss12, Bssp3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurotrypsin precursor (EC 3.4.21.-) (Motopsin) (Brain-specific serine protease 3) (BSSP-3). | |||||
|
MARCO_MOUSE
|
||||||
| NC score | 0.391268 (rank : 27) | θ value | 7.56453e-23 (rank : 26) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q60754 | Gene names | Marco | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage receptor MARCO (Macrophage receptor with collagenous structure). | |||||
|
CD5_HUMAN
|
||||||
| NC score | 0.385444 (rank : 28) | θ value | 0.47712 (rank : 33) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P06127 | Gene names | CD5, LEU1 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell surface glycoprotein CD5 precursor (Lymphocyte antigen T1/Leu- 1). | |||||
|
CFAI_MOUSE
|
||||||
| NC score | 0.116250 (rank : 29) | θ value | 0.00035302 (rank : 27) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q61129, Q9WU07 | Gene names | Cfi, If | |||
|
Domain Architecture |

|
|||||
| Description | Complement factor I precursor (EC 3.4.21.45) (C3B/C4B inactivator) [Contains: Complement factor I heavy chain; Complement factor I light chain]. | |||||
|
CFAI_HUMAN
|
||||||
| NC score | 0.115302 (rank : 30) | θ value | 0.00102713 (rank : 28) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P05156, O60442 | Gene names | CFI, IF | |||
|
Domain Architecture |

|
|||||
| Description | Complement factor I precursor (EC 3.4.21.45) (C3B/C4B inactivator) [Contains: Complement factor I heavy chain; Complement factor I light chain]. | |||||
|
ENTK_MOUSE
|
||||||
| NC score | 0.099133 (rank : 31) | θ value | 0.0252991 (rank : 29) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 338 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P97435 | Gene names | Prss7, Entk | |||
|
Domain Architecture |

|
|||||
| Description | Enteropeptidase (EC 3.4.21.9) (Enterokinase) [Contains: Enteropeptidase non-catalytic heavy chain; Enteropeptidase catalytic light chain]. | |||||
|
CUZD1_MOUSE
|
||||||
| NC score | 0.087029 (rank : 32) | θ value | θ > 10 (rank : 51) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P70412, Q9CTZ7 | Gene names | Cuzd1, Itmap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CUB and zona pellucida-like domain-containing protein 1 precursor (Integral membrane-associated protein 1). | |||||
|
CUZD1_HUMAN
|
||||||
| NC score | 0.084189 (rank : 33) | θ value | θ > 10 (rank : 50) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q86UP6, Q7Z660, Q7Z661, Q86SG1, Q86UP5, Q9HAR7 | Gene names | CUZD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CUB and zona pellucida-like domain-containing protein 1 precursor (Transmembrane protein UO-44). | |||||
|
ENTK_HUMAN
|
||||||
| NC score | 0.083397 (rank : 34) | θ value | 0.813845 (rank : 34) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P98073 | Gene names | PRSS7, ENTK | |||
|
Domain Architecture |

|
|||||
| Description | Enteropeptidase precursor (EC 3.4.21.9) (Enterokinase) [Contains: Enteropeptidase non-catalytic heavy chain; Enteropeptidase catalytic light chain]. | |||||
|
TMPS3_HUMAN
|
||||||
| NC score | 0.079884 (rank : 35) | θ value | 0.163984 (rank : 31) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P57727, Q6ZMC3 | Gene names | TMPRSS3, ECHOS1, TADG12 | |||
|
Domain Architecture |

|
|||||
| Description | Transmembrane protease, serine 3 (EC 3.4.21.-) (Serine protease TADG- 12) (Tumor-associated differentially-expressed gene 12 protein). | |||||
|
TMPS3_MOUSE
|
||||||
| NC score | 0.077876 (rank : 36) | θ value | 0.21417 (rank : 32) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8K1T0, Q8VDE0 | Gene names | Tmprss3 | |||
|
Domain Architecture |

|
|||||
| Description | Transmembrane protease, serine 3 (EC 3.4.21.-). | |||||
|
TMPSD_MOUSE
|
||||||
| NC score | 0.069287 (rank : 37) | θ value | 1.38821 (rank : 36) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 334 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q5U405, Q8CFE0, Q91VQ8 | Gene names | Tmprss13, Msp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protease, serine 13 (EC 3.4.21.-) (Mosaic serine protease) (Membrane-type mosaic serine protease). | |||||
|
TMPS4_MOUSE
|
||||||
| NC score | 0.059917 (rank : 38) | θ value | 3.0926 (rank : 41) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8VCA5 | Gene names | Tmprss4, Cap2 | |||
|
Domain Architecture |

|
|||||
| Description | Transmembrane protease, serine 4 (EC 3.4.21.-) (Channel-activating protease 2) (mCAP2). | |||||
|
ST14_MOUSE
|
||||||
| NC score | 0.053838 (rank : 39) | θ value | θ > 10 (rank : 53) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P56677 | Gene names | St14, Prss14 | |||
|
Domain Architecture |

|
|||||
| Description | Suppressor of tumorigenicity protein 14 (EC 3.4.21.-) (Serine protease 14) (Epithin). | |||||
|
TMPS5_MOUSE
|
||||||
| NC score | 0.052229 (rank : 40) | θ value | θ > 10 (rank : 54) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9ER04, Q9ER02, Q9ER03 | Gene names | Tmprss5 | |||
|
Domain Architecture |

|
|||||
| Description | Transmembrane protease, serine 5 (EC 3.4.21.-) (Spinesin). | |||||
|
ST14_HUMAN
|
||||||
| NC score | 0.052127 (rank : 41) | θ value | θ > 10 (rank : 52) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9Y5Y6, Q9BS01, Q9H3S0, Q9HB36, Q9HCA3 | Gene names | ST14, PRSS14, SNC19, TADG15 | |||
|
Domain Architecture |

|
|||||
| Description | Suppressor of tumorigenicity protein 14 (EC 3.4.21.-) (Serine protease 14) (Matriptase) (Membrane-type serine protease 1) (MT-SP1) (Prostamin) (Serine protease TADG-15) (Tumor-associated differentially-expressed gene 15 protein). | |||||
|
TMPSD_HUMAN
|
||||||
| NC score | 0.051983 (rank : 42) | θ value | θ > 10 (rank : 55) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9BYE2, Q86YM4, Q96JY8, Q9BYE1 | Gene names | TMPRSS13, MSP, TMPRSS11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protease, serine 13 (EC 3.4.21.-) (Mosaic serine protease) (Membrane-type mosaic serine protease). | |||||
|
TMPS2_HUMAN
|
||||||
| NC score | 0.050792 (rank : 43) | θ value | 8.99809 (rank : 47) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 252 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O15393, Q9BXX1 | Gene names | TMPRSS2, PRSS10 | |||
|
Domain Architecture |

|
|||||
| Description | Transmembrane protease, serine 2 precursor (EC 3.4.21.-) [Contains: Transmembrane protease, serine 2 non-catalytic chain; Transmembrane protease, serine 2 catalytic chain]. | |||||
|
TMPS4_HUMAN
|
||||||
| NC score | 0.049304 (rank : 44) | θ value | 8.99809 (rank : 48) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9NRS4, Q5XKQ6, Q6UX37, Q9NZA5 | Gene names | TMPRSS4, TMPRSS3 | |||
|
Domain Architecture |

|
|||||
| Description | Transmembrane protease, serine 4 (EC 3.4.21.-) (Membrane-type serine protease 2) (MT-SP2). | |||||
|
LN28B_HUMAN
|
||||||
| NC score | 0.029219 (rank : 45) | θ value | 1.06291 (rank : 35) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6ZN17 | Gene names | LIN28B, CSDD2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lin-28 homolog B. | |||||
|
LN28A_HUMAN
|
||||||
| NC score | 0.027915 (rank : 46) | θ value | 4.03905 (rank : 43) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H9Z2 | Gene names | LIN28, CSDD1, LIN28A, ZCCHC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lin-28 homolog A (Zinc finger CCHC domain-containing protein 1). | |||||
|
LN28A_MOUSE
|
||||||
| NC score | 0.024119 (rank : 47) | θ value | 6.88961 (rank : 46) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8K3Y3, Q6NV62 | Gene names | Lin28, Lin28a, Tex17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lin-28 homolog A (Testis-expressed protein 17). | |||||
|
NRX1B_HUMAN
|
||||||
| NC score | 0.015623 (rank : 48) | θ value | 3.0926 (rank : 40) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P58400 | Gene names | NRXN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurexin-1-beta precursor (Neurexin I-beta). | |||||
|
TRMU_MOUSE
|
||||||
| NC score | 0.013527 (rank : 49) | θ value | 5.27518 (rank : 45) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 5 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9DAT5 | Gene names | Trmu, Trmt1 | |||
|
Domain Architecture |
|
|||||
| Description | tRNA (5-methylaminomethyl-2-thiouridylate)-methyltransferase (EC 2.1.1.61). | |||||
|
TRMU_HUMAN
|
||||||
| NC score | 0.011696 (rank : 50) | θ value | 8.99809 (rank : 49) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 3 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O75648, Q5W9C8 | Gene names | TRMU, MTU1, TRMT1 | |||
|
Domain Architecture |
|
|||||
| Description | tRNA (5-methylaminomethyl-2-thiouridylate)-methyltransferase (EC 2.1.1.61) (Mitochondrial tRNA-specific 2-thiouridylase 1) (MTO2 homolog). | |||||
|
NRX1A_MOUSE
|
||||||
| NC score | 0.010958 (rank : 51) | θ value | 1.81305 (rank : 38) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9CS84, O88722, Q80Y87, Q8CHE6 | Gene names | Nrxn1, Kiaa0578 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurexin-1-alpha precursor (Neurexin I-alpha). | |||||
|
NRX1A_HUMAN
|
||||||
| NC score | 0.010302 (rank : 52) | θ value | 3.0926 (rank : 39) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9ULB1, O60323, Q53TJ9, Q53TQ1, Q9C079, Q9C080, Q9C081, Q9H3M2, Q9UDM6 | Gene names | NRXN1, KIAA0578 | |||
|
Domain Architecture |

|
|||||
| Description | Neurexin-1-alpha precursor (Neurexin I-alpha). | |||||
|
UBP20_HUMAN
|
||||||
| NC score | 0.008920 (rank : 53) | θ value | 3.0926 (rank : 42) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y2K6, Q96LG5, Q9UQN8, Q9UQP0 | Gene names | USP20, KIAA1003, LSFR3A | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 20 (EC 3.1.2.15) (Ubiquitin thioesterase 20) (Ubiquitin-specific-processing protease 20) (Deubiquitinating enzyme 20). | |||||
|
ITA2_MOUSE
|
||||||
| NC score | 0.004987 (rank : 54) | θ value | 1.81305 (rank : 37) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q62469, Q62163 | Gene names | Itga2 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin alpha-2 precursor (Platelet membrane glycoprotein Ia) (GPIa) (Collagen receptor) (VLA-2 alpha chain) (CD49b antigen). | |||||
|
MIRO2_MOUSE
|
||||||
| NC score | 0.004545 (rank : 55) | θ value | 4.03905 (rank : 44) | |||
| Query Neighborhood Hits | 49 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8JZN7 | Gene names | Rhot2, Arht2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitochondrial Rho GTPase 2 (EC 3.6.5.-) (MIRO-2) (Ras homolog gene family member T2). | |||||