Please be patient as the page loads
|
IP3KB_HUMAN
|
||||||
| SwissProt Accessions | P27987, Q5VWL9, Q5VWM0, Q96BZ2, Q96JS1, Q9UH47 | Gene names | ITPKB | |||
|
Domain Architecture |

|
|||||
| Description | Inositol-trisphosphate 3-kinase B (EC 2.7.1.127) (Inositol 1,4,5- trisphosphate 3-kinase B) (IP3K B) (IP3 3-kinase B) (IP3K-B). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
IP3KB_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 124 | |
| SwissProt Accessions | P27987, Q5VWL9, Q5VWM0, Q96BZ2, Q96JS1, Q9UH47 | Gene names | ITPKB | |||
|
Domain Architecture |

|
|||||
| Description | Inositol-trisphosphate 3-kinase B (EC 2.7.1.127) (Inositol 1,4,5- trisphosphate 3-kinase B) (IP3K B) (IP3 3-kinase B) (IP3K-B). | |||||
|
IP3KC_HUMAN
|
||||||
| θ value | 1.21547e-113 (rank : 2) | NC score | 0.928394 (rank : 4) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q96DU7, Q9UE25, Q9Y475 | Gene names | ITPKC, IP3KC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol-trisphosphate 3-kinase C (EC 2.7.1.127) (Inositol 1,4,5- trisphosphate 3-kinase C) (InsP 3-kinase C) (IP3K-C). | |||||
|
IP3KC_MOUSE
|
||||||
| θ value | 4.32305e-111 (rank : 3) | NC score | 0.926571 (rank : 5) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7TS72, Q3U384 | Gene names | Itpkc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol-trisphosphate 3-kinase C (EC 2.7.1.127) (Inositol 1,4,5- trisphosphate 3-kinase C) (IP3K-C). | |||||
|
IP3KA_MOUSE
|
||||||
| θ value | 3.65967e-110 (rank : 4) | NC score | 0.943481 (rank : 2) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8R071 | Gene names | Itpka | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol-trisphosphate 3-kinase A (EC 2.7.1.127) (Inositol 1,4,5- trisphosphate 3-kinase A) (IP3K A) (IP3 3-kinase A). | |||||
|
IP3KA_HUMAN
|
||||||
| θ value | 4.77967e-110 (rank : 5) | NC score | 0.938154 (rank : 3) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P23677, Q8TAN3 | Gene names | ITPKA | |||
|
Domain Architecture |

|
|||||
| Description | Inositol-trisphosphate 3-kinase A (EC 2.7.1.127) (Inositol 1,4,5- trisphosphate 3-kinase A) (IP3K A) (IP3 3-kinase A). | |||||
|
IPMK_HUMAN
|
||||||
| θ value | 0.000461057 (rank : 6) | NC score | 0.225220 (rank : 6) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8NFU5 | Gene names | IPMK, IMPK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol polyphosphate multikinase (EC 2.7.1.151) (Inositol 1,3,4,6- tetrakisphosphate 5-kinase). | |||||
|
IPMK_MOUSE
|
||||||
| θ value | 0.00134147 (rank : 7) | NC score | 0.221436 (rank : 7) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7TT16, Q8BZ11, Q8BZA8, Q9CWM9 | Gene names | Ipmk, Impk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol polyphosphate multikinase (EC 2.7.1.151) (Inositol 1,3,4,6- tetrakisphosphate 5-kinase). | |||||
|
IP6K2_HUMAN
|
||||||
| θ value | 0.00298849 (rank : 8) | NC score | 0.144439 (rank : 10) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UHH9, Q6P0N8, Q9BSZ6, Q9BUW3, Q9H4P7, Q9NT63, Q9UFU6 | Gene names | IHPK2, IP6K2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol hexakisphosphate kinase 2 (EC 2.7.4.21) (InsP6 kinase 2) (Inositol hexakisphosphate kinase 2) (P(i)-uptake stimulator) (PiUS). | |||||
|
IP6K2_MOUSE
|
||||||
| θ value | 0.00298849 (rank : 9) | NC score | 0.143812 (rank : 11) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q80V72 | Gene names | Ihpk2, Ip6k2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol hexakisphosphate kinase 2 (EC 2.7.4.21) (InsP6 kinase 2) (Inositol hexakisphosphate kinase 2) (P(i)-uptake stimulator) (PiUS). | |||||
|
NFH_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 10) | NC score | 0.039947 (rank : 41) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
IP6K3_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 11) | NC score | 0.150472 (rank : 9) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BWD2 | Gene names | Ihpk3, Ip6k3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol hexaphosphate kinase 3 (EC 2.7.4.21) (InsP6 kinase 3) (Inositol hexakisphosphate kinase 3). | |||||
|
EPN3_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 12) | NC score | 0.052260 (rank : 20) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9H201, Q9BVN6, Q9NWK2 | Gene names | EPN3 | |||
|
Domain Architecture |
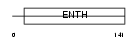
|
|||||
| Description | Epsin-3 (EPS-15-interacting protein 3). | |||||
|
COGA1_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 13) | NC score | 0.063687 (rank : 14) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q07092 | Gene names | COL16A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVI) chain precursor. | |||||
|
PDZD2_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 14) | NC score | 0.046913 (rank : 28) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O15018, Q9BXD4 | Gene names | PDZD2, AIPC, KIAA0300, PDZK3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 2 (PDZ domain-containing protein 3) (Activated in prostate cancer protein). | |||||
|
CEP35_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 15) | NC score | 0.041866 (rank : 38) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q5VT06, O75068, Q8TDK3, Q8WY20 | Gene names | CEP350, CAP350, GM133 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein 350 (Centrosome-associated protein of 350 kDa). | |||||
|
MILK1_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 16) | NC score | 0.038112 (rank : 48) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8N3F8, Q5TI16, Q7RTP5, Q8N3N8, Q9BVL9, Q9BY92, Q9UH43, Q9UH44, Q9UH45 | Gene names | MIRAB13, KIAA1668 | |||
|
Domain Architecture |
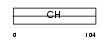
|
|||||
| Description | Molecule interacting with Rab13 (MIRab13) (MICAL-like protein 1). | |||||
|
IP6K3_HUMAN
|
||||||
| θ value | 0.125558 (rank : 17) | NC score | 0.155700 (rank : 8) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q96PC2, Q96MQ9 | Gene names | IHPK3, IP6K3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol hexaphosphate kinase 3 (EC 2.7.4.21) (InsP6 kinase 3) (Inositol hexakisphosphate kinase 3). | |||||
|
PRG4_HUMAN
|
||||||
| θ value | 0.163984 (rank : 18) | NC score | 0.045241 (rank : 32) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
ULK1_HUMAN
|
||||||
| θ value | 0.163984 (rank : 19) | NC score | 0.008627 (rank : 106) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1121 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O75385 | Gene names | ULK1 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase ULK1 (EC 2.7.11.1) (Unc-51-like kinase 1). | |||||
|
ABL1_MOUSE
|
||||||
| θ value | 0.21417 (rank : 20) | NC score | 0.006946 (rank : 111) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1178 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P00520, P97896, Q61252, Q61253, Q61254, Q61255, Q61256, Q61257, Q61258, Q61259, Q61260, Q61261, Q6PCM5, Q8C1X4 | Gene names | Abl1, Abl | |||
|
Domain Architecture |
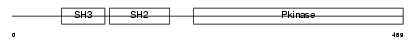
|
|||||
| Description | Proto-oncogene tyrosine-protein kinase ABL1 (EC 2.7.10.2) (p150) (c- ABL) (Abelson murine leukemia viral oncogene homolog 1). | |||||
|
ZDHC5_MOUSE
|
||||||
| θ value | 0.279714 (rank : 21) | NC score | 0.027491 (rank : 63) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8VDZ4, Q69ZB5, Q8R2X7 | Gene names | Zdhhc5, Kiaa1748 | |||
|
Domain Architecture |
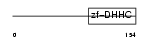
|
|||||
| Description | Probable palmitoyltransferase ZDHHC5 (EC 2.3.1.-) (Zinc finger DHHC domain-containing protein 5) (DHHC-5). | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 22) | NC score | 0.059337 (rank : 15) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
GP179_HUMAN
|
||||||
| θ value | 0.47712 (rank : 23) | NC score | 0.043034 (rank : 35) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 372 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6PRD1 | Gene names | GPR179, GPR158L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 179 precursor (Probable G-protein coupled receptor 158-like 1). | |||||
|
K1802_MOUSE
|
||||||
| θ value | 0.47712 (rank : 24) | NC score | 0.038834 (rank : 45) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
MAVS_MOUSE
|
||||||
| θ value | 0.47712 (rank : 25) | NC score | 0.050181 (rank : 23) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8VCF0 | Gene names | Mavs, Ips1, Visa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitochondrial antiviral signaling protein (Interferon-beta promoter stimulator protein 1) (IPS-1) (Virus-induced signaling adapter) (CARD adapter inducing interferon-beta) (Cardif). | |||||
|
PGCB_MOUSE
|
||||||
| θ value | 0.47712 (rank : 26) | NC score | 0.019174 (rank : 80) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 459 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q61361 | Gene names | Bcan | |||
|
Domain Architecture |

|
|||||
| Description | Brevican core protein precursor. | |||||
|
BIN1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 27) | NC score | 0.039515 (rank : 42) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O08539, Q62434 | Gene names | Bin1, Amphl, Sh3p9 | |||
|
Domain Architecture |
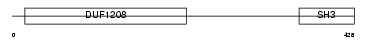
|
|||||
| Description | Myc box-dependent-interacting protein 1 (Bridging integrator 1) (Amphiphysin-like protein) (Amphiphysin II) (SH3-domain-containing protein 9). | |||||
|
CO7A1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 28) | NC score | 0.045231 (rank : 33) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 807 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q02388, Q14054, Q16507 | Gene names | COL7A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(VII) chain precursor (Long-chain collagen) (LC collagen). | |||||
|
FCN1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 29) | NC score | 0.025663 (rank : 68) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O00602, Q92596 | Gene names | FCN1 | |||
|
Domain Architecture |

|
|||||
| Description | Ficolin-1 precursor (Collagen/fibrinogen domain-containing protein 1) (Ficolin-A) (Ficolin A) (M-Ficolin). | |||||
|
PI5PA_HUMAN
|
||||||
| θ value | 0.813845 (rank : 30) | NC score | 0.038952 (rank : 44) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 749 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q15735, Q32M61, Q6ZTH6, Q8N902, Q9UDT9 | Gene names | PIB5PA, PIPP | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4,5-bisphosphate 5-phosphatase A (EC 3.1.3.56). | |||||
|
SRRM1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 31) | NC score | 0.047416 (rank : 27) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
ZN598_MOUSE
|
||||||
| θ value | 0.813845 (rank : 32) | NC score | 0.032855 (rank : 54) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q80YR4, Q6KAT0, Q8R3S1 | Gene names | Znf598, Zfp598 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 598. | |||||
|
ATX2L_HUMAN
|
||||||
| θ value | 1.06291 (rank : 33) | NC score | 0.029598 (rank : 60) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8WWM7, O95135, Q6NVJ8, Q6PJW6, Q8IU61, Q8IU95, Q8WWM3, Q8WWM4, Q8WWM5, Q8WWM6, Q99703 | Gene names | ATXN2L, A2D, A2LG, A2LP, A2RP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2-like protein (Ataxin-2 domain protein) (Ataxin-2-related protein). | |||||
|
CO1A1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 34) | NC score | 0.053217 (rank : 18) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 475 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P02452, P78441, Q13896, Q13902, Q13903, Q14037, Q14992, Q15176, Q15201, Q16050, Q7KZ30, Q7KZ34, Q8IVI5, Q9UML6, Q9UMM7 | Gene names | COL1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(I) chain precursor. | |||||
|
ETV4_MOUSE
|
||||||
| θ value | 1.06291 (rank : 35) | NC score | 0.015807 (rank : 92) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P28322 | Gene names | Etv4, Pea-3, Pea3 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 4 (Polyomavirus enhancer activator 3) (Protein PEA3). | |||||
|
PHLB1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 36) | NC score | 0.026885 (rank : 64) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 678 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q6PDH0, Q80TV2 | Gene names | Phldb1, Kiaa0638, Ll5a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family B member 1 (Protein LL5-alpha). | |||||
|
SDK1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 37) | NC score | 0.009429 (rank : 103) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 511 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q7Z5N4, Q8TEN9, Q8TEP5, Q96N44 | Gene names | SDK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein sidekick-1 precursor. | |||||
|
XKR7_HUMAN
|
||||||
| θ value | 1.06291 (rank : 38) | NC score | 0.027949 (rank : 62) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q5GH72, Q9NUG5 | Gene names | XKR7, C20orf159, XRG7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | XK-related protein 7. | |||||
|
CO1A1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 39) | NC score | 0.052128 (rank : 21) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P11087, Q53WT0, Q60635, Q61367, Q61427, Q63919, Q6PCL3, Q810J9 | Gene names | Col1a1, Cola1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(I) chain precursor. | |||||
|
JPH2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 40) | NC score | 0.030005 (rank : 58) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 639 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9BR39, O95913, Q5JY74, Q9UJN4 | Gene names | JPH2, JP2 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
CO8A2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 41) | NC score | 0.041751 (rank : 39) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P25318, Q68ED0, Q6KAQ4, Q6P1C4 | Gene names | Col8a2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-2(VIII) chain precursor (Endothelial collagen). | |||||
|
COHA1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 42) | NC score | 0.053084 (rank : 19) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q07563, Q99LK8 | Gene names | Col17a1, Bp180, Bpag2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVII) chain (Bullous pemphigoid antigen 2) (180 kDa bullous pemphigoid antigen 2). | |||||
|
TAF6_HUMAN
|
||||||
| θ value | 1.81305 (rank : 43) | NC score | 0.029836 (rank : 59) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P49848 | Gene names | TAF6, TAF2E, TAFII70 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 6 (Transcription initiation factor TFIID 70 kDa subunit) (TAF(II)70) (TAFII-70) (TAFII- 80) (TAFII80). | |||||
|
TR13B_MOUSE
|
||||||
| θ value | 1.81305 (rank : 44) | NC score | 0.038095 (rank : 49) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9ET35, Q9DBZ3 | Gene names | Tnfrsf13b, Taci | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor necrosis factor receptor superfamily member 13B (Transmembrane activator and CAML interactor). | |||||
|
CO3A1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 45) | NC score | 0.049971 (rank : 24) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P02461, Q15112, Q16403, Q6LDB3, Q6LDJ2, Q6LDJ3, Q7KZ56 | Gene names | COL3A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(III) chain precursor. | |||||
|
COBA2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 46) | NC score | 0.046696 (rank : 30) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 501 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q64739, Q61432, Q9Z1W0 | Gene names | Col11a2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(XI) chain precursor. | |||||
|
COHA1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 47) | NC score | 0.047729 (rank : 26) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 400 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9UMD9, Q02802, Q99018, Q9NQK9, Q9UC14 | Gene names | COL17A1, BP180, BPAG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XVII) chain (Bullous pemphigoid antigen 2) (180 kDa bullous pemphigoid antigen 2). | |||||
|
DPOG1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 48) | NC score | 0.023500 (rank : 72) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P54099, Q9JI28 | Gene names | Polg, Polg1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase subunit gamma 1 (EC 2.7.7.7) (Mitochondrial DNA polymerase catalytic subunit) (PolG-alpha). | |||||
|
IF4G1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 49) | NC score | 0.023426 (rank : 74) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 534 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q6NZJ6, Q6NZN8, Q8BW99 | Gene names | Eif4g1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1). | |||||
|
K1949_MOUSE
|
||||||
| θ value | 2.36792 (rank : 50) | NC score | 0.041910 (rank : 37) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8BQ30, Q3UDS6, Q8CEY9, Q8R3D8 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1949 homolog. | |||||
|
NCOA2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 51) | NC score | 0.021783 (rank : 76) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q61026, O09001, P97759 | Gene names | Ncoa2, Grip1, Tif2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2) (Glucocorticoid receptor-interacting protein 1) (GRIP-1). | |||||
|
PITM1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 52) | NC score | 0.015968 (rank : 91) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O00562, Q6T7X3, Q8TBN3, Q9BZ73 | Gene names | PITPNM1, DRES9, NIR2, PITPNM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated phosphatidylinositol transfer protein 1 (Phosphatidylinositol transfer protein, membrane-associated 1) (Pyk2 N-terminal domain-interacting receptor 2) (NIR-2) (Drosophila retinal degeneration B homolog). | |||||
|
SFR14_HUMAN
|
||||||
| θ value | 2.36792 (rank : 53) | NC score | 0.022698 (rank : 75) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IX01, O15071, O60369, Q5JPH7, Q8WUF7 | Gene names | SFRS14, KIAA0365 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative splicing factor, arginine/serine-rich 14 (Arginine/serine- rich-splicing factor 14). | |||||
|
CIC_HUMAN
|
||||||
| θ value | 3.0926 (rank : 54) | NC score | 0.025638 (rank : 69) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 992 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q96RK0, Q7LGI1, Q9UEG5, Q9Y6T1 | Gene names | CIC, KIAA0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
CO1A2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 55) | NC score | 0.050960 (rank : 22) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P08123, P02464, Q9UEB6, Q9UPH0 | Gene names | COL1A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(I) chain precursor. | |||||
|
CO2A1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 56) | NC score | 0.053250 (rank : 17) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 483 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P28481 | Gene names | Col2a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(II) chain precursor [Contains: Chondrocalcin]. | |||||
|
COBA2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 57) | NC score | 0.047962 (rank : 25) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P13942, Q07751, Q13271, Q13272, Q13273, Q99866, Q9UIP9 | Gene names | COL11A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(XI) chain precursor. | |||||
|
DAG1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 58) | NC score | 0.023433 (rank : 73) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q14118 | Gene names | DAG1 | |||
|
Domain Architecture |

|
|||||
| Description | Dystroglycan precursor (Dystrophin-associated glycoprotein 1) [Contains: Alpha-dystroglycan (Alpha-DG); Beta-dystroglycan (Beta- DG)]. | |||||
|
ENAM_MOUSE
|
||||||
| θ value | 3.0926 (rank : 59) | NC score | 0.028738 (rank : 61) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 229 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O55196 | Gene names | Enam | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Enamelin precursor. | |||||
|
NUP62_HUMAN
|
||||||
| θ value | 3.0926 (rank : 60) | NC score | 0.025764 (rank : 67) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 195 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P37198, Q503A4, Q96C43, Q9NSL1 | Gene names | NUP62 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore glycoprotein p62 (62 kDa nucleoporin). | |||||
|
PGCA_HUMAN
|
||||||
| θ value | 3.0926 (rank : 61) | NC score | 0.039470 (rank : 43) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P16112, Q13650, Q9UCP4, Q9UCP5, Q9UDE0 | Gene names | AGC1, CSPG1 | |||
|
Domain Architecture |

|
|||||
| Description | Aggrecan core protein precursor (Cartilage-specific proteoglycan core protein) (CSPCP) (Chondroitin sulfate proteoglycan core protein 1) [Contains: Aggrecan core protein 2]. | |||||
|
RAVR2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 62) | NC score | 0.014834 (rank : 94) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7TPD6 | Gene names | Raver2, Kiaa1579 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ribonucleoprotein PTB-binding 2 (Protein raver-2). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 63) | NC score | 0.044640 (rank : 34) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
ZN444_HUMAN
|
||||||
| θ value | 3.0926 (rank : 64) | NC score | 0.000494 (rank : 125) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 701 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8N0Y2, Q8TEQ9, Q8WU35, Q9NUU1 | Gene names | ZNF444, EZF2, ZSCAN17 | |||
|
Domain Architecture |
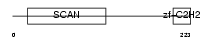
|
|||||
| Description | Zinc finger protein 444 (Endothelial zinc finger protein 2) (EZF-2) (Zinc finger and SCAN domain-containing protein 17). | |||||
|
BIN1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 65) | NC score | 0.037436 (rank : 50) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O00499, O00297, O00545, O43867, O60552, O60553, O60554, O60555, O75514, O75515, O75516, O75517, O75518, Q92944, Q99688 | Gene names | BIN1, AMPHL | |||
|
Domain Architecture |

|
|||||
| Description | Myc box-dependent-interacting protein 1 (Bridging integrator 1) (Amphiphysin-like protein) (Amphiphysin II) (Box-dependent myc- interacting protein 1). | |||||
|
CO2A1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 66) | NC score | 0.056231 (rank : 16) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P02458, Q12985, Q14009, Q14044, Q14046, Q14056, Q14058, Q16672, Q6LBY1, Q6LBY2, Q6LBY3, Q99227, Q9UE38, Q9UE39, Q9UE40, Q9UE41, Q9UE42, Q9UE43 | Gene names | COL2A1 | |||
|
Domain Architecture |
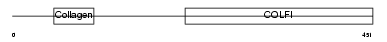
|
|||||
| Description | Collagen alpha-1(II) chain precursor [Contains: Chondrocalcin]. | |||||
|
CO4A6_HUMAN
|
||||||
| θ value | 4.03905 (rank : 67) | NC score | 0.046751 (rank : 29) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q14031, Q12823, Q14053, Q5JYH6, Q9NQM5, Q9NTX3, Q9UJ76, Q9UMG6, Q9Y4L4 | Gene names | COL4A6 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-6(IV) chain precursor. | |||||
|
EMID1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 68) | NC score | 0.038343 (rank : 47) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q96A84, Q6ICG1, Q86SS7 | Gene names | EMID1, EMU1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EMI domain-containing protein 1 precursor (Protein Emu1) (Emilin and multimerin domain-containing protein 1). | |||||
|
GSK3A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 69) | NC score | 0.002457 (rank : 123) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 835 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P49840, O14959 | Gene names | GSK3A | |||
|
Domain Architecture |

|
|||||
| Description | Glycogen synthase kinase-3 alpha (EC 2.7.11.26) (GSK-3 alpha). | |||||
|
IKBL_HUMAN
|
||||||
| θ value | 4.03905 (rank : 70) | NC score | 0.018048 (rank : 83) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UBC1, Q14625, Q9UBX4 | Gene names | NFKBIL1, IKBL | |||
|
Domain Architecture |
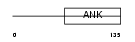
|
|||||
| Description | NF-kappa-B inhibitor-like protein 1 (Inhibitor of kappa B-like) (I- kappa-B-like) (IkappaBL) (Nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor-like 1). | |||||
|
INVS_MOUSE
|
||||||
| θ value | 4.03905 (rank : 71) | NC score | 0.009245 (rank : 104) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 609 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O89019, O88849 | Gene names | Invs, Inv, Nphp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inversin (Inversion of embryo turning protein) (Nephrocystin-2). | |||||
|
ITCH_HUMAN
|
||||||
| θ value | 4.03905 (rank : 72) | NC score | 0.006605 (rank : 115) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96J02, O43584, Q5TEL0, Q96F66, Q9BY75, Q9H451, Q9H4U5 | Gene names | ITCH | |||
|
Domain Architecture |

|
|||||
| Description | Itchy homolog E3 ubiquitin protein ligase (EC 6.3.2.-) (Itch) (Atrophin-1-interacting protein 4) (AIP4) (NFE2-associated polypeptide 1) (NAPP1). | |||||
|
PLIN_MOUSE
|
||||||
| θ value | 4.03905 (rank : 73) | NC score | 0.018836 (rank : 81) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CGN5 | Gene names | Plin, Peri | |||
|
Domain Architecture |

|
|||||
| Description | Perilipin (PERI) (Lipid droplet-associated protein). | |||||
|
TM115_MOUSE
|
||||||
| θ value | 4.03905 (rank : 74) | NC score | 0.024140 (rank : 71) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9WUH1, Q91VT6 | Gene names | Tmem115, Pl6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 115 (Protein PL6 homolog). | |||||
|
ZDHC5_HUMAN
|
||||||
| θ value | 4.03905 (rank : 75) | NC score | 0.020876 (rank : 77) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9C0B5, Q6ZMF0, Q8TAK8, Q9H923, Q9UFI7 | Gene names | ZDHHC5, KIAA1748, ZNF375 | |||
|
Domain Architecture |
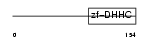
|
|||||
| Description | Probable palmitoyltransferase ZDHHC5 (EC 2.3.1.-) (Zinc finger DHHC domain-containing protein 5) (DHHC-5) (Zinc finger protein 375). | |||||
|
ARL4C_HUMAN
|
||||||
| θ value | 5.27518 (rank : 76) | NC score | 0.006864 (rank : 112) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P56559, Q9BVN1, Q9UQ34 | Gene names | ARL4C, ARL7 | |||
|
Domain Architecture |
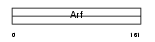
|
|||||
| Description | ADP-ribosylation factor-like protein 4C (ADP-ribosylation factor-like protein 7) (ADP-ribosylation factor-like protein LAK). | |||||
|
ARL4C_MOUSE
|
||||||
| θ value | 5.27518 (rank : 77) | NC score | 0.006864 (rank : 113) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P61208 | Gene names | Arl4c, Arl7 | |||
|
Domain Architecture |
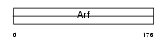
|
|||||
| Description | ADP-ribosylation factor-like protein 4C (ADP-ribosylation factor-like 7). | |||||
|
CAMLG_MOUSE
|
||||||
| θ value | 5.27518 (rank : 78) | NC score | 0.019776 (rank : 79) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P49070, Q99JU5 | Gene names | Camlg, Caml | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium signal-modulating cyclophilin ligand (CAML). | |||||
|
CO8A2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 79) | NC score | 0.040823 (rank : 40) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 299 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P25067, Q8TEJ5 | Gene names | COL8A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(VIII) chain precursor (Endothelial collagen). | |||||
|
COJA1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 80) | NC score | 0.046420 (rank : 31) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q14993, Q00559, Q05850, Q12885, Q13676, Q5JUF0, Q5T424, Q9H572, Q9NPZ2, Q9NQP2 | Gene names | COL19A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XIX) chain precursor (Collagen alpha-1(Y) chain). | |||||
|
K1949_HUMAN
|
||||||
| θ value | 5.27518 (rank : 81) | NC score | 0.033062 (rank : 53) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6NYC8, Q68CK8, Q6ZUJ6, Q8NDQ4, Q8TF52, Q9BRL9 | Gene names | KIAA1949 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1949. | |||||
|
RGS3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 82) | NC score | 0.025982 (rank : 66) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
RIOK2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 83) | NC score | 0.017225 (rank : 86) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9CQS5, Q91XF3 | Gene names | Riok2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase RIO2 (EC 2.7.11.1) (RIO kinase 2). | |||||
|
USBP1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 84) | NC score | 0.016614 (rank : 89) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8N6Y0, Q8NBX7, Q96KH3, Q9BYI8 | Gene names | USHBP1, AIEBP, MCC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USH1C-binding protein 1 (Usher syndrome type-1C protein-binding protein 1) (MCC-2) (AIE-75 binding protein). | |||||
|
XKR7_MOUSE
|
||||||
| θ value | 5.27518 (rank : 85) | NC score | 0.024380 (rank : 70) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q5GH64 | Gene names | Xkr7, Xrg7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | XK-related protein 7. | |||||
|
ZN358_HUMAN
|
||||||
| θ value | 5.27518 (rank : 86) | NC score | -0.000802 (rank : 126) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 840 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9NW07, Q9BTM7 | Gene names | ZNF358 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 358. | |||||
|
AP4B1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 87) | NC score | 0.013864 (rank : 96) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y6B7 | Gene names | AP4B1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-4 complex subunit beta-1 (Adapter-related protein complex 4 beta 1 subunit) (Beta subunit of AP-4) (AP-4 adapter complex beta subunit). | |||||
|
AP4B1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 88) | NC score | 0.013534 (rank : 97) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9WV76 | Gene names | Ap4b1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-4 complex subunit beta-1 (Adapter-related protein complex 4 beta-1 subunit) (Beta subunit of AP-4) (AP-4 adapter complex beta subunit). | |||||
|
ASPX_HUMAN
|
||||||
| θ value | 6.88961 (rank : 89) | NC score | 0.017321 (rank : 85) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P26436 | Gene names | ACRV1 | |||
|
Domain Architecture |

|
|||||
| Description | Acrosomal protein SP-10 precursor (Acrosomal vesicle protein 1). | |||||
|
BSPRY_MOUSE
|
||||||
| θ value | 6.88961 (rank : 90) | NC score | 0.007568 (rank : 109) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80YW5, Q3TU74, Q8BZF0, Q99KV7, Q9ER70 | Gene names | Bspry | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B box and SPRY domain-containing protein. | |||||
|
CO6A2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 91) | NC score | 0.038817 (rank : 46) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P12110, Q13909, Q13910, Q13911, Q14049, Q16259, Q16597, Q9UML3 | Gene names | COL6A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(VI) chain precursor. | |||||
|
COAA1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 92) | NC score | 0.036598 (rank : 51) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 316 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q03692 | Gene names | COL10A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(X) chain precursor. | |||||
|
COEA1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 93) | NC score | 0.030224 (rank : 57) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q80X19, Q8C6X3, Q9WV05 | Gene names | Col14a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XIV) chain precursor. | |||||
|
DAB2P_HUMAN
|
||||||
| θ value | 6.88961 (rank : 94) | NC score | 0.008966 (rank : 105) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5VWQ8, Q8TDL2, Q96SE1, Q9C0C0 | Gene names | DAB2IP, AF9Q34, AIP1, KIAA1743 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Disabled homolog 2-interacting protein (DAB2-interacting protein) (DAB2 interaction protein) (ASK-interacting protein 1). | |||||
|
EFHB_MOUSE
|
||||||
| θ value | 6.88961 (rank : 95) | NC score | 0.016620 (rank : 88) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CDU5 | Gene names | Efhb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EF-hand domain-containing family member B. | |||||
|
ERR1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 96) | NC score | 0.005642 (rank : 119) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P11474, Q14514 | Gene names | ESRRA, ERR1, ESRL1, NR3B1 | |||
|
Domain Architecture |
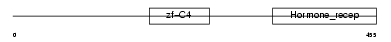
|
|||||
| Description | Steroid hormone receptor ERR1 (Estrogen-related receptor, alpha) (ERR- alpha) (Estrogen receptor-like 1). | |||||
|
FMN2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 97) | NC score | 0.017081 (rank : 87) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 484 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9NZ56, Q59GF6, Q5VU37, Q9NZ55 | Gene names | FMN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Formin-2. | |||||
|
GA2L2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 98) | NC score | 0.020277 (rank : 78) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q5SSG4 | Gene names | Gas2l2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GAS2-like protein 2 (Growth arrest-specific 2-like 2). | |||||
|
GDF7_MOUSE
|
||||||
| θ value | 6.88961 (rank : 99) | NC score | 0.005063 (rank : 120) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P43029, Q7TNX4, Q99MY1 | Gene names | Gdf7, Gdf-7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Growth/differentiation factor 7 precursor (GDF-7). | |||||
|
IF39_MOUSE
|
||||||
| θ value | 6.88961 (rank : 100) | NC score | 0.017745 (rank : 84) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8JZQ9, Q922K2 | Gene names | Eif3s9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 9 (eIF-3 eta) (eIF3 p116). | |||||
|
K0310_HUMAN
|
||||||
| θ value | 6.88961 (rank : 101) | NC score | 0.026441 (rank : 65) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O15027, Q5SXP0, Q5SXP1, Q8N347, Q96HP1 | Gene names | KIAA0310 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0310. | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 6.88961 (rank : 102) | NC score | 0.033895 (rank : 52) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
NCOR1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 103) | NC score | 0.014813 (rank : 95) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 521 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q60974, Q60812 | Gene names | Ncor1, Rxrip13 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR) (Retinoid X receptor- interacting protein 13) (RIP13). | |||||
|
PEF1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 104) | NC score | 0.010322 (rank : 101) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8BFY6, Q8VCT5, Q9CYW8, Q9D934 | Gene names | Pef1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peflin (PEF protein with a long N-terminal hydrophobic domain) (Penta- EF hand domain-containing protein 1). | |||||
|
PRGR_HUMAN
|
||||||
| θ value | 6.88961 (rank : 105) | NC score | 0.005846 (rank : 118) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P06401, Q9UPF7 | Gene names | PGR, NR3C3 | |||
|
Domain Architecture |

|
|||||
| Description | Progesterone receptor (PR). | |||||
|
S45A1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 106) | NC score | 0.012388 (rank : 99) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BIV7, Q3TZ98 | Gene names | Slc45a1, Dnb5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proton-associated sugar transporter A (PAST-A) (Solute carrier family 45 member 1) (Deleted in neuroblastoma 5 protein homolog) (DNb-5 homolog). | |||||
|
SSFA2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 107) | NC score | 0.018234 (rank : 82) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q922B9, Q8VEF7 | Gene names | Ssfa2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-specific antigen 2. | |||||
|
ST32C_HUMAN
|
||||||
| θ value | 6.88961 (rank : 108) | NC score | 0.001176 (rank : 124) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 832 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q86UX6, Q5T0Q5, Q86UE1 | Gene names | STK32C | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase 32C (EC 2.7.11.1) (PKE) (YANK3). | |||||
|
ZN592_HUMAN
|
||||||
| θ value | 6.88961 (rank : 109) | NC score | 0.003464 (rank : 122) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 762 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q92610 | Gene names | ZNF592, KIAA0211 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 592. | |||||
|
AZI1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 110) | NC score | 0.006265 (rank : 117) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1000 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UPN4, Q96F50 | Gene names | AZI1, CEP131, KIAA1118 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 5-azacytidine-induced protein 1 (Pre-acrosome localization protein 1) (Centrosomal protein of 131 kDa) (Cep131 protein). | |||||
|
BSN_MOUSE
|
||||||
| θ value | 8.99809 (rank : 111) | NC score | 0.030268 (rank : 56) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
CCD13_HUMAN
|
||||||
| θ value | 8.99809 (rank : 112) | NC score | 0.003930 (rank : 121) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 721 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8IYE1 | Gene names | CCDC13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 13. | |||||
|
CO4A4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 113) | NC score | 0.042563 (rank : 36) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 440 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P53420 | Gene names | COL4A4 | |||
|
Domain Architecture |
|
|||||
| Description | Collagen alpha-4(IV) chain precursor. | |||||
|
COEA1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 114) | NC score | 0.032542 (rank : 55) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q05707, O00260, O00261, O00262, Q05708, Q5XJ18, Q96C67 | Gene names | COL14A1, UND | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XIV) chain precursor (Undulin). | |||||
|
EXO1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 115) | NC score | 0.012178 (rank : 100) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UQ84, O60545, O75214, O75466, Q5T396, Q96IJ1, Q9UG38, Q9UNW0 | Gene names | EXO1, EXOI, HEX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Exonuclease 1 (EC 3.1.-.-) (hExo1) (Exonuclease I) (hExoI). | |||||
|
FBX30_MOUSE
|
||||||
| θ value | 8.99809 (rank : 116) | NC score | 0.009920 (rank : 102) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BJL1, Q8CI21, Q9D9X5 | Gene names | Fbxo30 | |||
|
Domain Architecture |

|
|||||
| Description | F-box only protein 30. | |||||
|
FXL22_HUMAN
|
||||||
| θ value | 8.99809 (rank : 117) | NC score | 0.016354 (rank : 90) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6P050 | Gene names | FBXL22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 22 (F-box and leucine-rich repeat protein 22). | |||||
|
PER1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 118) | NC score | 0.013251 (rank : 98) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O35973 | Gene names | Per1, Per, Rigui | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (mPER) (M-Rigui). | |||||
|
PODO_HUMAN
|
||||||
| θ value | 8.99809 (rank : 119) | NC score | 0.008564 (rank : 107) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NP85, Q8N6Q5 | Gene names | NPHS2 | |||
|
Domain Architecture |
|
|||||
| Description | Podocin. | |||||
|
RED1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 120) | NC score | 0.007947 (rank : 108) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q91ZS8, Q8K3X1, Q91ZS6, Q91ZS7, Q91ZS9, Q99MU8 | Gene names | Adarb1, Adar2, Red1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double-stranded RNA-specific editase 1 (EC 3.5.-.-) (dsRNA adenosine deaminase) (RNA-editing deaminase 1) (RNA-editing enzyme 1). | |||||
|
SPEG_MOUSE
|
||||||
| θ value | 8.99809 (rank : 121) | NC score | 0.006426 (rank : 116) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
TCGAP_HUMAN
|
||||||
| θ value | 8.99809 (rank : 122) | NC score | 0.015605 (rank : 93) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 872 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O14559, O14552, O14560, Q6ZSP6, Q96CP3, Q9NT23 | Gene names | SNX26, TCGAP | |||
|
Domain Architecture |

|
|||||
| Description | TC10/CDC42 GTPase-activating protein (Sorting nexin-26). | |||||
|
TSH3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 123) | NC score | 0.006761 (rank : 114) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 564 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q63HK5, Q9H0G6, Q9P254 | Gene names | TSHZ3, KIAA1474, TSH3, ZNF537 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 3 (Zinc finger protein 537). | |||||
|
UBP29_MOUSE
|
||||||
| θ value | 8.99809 (rank : 124) | NC score | 0.007173 (rank : 110) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9ES63 | Gene names | Usp29 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 29 (EC 3.1.2.15) (Ubiquitin thioesterase 29) (Ubiquitin-specific-processing protease 29) (Deubiquitinating enzyme 29). | |||||
|
IP6K1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 125) | NC score | 0.096990 (rank : 13) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q92551, Q7L3I7, Q96E38 | Gene names | IHPK1, IP6K1, KIAA0263 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol hexaphosphate kinase 1 (EC 2.7.4.21) (InsP6 kinase 1) (Inositol hexakisphosphate kinase 1). | |||||
|
IP6K1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 126) | NC score | 0.097171 (rank : 12) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6PD10, Q9QXV6 | Gene names | Ihpk1, Ip6k1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol hexaphosphate kinase 1 (EC 2.7.4.21) (InsP6 kinase 1) (Inositol hexakisphosphate kinase 1). | |||||
|
IP3KB_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 124 | |
| SwissProt Accessions | P27987, Q5VWL9, Q5VWM0, Q96BZ2, Q96JS1, Q9UH47 | Gene names | ITPKB | |||
|
Domain Architecture |

|
|||||
| Description | Inositol-trisphosphate 3-kinase B (EC 2.7.1.127) (Inositol 1,4,5- trisphosphate 3-kinase B) (IP3K B) (IP3 3-kinase B) (IP3K-B). | |||||
|
IP3KA_MOUSE
|
||||||
| NC score | 0.943481 (rank : 2) | θ value | 3.65967e-110 (rank : 4) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8R071 | Gene names | Itpka | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol-trisphosphate 3-kinase A (EC 2.7.1.127) (Inositol 1,4,5- trisphosphate 3-kinase A) (IP3K A) (IP3 3-kinase A). | |||||
|
IP3KA_HUMAN
|
||||||
| NC score | 0.938154 (rank : 3) | θ value | 4.77967e-110 (rank : 5) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P23677, Q8TAN3 | Gene names | ITPKA | |||
|
Domain Architecture |

|
|||||
| Description | Inositol-trisphosphate 3-kinase A (EC 2.7.1.127) (Inositol 1,4,5- trisphosphate 3-kinase A) (IP3K A) (IP3 3-kinase A). | |||||
|
IP3KC_HUMAN
|
||||||
| NC score | 0.928394 (rank : 4) | θ value | 1.21547e-113 (rank : 2) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q96DU7, Q9UE25, Q9Y475 | Gene names | ITPKC, IP3KC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol-trisphosphate 3-kinase C (EC 2.7.1.127) (Inositol 1,4,5- trisphosphate 3-kinase C) (InsP 3-kinase C) (IP3K-C). | |||||
|
IP3KC_MOUSE
|
||||||
| NC score | 0.926571 (rank : 5) | θ value | 4.32305e-111 (rank : 3) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7TS72, Q3U384 | Gene names | Itpkc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol-trisphosphate 3-kinase C (EC 2.7.1.127) (Inositol 1,4,5- trisphosphate 3-kinase C) (IP3K-C). | |||||
|
IPMK_HUMAN
|
||||||
| NC score | 0.225220 (rank : 6) | θ value | 0.000461057 (rank : 6) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8NFU5 | Gene names | IPMK, IMPK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol polyphosphate multikinase (EC 2.7.1.151) (Inositol 1,3,4,6- tetrakisphosphate 5-kinase). | |||||
|
IPMK_MOUSE
|
||||||
| NC score | 0.221436 (rank : 7) | θ value | 0.00134147 (rank : 7) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7TT16, Q8BZ11, Q8BZA8, Q9CWM9 | Gene names | Ipmk, Impk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol polyphosphate multikinase (EC 2.7.1.151) (Inositol 1,3,4,6- tetrakisphosphate 5-kinase). | |||||
|
IP6K3_HUMAN
|
||||||
| NC score | 0.155700 (rank : 8) | θ value | 0.125558 (rank : 17) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q96PC2, Q96MQ9 | Gene names | IHPK3, IP6K3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol hexaphosphate kinase 3 (EC 2.7.4.21) (InsP6 kinase 3) (Inositol hexakisphosphate kinase 3). | |||||
|
IP6K3_MOUSE
|
||||||
| NC score | 0.150472 (rank : 9) | θ value | 0.0113563 (rank : 11) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BWD2 | Gene names | Ihpk3, Ip6k3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol hexaphosphate kinase 3 (EC 2.7.4.21) (InsP6 kinase 3) (Inositol hexakisphosphate kinase 3). | |||||
|
IP6K2_HUMAN
|
||||||
| NC score | 0.144439 (rank : 10) | θ value | 0.00298849 (rank : 8) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UHH9, Q6P0N8, Q9BSZ6, Q9BUW3, Q9H4P7, Q9NT63, Q9UFU6 | Gene names | IHPK2, IP6K2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol hexakisphosphate kinase 2 (EC 2.7.4.21) (InsP6 kinase 2) (Inositol hexakisphosphate kinase 2) (P(i)-uptake stimulator) (PiUS). | |||||
|
IP6K2_MOUSE
|
||||||
| NC score | 0.143812 (rank : 11) | θ value | 0.00298849 (rank : 9) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q80V72 | Gene names | Ihpk2, Ip6k2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol hexakisphosphate kinase 2 (EC 2.7.4.21) (InsP6 kinase 2) (Inositol hexakisphosphate kinase 2) (P(i)-uptake stimulator) (PiUS). | |||||
|
IP6K1_MOUSE
|
||||||
| NC score | 0.097171 (rank : 12) | θ value | θ > 10 (rank : 126) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6PD10, Q9QXV6 | Gene names | Ihpk1, Ip6k1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol hexaphosphate kinase 1 (EC 2.7.4.21) (InsP6 kinase 1) (Inositol hexakisphosphate kinase 1). | |||||
|
IP6K1_HUMAN
|
||||||
| NC score | 0.096990 (rank : 13) | θ value | θ > 10 (rank : 125) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q92551, Q7L3I7, Q96E38 | Gene names | IHPK1, IP6K1, KIAA0263 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol hexaphosphate kinase 1 (EC 2.7.4.21) (InsP6 kinase 1) (Inositol hexakisphosphate kinase 1). | |||||
|
COGA1_HUMAN
|
||||||
| NC score | 0.063687 (rank : 14) | θ value | 0.0252991 (rank : 13) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q07092 | Gene names | COL16A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVI) chain precursor. | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.059337 (rank : 15) | θ value | 0.365318 (rank : 22) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
CO2A1_HUMAN
|
||||||
| NC score | 0.056231 (rank : 16) | θ value | 4.03905 (rank : 66) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P02458, Q12985, Q14009, Q14044, Q14046, Q14056, Q14058, Q16672, Q6LBY1, Q6LBY2, Q6LBY3, Q99227, Q9UE38, Q9UE39, Q9UE40, Q9UE41, Q9UE42, Q9UE43 | Gene names | COL2A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(II) chain precursor [Contains: Chondrocalcin]. | |||||
|
CO2A1_MOUSE
|
||||||
| NC score | 0.053250 (rank : 17) | θ value | 3.0926 (rank : 56) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 483 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P28481 | Gene names | Col2a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(II) chain precursor [Contains: Chondrocalcin]. | |||||
|
CO1A1_HUMAN
|
||||||
| NC score | 0.053217 (rank : 18) | θ value | 1.06291 (rank : 34) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 475 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P02452, P78441, Q13896, Q13902, Q13903, Q14037, Q14992, Q15176, Q15201, Q16050, Q7KZ30, Q7KZ34, Q8IVI5, Q9UML6, Q9UMM7 | Gene names | COL1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(I) chain precursor. | |||||
|
COHA1_MOUSE
|
||||||
| NC score | 0.053084 (rank : 19) | θ value | 1.81305 (rank : 42) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q07563, Q99LK8 | Gene names | Col17a1, Bp180, Bpag2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVII) chain (Bullous pemphigoid antigen 2) (180 kDa bullous pemphigoid antigen 2). | |||||
|
EPN3_HUMAN
|
||||||
| NC score | 0.052260 (rank : 20) | θ value | 0.0148317 (rank : 12) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9H201, Q9BVN6, Q9NWK2 | Gene names | EPN3 | |||
|
Domain Architecture |

|
|||||
| Description | Epsin-3 (EPS-15-interacting protein 3). | |||||
|
CO1A1_MOUSE
|
||||||
| NC score | 0.052128 (rank : 21) | θ value | 1.38821 (rank : 39) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P11087, Q53WT0, Q60635, Q61367, Q61427, Q63919, Q6PCL3, Q810J9 | Gene names | Col1a1, Cola1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(I) chain precursor. | |||||
|
CO1A2_HUMAN
|
||||||
| NC score | 0.050960 (rank : 22) | θ value | 3.0926 (rank : 55) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P08123, P02464, Q9UEB6, Q9UPH0 | Gene names | COL1A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(I) chain precursor. | |||||
|
MAVS_MOUSE
|
||||||
| NC score | 0.050181 (rank : 23) | θ value | 0.47712 (rank : 25) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8VCF0 | Gene names | Mavs, Ips1, Visa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitochondrial antiviral signaling protein (Interferon-beta promoter stimulator protein 1) (IPS-1) (Virus-induced signaling adapter) (CARD adapter inducing interferon-beta) (Cardif). | |||||
|
CO3A1_HUMAN
|
||||||
| NC score | 0.049971 (rank : 24) | θ value | 2.36792 (rank : 45) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P02461, Q15112, Q16403, Q6LDB3, Q6LDJ2, Q6LDJ3, Q7KZ56 | Gene names | COL3A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(III) chain precursor. | |||||
|
COBA2_HUMAN
|
||||||
| NC score | 0.047962 (rank : 25) | θ value | 3.0926 (rank : 57) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P13942, Q07751, Q13271, Q13272, Q13273, Q99866, Q9UIP9 | Gene names | COL11A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(XI) chain precursor. | |||||
|
COHA1_HUMAN
|
||||||
| NC score | 0.047729 (rank : 26) | θ value | 2.36792 (rank : 47) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 400 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9UMD9, Q02802, Q99018, Q9NQK9, Q9UC14 | Gene names | COL17A1, BP180, BPAG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XVII) chain (Bullous pemphigoid antigen 2) (180 kDa bullous pemphigoid antigen 2). | |||||
|
SRRM1_MOUSE
|
||||||
| NC score | 0.047416 (rank : 27) | θ value | 0.813845 (rank : 31) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
PDZD2_HUMAN
|
||||||
| NC score | 0.046913 (rank : 28) | θ value | 0.0252991 (rank : 14) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O15018, Q9BXD4 | Gene names | PDZD2, AIPC, KIAA0300, PDZK3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 2 (PDZ domain-containing protein 3) (Activated in prostate cancer protein). | |||||
|
CO4A6_HUMAN
|
||||||
| NC score | 0.046751 (rank : 29) | θ value | 4.03905 (rank : 67) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q14031, Q12823, Q14053, Q5JYH6, Q9NQM5, Q9NTX3, Q9UJ76, Q9UMG6, Q9Y4L4 | Gene names | COL4A6 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-6(IV) chain precursor. | |||||
|
COBA2_MOUSE
|
||||||
| NC score | 0.046696 (rank : 30) | θ value | 2.36792 (rank : 46) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 501 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q64739, Q61432, Q9Z1W0 | Gene names | Col11a2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(XI) chain precursor. | |||||
|
COJA1_HUMAN
|
||||||
| NC score | 0.046420 (rank : 31) | θ value | 5.27518 (rank : 80) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q14993, Q00559, Q05850, Q12885, Q13676, Q5JUF0, Q5T424, Q9H572, Q9NPZ2, Q9NQP2 | Gene names | COL19A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XIX) chain precursor (Collagen alpha-1(Y) chain). | |||||
|
PRG4_HUMAN
|
||||||
| NC score | 0.045241 (rank : 32) | θ value | 0.163984 (rank : 18) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
CO7A1_HUMAN
|
||||||
| NC score | 0.045231 (rank : 33) | θ value | 0.813845 (rank : 28) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 807 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q02388, Q14054, Q16507 | Gene names | COL7A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(VII) chain precursor (Long-chain collagen) (LC collagen). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.044640 (rank : 34) | θ value | 3.0926 (rank : 63) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
GP179_HUMAN
|
||||||
| NC score | 0.043034 (rank : 35) | θ value | 0.47712 (rank : 23) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 372 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6PRD1 | Gene names | GPR179, GPR158L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 179 precursor (Probable G-protein coupled receptor 158-like 1). | |||||
|
CO4A4_HUMAN
|
||||||
| NC score | 0.042563 (rank : 36) | θ value | 8.99809 (rank : 113) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 440 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P53420 | Gene names | COL4A4 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-4(IV) chain precursor. | |||||
|
K1949_MOUSE
|
||||||
| NC score | 0.041910 (rank : 37) | θ value | 2.36792 (rank : 50) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8BQ30, Q3UDS6, Q8CEY9, Q8R3D8 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1949 homolog. | |||||
|
CEP35_HUMAN
|
||||||
| NC score | 0.041866 (rank : 38) | θ value | 0.0736092 (rank : 15) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q5VT06, O75068, Q8TDK3, Q8WY20 | Gene names | CEP350, CAP350, GM133 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein 350 (Centrosome-associated protein of 350 kDa). | |||||
|
CO8A2_MOUSE
|
||||||
| NC score | 0.041751 (rank : 39) | θ value | 1.81305 (rank : 41) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P25318, Q68ED0, Q6KAQ4, Q6P1C4 | Gene names | Col8a2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-2(VIII) chain precursor (Endothelial collagen). | |||||
|
CO8A2_HUMAN
|
||||||
| NC score | 0.040823 (rank : 40) | θ value | 5.27518 (rank : 79) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 299 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P25067, Q8TEJ5 | Gene names | COL8A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(VIII) chain precursor (Endothelial collagen). | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.039947 (rank : 41) | θ value | 0.00390308 (rank : 10) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
BIN1_MOUSE
|
||||||
| NC score | 0.039515 (rank : 42) | θ value | 0.813845 (rank : 27) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O08539, Q62434 | Gene names | Bin1, Amphl, Sh3p9 | |||
|
Domain Architecture |

|
|||||
| Description | Myc box-dependent-interacting protein 1 (Bridging integrator 1) (Amphiphysin-like protein) (Amphiphysin II) (SH3-domain-containing protein 9). | |||||
|
PGCA_HUMAN
|
||||||
| NC score | 0.039470 (rank : 43) | θ value | 3.0926 (rank : 61) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P16112, Q13650, Q9UCP4, Q9UCP5, Q9UDE0 | Gene names | AGC1, CSPG1 | |||
|
Domain Architecture |

|
|||||
| Description | Aggrecan core protein precursor (Cartilage-specific proteoglycan core protein) (CSPCP) (Chondroitin sulfate proteoglycan core protein 1) [Contains: Aggrecan core protein 2]. | |||||
|
PI5PA_HUMAN
|
||||||
| NC score | 0.038952 (rank : 44) | θ value | 0.813845 (rank : 30) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 749 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q15735, Q32M61, Q6ZTH6, Q8N902, Q9UDT9 | Gene names | PIB5PA, PIPP | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4,5-bisphosphate 5-phosphatase A (EC 3.1.3.56). | |||||
|
K1802_MOUSE
|
||||||
| NC score | 0.038834 (rank : 45) | θ value | 0.47712 (rank : 24) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
CO6A2_HUMAN
|
||||||
| NC score | 0.038817 (rank : 46) | θ value | 6.88961 (rank : 91) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P12110, Q13909, Q13910, Q13911, Q14049, Q16259, Q16597, Q9UML3 | Gene names | COL6A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(VI) chain precursor. | |||||
|
EMID1_HUMAN
|
||||||
| NC score | 0.038343 (rank : 47) | θ value | 4.03905 (rank : 68) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q96A84, Q6ICG1, Q86SS7 | Gene names | EMID1, EMU1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EMI domain-containing protein 1 precursor (Protein Emu1) (Emilin and multimerin domain-containing protein 1). | |||||
|
MILK1_HUMAN
|
||||||
| NC score | 0.038112 (rank : 48) | θ value | 0.0736092 (rank : 16) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8N3F8, Q5TI16, Q7RTP5, Q8N3N8, Q9BVL9, Q9BY92, Q9UH43, Q9UH44, Q9UH45 | Gene names | MIRAB13, KIAA1668 | |||
|
Domain Architecture |

|
|||||
| Description | Molecule interacting with Rab13 (MIRab13) (MICAL-like protein 1). | |||||
|
TR13B_MOUSE
|
||||||
| NC score | 0.038095 (rank : 49) | θ value | 1.81305 (rank : 44) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9ET35, Q9DBZ3 | Gene names | Tnfrsf13b, Taci | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor necrosis factor receptor superfamily member 13B (Transmembrane activator and CAML interactor). | |||||
|
BIN1_HUMAN
|
||||||
| NC score | 0.037436 (rank : 50) | θ value | 4.03905 (rank : 65) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O00499, O00297, O00545, O43867, O60552, O60553, O60554, O60555, O75514, O75515, O75516, O75517, O75518, Q92944, Q99688 | Gene names | BIN1, AMPHL | |||
|
Domain Architecture |

|
|||||
| Description | Myc box-dependent-interacting protein 1 (Bridging integrator 1) (Amphiphysin-like protein) (Amphiphysin II) (Box-dependent myc- interacting protein 1). | |||||
|
COAA1_HUMAN
|
||||||
| NC score | 0.036598 (rank : 51) | θ value | 6.88961 (rank : 92) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 316 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q03692 | Gene names | COL10A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(X) chain precursor. | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.033895 (rank : 52) | θ value | 6.88961 (rank : 102) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
K1949_HUMAN
|
||||||
| NC score | 0.033062 (rank : 53) | θ value | 5.27518 (rank : 81) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6NYC8, Q68CK8, Q6ZUJ6, Q8NDQ4, Q8TF52, Q9BRL9 | Gene names | KIAA1949 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1949. | |||||
|
ZN598_MOUSE
|
||||||
| NC score | 0.032855 (rank : 54) | θ value | 0.813845 (rank : 32) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q80YR4, Q6KAT0, Q8R3S1 | Gene names | Znf598, Zfp598 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 598. | |||||
|
COEA1_HUMAN
|
||||||
| NC score | 0.032542 (rank : 55) | θ value | 8.99809 (rank : 114) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q05707, O00260, O00261, O00262, Q05708, Q5XJ18, Q96C67 | Gene names | COL14A1, UND | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XIV) chain precursor (Undulin). | |||||
|
BSN_MOUSE
|
||||||
| NC score | 0.030268 (rank : 56) | θ value | 8.99809 (rank : 111) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
COEA1_MOUSE
|
||||||
| NC score | 0.030224 (rank : 57) | θ value | 6.88961 (rank : 93) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q80X19, Q8C6X3, Q9WV05 | Gene names | Col14a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XIV) chain precursor. | |||||
|
JPH2_HUMAN
|
||||||
| NC score | 0.030005 (rank : 58) | θ value | 1.38821 (rank : 40) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 639 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9BR39, O95913, Q5JY74, Q9UJN4 | Gene names | JPH2, JP2 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
TAF6_HUMAN
|
||||||
| NC score | 0.029836 (rank : 59) | θ value | 1.81305 (rank : 43) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P49848 | Gene names | TAF6, TAF2E, TAFII70 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 6 (Transcription initiation factor TFIID 70 kDa subunit) (TAF(II)70) (TAFII-70) (TAFII- 80) (TAFII80). | |||||
|
ATX2L_HUMAN
|
||||||
| NC score | 0.029598 (rank : 60) | θ value | 1.06291 (rank : 33) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8WWM7, O95135, Q6NVJ8, Q6PJW6, Q8IU61, Q8IU95, Q8WWM3, Q8WWM4, Q8WWM5, Q8WWM6, Q99703 | Gene names | ATXN2L, A2D, A2LG, A2LP, A2RP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2-like protein (Ataxin-2 domain protein) (Ataxin-2-related protein). | |||||
|
ENAM_MOUSE
|
||||||
| NC score | 0.028738 (rank : 61) | θ value | 3.0926 (rank : 59) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 229 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O55196 | Gene names | Enam | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Enamelin precursor. | |||||
|
XKR7_HUMAN
|
||||||
| NC score | 0.027949 (rank : 62) | θ value | 1.06291 (rank : 38) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q5GH72, Q9NUG5 | Gene names | XKR7, C20orf159, XRG7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | XK-related protein 7. | |||||
|
ZDHC5_MOUSE
|
||||||
| NC score | 0.027491 (rank : 63) | θ value | 0.279714 (rank : 21) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8VDZ4, Q69ZB5, Q8R2X7 | Gene names | Zdhhc5, Kiaa1748 | |||
|
Domain Architecture |

|
|||||
| Description | Probable palmitoyltransferase ZDHHC5 (EC 2.3.1.-) (Zinc finger DHHC domain-containing protein 5) (DHHC-5). | |||||
|
PHLB1_MOUSE
|
||||||
| NC score | 0.026885 (rank : 64) | θ value | 1.06291 (rank : 36) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 678 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q6PDH0, Q80TV2 | Gene names | Phldb1, Kiaa0638, Ll5a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family B member 1 (Protein LL5-alpha). | |||||
|
K0310_HUMAN
|
||||||
| NC score | 0.026441 (rank : 65) | θ value | 6.88961 (rank : 101) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O15027, Q5SXP0, Q5SXP1, Q8N347, Q96HP1 | Gene names | KIAA0310 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0310. | |||||
|
RGS3_HUMAN
|
||||||
| NC score | 0.025982 (rank : 66) | θ value | 5.27518 (rank : 82) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
NUP62_HUMAN
|
||||||
| NC score | 0.025764 (rank : 67) | θ value | 3.0926 (rank : 60) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 195 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P37198, Q503A4, Q96C43, Q9NSL1 | Gene names | NUP62 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore glycoprotein p62 (62 kDa nucleoporin). | |||||
|
FCN1_HUMAN
|
||||||
| NC score | 0.025663 (rank : 68) | θ value | 0.813845 (rank : 29) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O00602, Q92596 | Gene names | FCN1 | |||
|
Domain Architecture |

|
|||||
| Description | Ficolin-1 precursor (Collagen/fibrinogen domain-containing protein 1) (Ficolin-A) (Ficolin A) (M-Ficolin). | |||||
|
CIC_HUMAN
|
||||||
| NC score | 0.025638 (rank : 69) | θ value | 3.0926 (rank : 54) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 992 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q96RK0, Q7LGI1, Q9UEG5, Q9Y6T1 | Gene names | CIC, KIAA0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
XKR7_MOUSE
|
||||||
| NC score | 0.024380 (rank : 70) | θ value | 5.27518 (rank : 85) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q5GH64 | Gene names | Xkr7, Xrg7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | XK-related protein 7. | |||||
|
TM115_MOUSE
|
||||||
| NC score | 0.024140 (rank : 71) | θ value | 4.03905 (rank : 74) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9WUH1, Q91VT6 | Gene names | Tmem115, Pl6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 115 (Protein PL6 homolog). | |||||
|
DPOG1_MOUSE
|
||||||
| NC score | 0.023500 (rank : 72) | θ value | 2.36792 (rank : 48) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P54099, Q9JI28 | Gene names | Polg, Polg1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase subunit gamma 1 (EC 2.7.7.7) (Mitochondrial DNA polymerase catalytic subunit) (PolG-alpha). | |||||
|
DAG1_HUMAN
|
||||||
| NC score | 0.023433 (rank : 73) | θ value | 3.0926 (rank : 58) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q14118 | Gene names | DAG1 | |||
|
Domain Architecture |

|
|||||
| Description | Dystroglycan precursor (Dystrophin-associated glycoprotein 1) [Contains: Alpha-dystroglycan (Alpha-DG); Beta-dystroglycan (Beta- DG)]. | |||||
|
IF4G1_MOUSE
|
||||||
| NC score | 0.023426 (rank : 74) | θ value | 2.36792 (rank : 49) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 534 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q6NZJ6, Q6NZN8, Q8BW99 | Gene names | Eif4g1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1). | |||||
|
SFR14_HUMAN
|
||||||
| NC score | 0.022698 (rank : 75) | θ value | 2.36792 (rank : 53) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IX01, O15071, O60369, Q5JPH7, Q8WUF7 | Gene names | SFRS14, KIAA0365 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative splicing factor, arginine/serine-rich 14 (Arginine/serine- rich-splicing factor 14). | |||||
|
NCOA2_MOUSE
|
||||||
| NC score | 0.021783 (rank : 76) | θ value | 2.36792 (rank : 51) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q61026, O09001, P97759 | Gene names | Ncoa2, Grip1, Tif2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2) (Glucocorticoid receptor-interacting protein 1) (GRIP-1). | |||||
|
ZDHC5_HUMAN
|
||||||
| NC score | 0.020876 (rank : 77) | θ value | 4.03905 (rank : 75) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9C0B5, Q6ZMF0, Q8TAK8, Q9H923, Q9UFI7 | Gene names | ZDHHC5, KIAA1748, ZNF375 | |||
|
Domain Architecture |

|
|||||
| Description | Probable palmitoyltransferase ZDHHC5 (EC 2.3.1.-) (Zinc finger DHHC domain-containing protein 5) (DHHC-5) (Zinc finger protein 375). | |||||
|
GA2L2_MOUSE
|
||||||
| NC score | 0.020277 (rank : 78) | θ value | 6.88961 (rank : 98) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q5SSG4 | Gene names | Gas2l2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GAS2-like protein 2 (Growth arrest-specific 2-like 2). | |||||
|
CAMLG_MOUSE
|
||||||
| NC score | 0.019776 (rank : 79) | θ value | 5.27518 (rank : 78) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P49070, Q99JU5 | Gene names | Camlg, Caml | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium signal-modulating cyclophilin ligand (CAML). | |||||
|
PGCB_MOUSE
|
||||||
| NC score | 0.019174 (rank : 80) | θ value | 0.47712 (rank : 26) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 459 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q61361 | Gene names | Bcan | |||
|
Domain Architecture |

|
|||||
| Description | Brevican core protein precursor. | |||||
|
PLIN_MOUSE
|
||||||
| NC score | 0.018836 (rank : 81) | θ value | 4.03905 (rank : 73) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CGN5 | Gene names | Plin, Peri | |||
|
Domain Architecture |

|
|||||
| Description | Perilipin (PERI) (Lipid droplet-associated protein). | |||||
|
SSFA2_MOUSE
|
||||||
| NC score | 0.018234 (rank : 82) | θ value | 6.88961 (rank : 107) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q922B9, Q8VEF7 | Gene names | Ssfa2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-specific antigen 2. | |||||
|
IKBL_HUMAN
|
||||||
| NC score | 0.018048 (rank : 83) | θ value | 4.03905 (rank : 70) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UBC1, Q14625, Q9UBX4 | Gene names | NFKBIL1, IKBL | |||
|
Domain Architecture |

|
|||||
| Description | NF-kappa-B inhibitor-like protein 1 (Inhibitor of kappa B-like) (I- kappa-B-like) (IkappaBL) (Nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor-like 1). | |||||
|
IF39_MOUSE
|
||||||
| NC score | 0.017745 (rank : 84) | θ value | 6.88961 (rank : 100) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8JZQ9, Q922K2 | Gene names | Eif3s9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 9 (eIF-3 eta) (eIF3 p116). | |||||
|
ASPX_HUMAN
|
||||||
| NC score | 0.017321 (rank : 85) | θ value | 6.88961 (rank : 89) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P26436 | Gene names | ACRV1 | |||
|
Domain Architecture |

|
|||||
| Description | Acrosomal protein SP-10 precursor (Acrosomal vesicle protein 1). | |||||
|
RIOK2_MOUSE
|
||||||
| NC score | 0.017225 (rank : 86) | θ value | 5.27518 (rank : 83) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9CQS5, Q91XF3 | Gene names | Riok2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase RIO2 (EC 2.7.11.1) (RIO kinase 2). | |||||
|
FMN2_HUMAN
|
||||||
| NC score | 0.017081 (rank : 87) | θ value | 6.88961 (rank : 97) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 484 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9NZ56, Q59GF6, Q5VU37, Q9NZ55 | Gene names | FMN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Formin-2. | |||||
|
EFHB_MOUSE
|
||||||
| NC score | 0.016620 (rank : 88) | θ value | 6.88961 (rank : 95) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CDU5 | Gene names | Efhb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EF-hand domain-containing family member B. | |||||
|
USBP1_HUMAN
|
||||||
| NC score | 0.016614 (rank : 89) | θ value | 5.27518 (rank : 84) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8N6Y0, Q8NBX7, Q96KH3, Q9BYI8 | Gene names | USHBP1, AIEBP, MCC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USH1C-binding protein 1 (Usher syndrome type-1C protein-binding protein 1) (MCC-2) (AIE-75 binding protein). | |||||
|
FXL22_HUMAN
|
||||||
| NC score | 0.016354 (rank : 90) | θ value | 8.99809 (rank : 117) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6P050 | Gene names | FBXL22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 22 (F-box and leucine-rich repeat protein 22). | |||||
|
PITM1_HUMAN
|
||||||
| NC score | 0.015968 (rank : 91) | θ value | 2.36792 (rank : 52) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O00562, Q6T7X3, Q8TBN3, Q9BZ73 | Gene names | PITPNM1, DRES9, NIR2, PITPNM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated phosphatidylinositol transfer protein 1 (Phosphatidylinositol transfer protein, membrane-associated 1) (Pyk2 N-terminal domain-interacting receptor 2) (NIR-2) (Drosophila retinal degeneration B homolog). | |||||
|
ETV4_MOUSE
|
||||||
| NC score | 0.015807 (rank : 92) | θ value | 1.06291 (rank : 35) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P28322 | Gene names | Etv4, Pea-3, Pea3 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 4 (Polyomavirus enhancer activator 3) (Protein PEA3). | |||||
|
TCGAP_HUMAN
|
||||||
| NC score | 0.015605 (rank : 93) | θ value | 8.99809 (rank : 122) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 872 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O14559, O14552, O14560, Q6ZSP6, Q96CP3, Q9NT23 | Gene names | SNX26, TCGAP | |||
|
Domain Architecture |

|
|||||
| Description | TC10/CDC42 GTPase-activating protein (Sorting nexin-26). | |||||
|
RAVR2_MOUSE
|
||||||
| NC score | 0.014834 (rank : 94) | θ value | 3.0926 (rank : 62) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7TPD6 | Gene names | Raver2, Kiaa1579 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ribonucleoprotein PTB-binding 2 (Protein raver-2). | |||||
|
NCOR1_MOUSE
|
||||||
| NC score | 0.014813 (rank : 95) | θ value | 6.88961 (rank : 103) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 521 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q60974, Q60812 | Gene names | Ncor1, Rxrip13 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR) (Retinoid X receptor- interacting protein 13) (RIP13). | |||||
|
AP4B1_HUMAN
|
||||||
| NC score | 0.013864 (rank : 96) | θ value | 6.88961 (rank : 87) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y6B7 | Gene names | AP4B1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-4 complex subunit beta-1 (Adapter-related protein complex 4 beta 1 subunit) (Beta subunit of AP-4) (AP-4 adapter complex beta subunit). | |||||
|
AP4B1_MOUSE
|
||||||
| NC score | 0.013534 (rank : 97) | θ value | 6.88961 (rank : 88) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9WV76 | Gene names | Ap4b1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-4 complex subunit beta-1 (Adapter-related protein complex 4 beta-1 subunit) (Beta subunit of AP-4) (AP-4 adapter complex beta subunit). | |||||
|
PER1_MOUSE
|
||||||
| NC score | 0.013251 (rank : 98) | θ value | 8.99809 (rank : 118) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O35973 | Gene names | Per1, Per, Rigui | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (mPER) (M-Rigui). | |||||
|
S45A1_MOUSE
|
||||||
| NC score | 0.012388 (rank : 99) | θ value | 6.88961 (rank : 106) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BIV7, Q3TZ98 | Gene names | Slc45a1, Dnb5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proton-associated sugar transporter A (PAST-A) (Solute carrier family 45 member 1) (Deleted in neuroblastoma 5 protein homolog) (DNb-5 homolog). | |||||
|
EXO1_HUMAN
|
||||||
| NC score | 0.012178 (rank : 100) | θ value | 8.99809 (rank : 115) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UQ84, O60545, O75214, O75466, Q5T396, Q96IJ1, Q9UG38, Q9UNW0 | Gene names | EXO1, EXOI, HEX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Exonuclease 1 (EC 3.1.-.-) (hExo1) (Exonuclease I) (hExoI). | |||||
|
PEF1_MOUSE
|
||||||
| NC score | 0.010322 (rank : 101) | θ value | 6.88961 (rank : 104) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8BFY6, Q8VCT5, Q9CYW8, Q9D934 | Gene names | Pef1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peflin (PEF protein with a long N-terminal hydrophobic domain) (Penta- EF hand domain-containing protein 1). | |||||
|
FBX30_MOUSE
|
||||||
| NC score | 0.009920 (rank : 102) | θ value | 8.99809 (rank : 116) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BJL1, Q8CI21, Q9D9X5 | Gene names | Fbxo30 | |||
|
Domain Architecture |

|
|||||
| Description | F-box only protein 30. | |||||
|
SDK1_HUMAN
|
||||||
| NC score | 0.009429 (rank : 103) | θ value | 1.06291 (rank : 37) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 511 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q7Z5N4, Q8TEN9, Q8TEP5, Q96N44 | Gene names | SDK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein sidekick-1 precursor. | |||||
|
INVS_MOUSE
|
||||||
| NC score | 0.009245 (rank : 104) | θ value | 4.03905 (rank : 71) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 609 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O89019, O88849 | Gene names | Invs, Inv, Nphp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inversin (Inversion of embryo turning protein) (Nephrocystin-2). | |||||
|
DAB2P_HUMAN
|
||||||
| NC score | 0.008966 (rank : 105) | θ value | 6.88961 (rank : 94) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5VWQ8, Q8TDL2, Q96SE1, Q9C0C0 | Gene names | DAB2IP, AF9Q34, AIP1, KIAA1743 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Disabled homolog 2-interacting protein (DAB2-interacting protein) (DAB2 interaction protein) (ASK-interacting protein 1). | |||||
|
ULK1_HUMAN
|
||||||
| NC score | 0.008627 (rank : 106) | θ value | 0.163984 (rank : 19) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1121 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O75385 | Gene names | ULK1 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase ULK1 (EC 2.7.11.1) (Unc-51-like kinase 1). | |||||
|
PODO_HUMAN
|
||||||
| NC score | 0.008564 (rank : 107) | θ value | 8.99809 (rank : 119) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NP85, Q8N6Q5 | Gene names | NPHS2 | |||
|
Domain Architecture |

|
|||||
| Description | Podocin. | |||||
|
RED1_MOUSE
|
||||||
| NC score | 0.007947 (rank : 108) | θ value | 8.99809 (rank : 120) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q91ZS8, Q8K3X1, Q91ZS6, Q91ZS7, Q91ZS9, Q99MU8 | Gene names | Adarb1, Adar2, Red1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double-stranded RNA-specific editase 1 (EC 3.5.-.-) (dsRNA adenosine deaminase) (RNA-editing deaminase 1) (RNA-editing enzyme 1). | |||||
|
BSPRY_MOUSE
|
||||||
| NC score | 0.007568 (rank : 109) | θ value | 6.88961 (rank : 90) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80YW5, Q3TU74, Q8BZF0, Q99KV7, Q9ER70 | Gene names | Bspry | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B box and SPRY domain-containing protein. | |||||
|
UBP29_MOUSE
|
||||||
| NC score | 0.007173 (rank : 110) | θ value | 8.99809 (rank : 124) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9ES63 | Gene names | Usp29 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 29 (EC 3.1.2.15) (Ubiquitin thioesterase 29) (Ubiquitin-specific-processing protease 29) (Deubiquitinating enzyme 29). | |||||
|
ABL1_MOUSE
|
||||||
| NC score | 0.006946 (rank : 111) | θ value | 0.21417 (rank : 20) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1178 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P00520, P97896, Q61252, Q61253, Q61254, Q61255, Q61256, Q61257, Q61258, Q61259, Q61260, Q61261, Q6PCM5, Q8C1X4 | Gene names | Abl1, Abl | |||
|
Domain Architecture |

|
|||||
| Description | Proto-oncogene tyrosine-protein kinase ABL1 (EC 2.7.10.2) (p150) (c- ABL) (Abelson murine leukemia viral oncogene homolog 1). | |||||
|
ARL4C_HUMAN
|
||||||
| NC score | 0.006864 (rank : 112) | θ value | 5.27518 (rank : 76) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P56559, Q9BVN1, Q9UQ34 | Gene names | ARL4C, ARL7 | |||
|
Domain Architecture |

|
|||||
| Description | ADP-ribosylation factor-like protein 4C (ADP-ribosylation factor-like protein 7) (ADP-ribosylation factor-like protein LAK). | |||||
|
ARL4C_MOUSE
|
||||||
| NC score | 0.006864 (rank : 113) | θ value | 5.27518 (rank : 77) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P61208 | Gene names | Arl4c, Arl7 | |||
|
Domain Architecture |

|
|||||
| Description | ADP-ribosylation factor-like protein 4C (ADP-ribosylation factor-like 7). | |||||
|
TSH3_HUMAN
|
||||||
| NC score | 0.006761 (rank : 114) | θ value | 8.99809 (rank : 123) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 564 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q63HK5, Q9H0G6, Q9P254 | Gene names | TSHZ3, KIAA1474, TSH3, ZNF537 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 3 (Zinc finger protein 537). | |||||
|
ITCH_HUMAN
|
||||||
| NC score | 0.006605 (rank : 115) | θ value | 4.03905 (rank : 72) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96J02, O43584, Q5TEL0, Q96F66, Q9BY75, Q9H451, Q9H4U5 | Gene names | ITCH | |||
|
Domain Architecture |

|
|||||
| Description | Itchy homolog E3 ubiquitin protein ligase (EC 6.3.2.-) (Itch) (Atrophin-1-interacting protein 4) (AIP4) (NFE2-associated polypeptide 1) (NAPP1). | |||||
|
SPEG_MOUSE
|
||||||
| NC score | 0.006426 (rank : 116) | θ value | 8.99809 (rank : 121) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
AZI1_HUMAN
|
||||||
| NC score | 0.006265 (rank : 117) | θ value | 8.99809 (rank : 110) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1000 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UPN4, Q96F50 | Gene names | AZI1, CEP131, KIAA1118 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 5-azacytidine-induced protein 1 (Pre-acrosome localization protein 1) (Centrosomal protein of 131 kDa) (Cep131 protein). | |||||
|
PRGR_HUMAN
|
||||||
| NC score | 0.005846 (rank : 118) | θ value | 6.88961 (rank : 105) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P06401, Q9UPF7 | Gene names | PGR, NR3C3 | |||
|
Domain Architecture |

|
|||||
| Description | Progesterone receptor (PR). | |||||
|
ERR1_HUMAN
|
||||||
| NC score | 0.005642 (rank : 119) | θ value | 6.88961 (rank : 96) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P11474, Q14514 | Gene names | ESRRA, ERR1, ESRL1, NR3B1 | |||
|
Domain Architecture |

|
|||||
| Description | Steroid hormone receptor ERR1 (Estrogen-related receptor, alpha) (ERR- alpha) (Estrogen receptor-like 1). | |||||
|
GDF7_MOUSE
|
||||||
| NC score | 0.005063 (rank : 120) | θ value | 6.88961 (rank : 99) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P43029, Q7TNX4, Q99MY1 | Gene names | Gdf7, Gdf-7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Growth/differentiation factor 7 precursor (GDF-7). | |||||
|
CCD13_HUMAN
|
||||||
| NC score | 0.003930 (rank : 121) | θ value | 8.99809 (rank : 112) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 721 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8IYE1 | Gene names | CCDC13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 13. | |||||
|
ZN592_HUMAN
|
||||||
| NC score | 0.003464 (rank : 122) | θ value | 6.88961 (rank : 109) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 762 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q92610 | Gene names | ZNF592, KIAA0211 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 592. | |||||
|
GSK3A_HUMAN
|
||||||
| NC score | 0.002457 (rank : 123) | θ value | 4.03905 (rank : 69) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 835 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P49840, O14959 | Gene names | GSK3A | |||
|
Domain Architecture |

|
|||||
| Description | Glycogen synthase kinase-3 alpha (EC 2.7.11.26) (GSK-3 alpha). | |||||
|
ST32C_HUMAN
|
||||||
| NC score | 0.001176 (rank : 124) | θ value | 6.88961 (rank : 108) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 832 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q86UX6, Q5T0Q5, Q86UE1 | Gene names | STK32C | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase 32C (EC 2.7.11.1) (PKE) (YANK3). | |||||
|
ZN444_HUMAN
|
||||||
| NC score | 0.000494 (rank : 125) | θ value | 3.0926 (rank : 64) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 701 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8N0Y2, Q8TEQ9, Q8WU35, Q9NUU1 | Gene names | ZNF444, EZF2, ZSCAN17 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 444 (Endothelial zinc finger protein 2) (EZF-2) (Zinc finger and SCAN domain-containing protein 17). | |||||
|
ZN358_HUMAN
|
||||||
| NC score | -0.000802 (rank : 126) | θ value | 5.27518 (rank : 86) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 840 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9NW07, Q9BTM7 | Gene names | ZNF358 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 358. | |||||