Please be patient as the page loads
|
GMEB2_MOUSE
|
||||||
| SwissProt Accessions | P58929, Q6PCY0 | Gene names | Gmeb2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glucocorticoid modulatory element-binding protein 2 (GMEB-2). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
GMEB2_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.987619 (rank : 2) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9UKD1, Q5TDS0, Q9H431, Q9H4X7, Q9H4X8, Q9UF78, Q9ULF1 | Gene names | GMEB2, KIAA1269 | |||
|
Domain Architecture |
|
|||||
| Description | Glucocorticoid modulatory element-binding protein 2 (GMEB-2) (Parvovirus initiation factor p79) (PIF p79) (DNA-binding protein p79PIF). | |||||
|
GMEB2_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | P58929, Q6PCY0 | Gene names | Gmeb2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glucocorticoid modulatory element-binding protein 2 (GMEB-2). | |||||
|
GMEB1_MOUSE
|
||||||
| θ value | 1.22395e-89 (rank : 3) | NC score | 0.920203 (rank : 3) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9JL60 | Gene names | Gmeb1 | |||
|
Domain Architecture |
|
|||||
| Description | Glucocorticoid modulatory element-binding protein 1 (GMEB-1). | |||||
|
GMEB1_HUMAN
|
||||||
| θ value | 4.65103e-89 (rank : 4) | NC score | 0.919711 (rank : 4) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Y692, Q9NWH1, Q9UKD0 | Gene names | GMEB1 | |||
|
Domain Architecture |
|
|||||
| Description | Glucocorticoid modulatory element-binding protein 1 (GMEB-1) (Parvovirus initiation factor p96) (PIF p96) (DNA-binding protein p96PIF). | |||||
|
IPR1_MOUSE
|
||||||
| θ value | 1.29631e-06 (rank : 5) | NC score | 0.384662 (rank : 5) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8BVK9, Q3UCV7, Q80V00 | Gene names | Ipr1, Ifi75 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intracellular pathogen resistance protein 1. | |||||
|
DEAF1_HUMAN
|
||||||
| θ value | 2.21117e-06 (rank : 6) | NC score | 0.342586 (rank : 6) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O75398, O15152, O75399, O75510, O75511, O75512, O75513, Q9UET1 | Gene names | DEAF1, SPN, ZMYND5 | |||
|
Domain Architecture |

|
|||||
| Description | Deformed epidermal autoregulatory factor 1 homolog (Nuclear DEAF-1- related transcriptional regulator) (NUDR) (Suppressin) (Zinc finger MYND domain-containing protein 5). | |||||
|
DEAF1_MOUSE
|
||||||
| θ value | 2.21117e-06 (rank : 7) | NC score | 0.339768 (rank : 7) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Z1T5 | Gene names | Deaf1 | |||
|
Domain Architecture |

|
|||||
| Description | Deformed epidermal autoregulatory factor 1 homolog (Nuclear DEAF-1- related transcriptional regulator) (NUDR). | |||||
|
LY10_HUMAN
|
||||||
| θ value | 0.00020696 (rank : 8) | NC score | 0.252916 (rank : 9) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q13342, Q13341, Q92881, Q96TG3 | Gene names | SP140, LYSP100 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear body protein SP140 (Nuclear autoantigen Sp-140) (Speckled 140 kDa) (LYSp100 protein) (Lymphoid-restricted homolog of Sp100). | |||||
|
SP110_HUMAN
|
||||||
| θ value | 0.000270298 (rank : 9) | NC score | 0.273446 (rank : 8) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9HB58, Q14976, Q14977, Q8WUZ6, Q9HCT8 | Gene names | SP110 | |||
|
Domain Architecture |

|
|||||
| Description | Sp110 nuclear body protein (Speckled 110 kDa) (Transcriptional coactivator Sp110) (Interferon-induced protein 41/75). | |||||
|
SP100_HUMAN
|
||||||
| θ value | 0.00102713 (rank : 10) | NC score | 0.201246 (rank : 10) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P23497, O75450, Q13343, Q96F70, Q96T24, Q96T95, Q9NP33, Q9UE32 | Gene names | SP100 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear autoantigen Sp-100 (Speckled 100 kDa) (Nuclear dot-associated Sp100 protein) (Lysp100b). | |||||
|
ATS12_HUMAN
|
||||||
| θ value | 0.125558 (rank : 11) | NC score | 0.024838 (rank : 35) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P58397, Q6UWL3 | Gene names | ADAMTS12 | |||
|
Domain Architecture |

|
|||||
| Description | ADAMTS-12 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase with thrombospondin motifs 12) (ADAM-TS 12) (ADAM-TS12). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 0.163984 (rank : 12) | NC score | 0.034834 (rank : 22) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
CP250_HUMAN
|
||||||
| θ value | 0.279714 (rank : 13) | NC score | 0.024912 (rank : 34) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1805 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9BV73, O14812, O60588, Q9H450 | Gene names | CEP250, CEP2, CNAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1). | |||||
|
NELFA_MOUSE
|
||||||
| θ value | 0.279714 (rank : 14) | NC score | 0.065903 (rank : 13) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 195 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BG30, Q8BVE4, Q8VEI7, Q9CSJ9, Q9Z1V9 | Gene names | Whsc2, Nelfa, Whsc2h | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Negative elongation factor A (NELF-A) (Wolf-Hirschhorn syndrome candidate 2 homolog) (mWHSC2). | |||||
|
RHG25_HUMAN
|
||||||
| θ value | 0.279714 (rank : 15) | NC score | 0.035462 (rank : 20) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 550 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P42331, Q8IXQ2 | Gene names | ARHGAP25, KIAA0053 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 25. | |||||
|
KIF5A_HUMAN
|
||||||
| θ value | 0.365318 (rank : 16) | NC score | 0.028104 (rank : 23) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 843 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q12840, Q4LE26 | Gene names | KIF5A, NKHC1 | |||
|
Domain Architecture |
|
|||||
| Description | Kinesin heavy chain isoform 5A (Neuronal kinesin heavy chain) (NKHC) (Kinesin heavy chain neuron-specific 1). | |||||
|
NELFA_HUMAN
|
||||||
| θ value | 0.62314 (rank : 17) | NC score | 0.062344 (rank : 15) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H3P2, O95392 | Gene names | WHSC2, NELFA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Negative elongation factor A (NELF-A) (Wolf-Hirschhorn syndrome candidate 2 protein). | |||||
|
FYCO1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 18) | NC score | 0.035004 (rank : 21) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1149 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9BQS8, Q3MJE6, Q86T41, Q86TB1, Q8TEF9, Q96IV5, Q9H8P9 | Gene names | FYCO1, ZFYVE7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE and coiled-coil domain-containing protein 1 (Zinc finger FYVE domain-containing protein 7). | |||||
|
KIF5A_MOUSE
|
||||||
| θ value | 0.813845 (rank : 19) | NC score | 0.026178 (rank : 28) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 859 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P33175, Q5DTP1, Q6PDY7, Q9Z2F9 | Gene names | Kif5a, Kiaa4086, Kif5, Nkhc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin heavy chain isoform 5A (Neuronal kinesin heavy chain) (NKHC) (Kinesin heavy chain neuron-specific 1). | |||||
|
EP400_HUMAN
|
||||||
| θ value | 1.06291 (rank : 20) | NC score | 0.018333 (rank : 51) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96L91, O15411, Q6P2F5, Q8N8Q7, Q8NE05, Q96JK7, Q9P230 | Gene names | EP400, CAGH32, KIAA1498, KIAA1818, TNRC12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E1A-binding protein p400 (EC 3.6.1.-) (p400 kDa SWI2/SNF2-related protein) (Domino homolog) (hDomino) (CAG repeat protein 32) (Trinucleotide repeat-containing gene 12 protein). | |||||
|
FA29A_HUMAN
|
||||||
| θ value | 1.06291 (rank : 21) | NC score | 0.063690 (rank : 14) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7Z4H7, Q6IQ10, Q6NZX5, Q8IZQ4, Q96FN0, Q9H950, Q9H998, Q9HCJ8, Q9NXT8 | Gene names | FAM29A, KIAA1574 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM29A. | |||||
|
PRDM2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 22) | NC score | 0.011703 (rank : 58) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q13029, Q13149, Q14550, Q5VUL9 | Gene names | PRDM2, RIZ | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 2 (PR domain-containing protein 2) (Retinoblastoma protein-interacting zinc-finger protein) (Zinc finger protein RIZ) (MTE-binding protein) (MTB-ZF) (GATA-3-binding protein G3B). | |||||
|
RB6I2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 23) | NC score | 0.027899 (rank : 25) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1584 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8IUD2, Q6NVK2, Q8IUD3, Q8IUD4, Q8IUD5, Q8NAS1, Q9NXN5, Q9UIK7, Q9UPS1 | Gene names | ERC1, ELKS, KIAA1081, RAB6IP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ELKS/RAB6-interacting/CAST family member 1 (RAB6-interacting protein 2) (ERC protein 1). | |||||
|
TM108_MOUSE
|
||||||
| θ value | 1.06291 (rank : 24) | NC score | 0.038446 (rank : 18) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 336 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BHE4, Q80WR9 | Gene names | Tmem108 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 108 precursor. | |||||
|
MYH6_HUMAN
|
||||||
| θ value | 1.38821 (rank : 25) | NC score | 0.023304 (rank : 38) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1426 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P13533, Q13943, Q14906, Q14907 | Gene names | MYH6, MYHCA | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
MYH7_HUMAN
|
||||||
| θ value | 1.38821 (rank : 26) | NC score | 0.023079 (rank : 41) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P12883, Q14836, Q14837, Q14904, Q16579, Q92679, Q9H1D5, Q9UMM8 | Gene names | MYH7, MYHCB | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle beta isoform (MyHC-beta). | |||||
|
ZHX1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 27) | NC score | 0.018612 (rank : 50) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 334 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P70121, Q8BQ68, Q8C6T4, Q8CJG3 | Gene names | Zhx1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc fingers and homeoboxes protein 1. | |||||
|
BTNL1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 28) | NC score | 0.015431 (rank : 54) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7TST0, O70356 | Gene names | Btnl1, Btnl3, Gm316, Ng10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Butyrophilin-like protein 1 precursor. | |||||
|
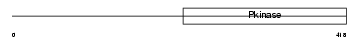
IRAK1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 29) | NC score | 0.005733 (rank : 63) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 820 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P51617, Q7Z5V4, Q96RL2 | Gene names | IRAK1, IRAK | |||
|
Domain Architecture |
|
|||||
| Description | Interleukin-1 receptor-associated kinase 1 (EC 2.7.11.1) (IRAK-1). | |||||
|
NICA_HUMAN
|
||||||
| θ value | 1.81305 (rank : 30) | NC score | 0.050318 (rank : 16) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q92542, Q86VV5 | Gene names | NCSTN, KIAA0253 | |||
|
Domain Architecture |

|
|||||
| Description | Nicastrin precursor. | |||||
|
NICA_MOUSE
|
||||||
| θ value | 1.81305 (rank : 31) | NC score | 0.050156 (rank : 17) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P57716, Q8VE20 | Gene names | Ncstn | |||
|
Domain Architecture |

|
|||||
| Description | Nicastrin precursor. | |||||
|
AHTF1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 32) | NC score | 0.022737 (rank : 44) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8WYP5, Q7Z4E3, Q8IZA4, Q96EH9, Q9Y4Q6 | Gene names | AHCTF1, ELYS, TMBS62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-hook-containing transcription factor 1 (Embryonic large molecule derived from yolk sac). | |||||
|
ALEX_HUMAN
|
||||||
| θ value | 2.36792 (rank : 33) | NC score | 0.028010 (rank : 24) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P84996 | Gene names | GNAS, GNAS1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ALEX (Alternative gene product encoded by XL-exon). | |||||
|
CING_MOUSE
|
||||||
| θ value | 2.36792 (rank : 34) | NC score | 0.026806 (rank : 27) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1432 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P59242 | Gene names | Cgn | |||
|
Domain Architecture |

|
|||||
| Description | Cingulin. | |||||
|
FLIP1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 35) | NC score | 0.022924 (rank : 42) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q7Z7B0, Q5VUL6, Q86TC3, Q8N8B9, Q96SK6, Q9NVI8, Q9ULE5 | Gene names | FILIP1, KIAA1275 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-A-interacting protein 1 (FILIP). | |||||
|
WDFY3_HUMAN
|
||||||
| θ value | 2.36792 (rank : 36) | NC score | 0.018115 (rank : 52) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 348 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8IZQ1, Q4W5K5, Q6P0Q5, Q8N1T2, Q96BS7, Q96N85, Q9Y2J7 | Gene names | WDFY3, KIAA0993 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat and FYVE domain-containing protein 3 (Autophagy-linked FYVE protein) (Alfy). | |||||
|
PME17_HUMAN
|
||||||
| θ value | 3.0926 (rank : 37) | NC score | 0.023175 (rank : 39) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P40967, Q12763, Q14448, Q14817, Q16565 | Gene names | SILV, D12S53E, PMEL17 | |||
|
Domain Architecture |

|
|||||
| Description | Melanocyte protein Pmel 17 precursor (Melanocyte lineage-specific antigen GP100) (Melanoma-associated ME20 antigen) (ME20M/ME20S) (ME20- M/ME20-S) (95 kDa melanocyte-specific secreted glycoprotein). | |||||
|
RB6I2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 38) | NC score | 0.024987 (rank : 33) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1433 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q99MI1, Q80TK7, Q8BPL1, Q8C7Y1, Q99MI2 | Gene names | Erc1, Cast2, Kiaa1081, Rab6ip2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ELKS/RAB6-interacting/CAST family member 1 (RAB6-interacting protein 2) (ERC protein 1) (ERC1) (CAZ-associated structural protein 2) (CAST2). | |||||
|
KNTC2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 39) | NC score | 0.025048 (rank : 32) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 692 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9D0F1, Q3TQT6, Q3UWM5, Q99P70 | Gene names | Kntc2, Hec1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinetochore protein Hec1 (Kinetochore-associated protein 2). | |||||
|
MYH6_MOUSE
|
||||||
| θ value | 4.03905 (rank : 40) | NC score | 0.021452 (rank : 48) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1501 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q02566, Q64258, Q64738 | Gene names | Myh6, Myhca | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
RBL1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 41) | NC score | 0.025250 (rank : 30) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P28749, Q4VXA0, Q8N5K6, Q9H1L5, Q9H1M1 | Gene names | RBL1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinoblastoma-like protein 1 (107 kDa retinoblastoma-associated protein) (PRB1) (P107). | |||||
|
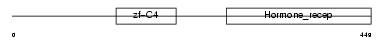
RXRA_HUMAN
|
||||||
| θ value | 4.03905 (rank : 42) | NC score | 0.009486 (rank : 59) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P19793, Q2V504 | Gene names | RXRA, NR2B1 | |||
|
Domain Architecture |
|
|||||
| Description | Retinoic acid receptor RXR-alpha (Retinoid X receptor alpha). | |||||
|
BRD8_MOUSE
|
||||||
| θ value | 5.27518 (rank : 43) | NC score | 0.036406 (rank : 19) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 422 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8R3B7, Q8C049, Q8R583, Q8VDP0 | Gene names | Brd8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 8. | |||||
|
DCC_MOUSE
|
||||||
| θ value | 5.27518 (rank : 44) | NC score | 0.005634 (rank : 64) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P70211 | Gene names | Dcc | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC). | |||||
|
FLIP1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 45) | NC score | 0.022790 (rank : 43) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1176 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9CS72 | Gene names | Filip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-A-interacting protein 1 (FILIP). | |||||
|
MYH1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 46) | NC score | 0.023108 (rank : 40) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1478 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P12882, Q9Y622 | Gene names | MYH1 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-1 (Myosin heavy chain, skeletal muscle, adult 1) (Myosin heavy chain IIx/d) (MyHC-IIx/d). | |||||
|
NIN_HUMAN
|
||||||
| θ value | 5.27518 (rank : 47) | NC score | 0.023520 (rank : 37) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1416 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8N4C6, Q6P0P6, Q9BWU6, Q9C012, Q9C013, Q9C014, Q9H5I6, Q9HAT7, Q9HBY5, Q9HCK7, Q9UH61 | Gene names | NIN, KIAA1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein (hNinein) (Glycogen synthase kinase 3 beta-interacting protein) (GSK3B-interacting protein). | |||||
|
RBL2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 48) | NC score | 0.025198 (rank : 31) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q08999, Q15073, Q16084, Q92812 | Gene names | RBL2, RB2 | |||
|
Domain Architecture |

|
|||||
| Description | Retinoblastoma-like protein 2 (130 kDa retinoblastoma-associated protein) (PRB2) (P130) (RBR-2). | |||||
|
RBL2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 49) | NC score | 0.025788 (rank : 29) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q64700 | Gene names | Rbl2 | |||
|
Domain Architecture |

|
|||||
| Description | Retinoblastoma-like protein 2 (130 kDa retinoblastoma-associated protein) (PRB2) (P130) (RBR-2). | |||||
|
CING_HUMAN
|
||||||
| θ value | 6.88961 (rank : 50) | NC score | 0.024113 (rank : 36) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1275 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9P2M7, Q5T386, Q9NR25 | Gene names | CGN, KIAA1319 | |||
|
Domain Architecture |

|
|||||
| Description | Cingulin. | |||||
|
DEND_HUMAN
|
||||||
| θ value | 6.88961 (rank : 51) | NC score | 0.015161 (rank : 55) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O94850 | Gene names | DDN, KIAA0749 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dendrin. | |||||
|
MYH13_HUMAN
|
||||||
| θ value | 6.88961 (rank : 52) | NC score | 0.020664 (rank : 49) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1504 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UKX3, O95252 | Gene names | MYH13 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-13 (Myosin heavy chain, skeletal muscle, extraocular) (MyHC- eo). | |||||
|
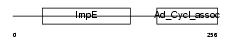
TTC28_HUMAN
|
||||||
| θ value | 6.88961 (rank : 53) | NC score | 0.012720 (rank : 57) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96AY4, O95928, O95929, Q9NTE4, Q9UG31, Q9UGG5, Q9UPV8, Q9Y3S5 | Gene names | TTC28, KIAA1043 | |||
|
Domain Architecture |
|
|||||
| Description | Tetratricopeptide repeat protein 28 (TPR repeat protein 28). | |||||
|
AZI1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 54) | NC score | 0.021983 (rank : 46) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 990 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q62036 | Gene names | Azi1, Az1, Azi | |||
|
Domain Architecture |

|
|||||
| Description | 5-azacytidine-induced protein 1 (Pre-acrosome localization protein 1). | |||||
|
CO039_MOUSE
|
||||||
| θ value | 8.99809 (rank : 55) | NC score | 0.021584 (rank : 47) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q3TEI4, Q3TMF2, Q3U253, Q3UM75, Q9DA93 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C15orf39 homolog. | |||||
|
GNRP_MOUSE
|
||||||
| θ value | 8.99809 (rank : 56) | NC score | 0.007304 (rank : 61) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P27671 | Gene names | Rasgrf1, Cdc25, Grf1 | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide-releasing protein (GNRP) (Ras-specific nucleotide exchange factor CDC25) (CDC25Mm). | |||||
|
K1377_HUMAN
|
||||||
| θ value | 8.99809 (rank : 57) | NC score | 0.015815 (rank : 53) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9P2H0, Q4G0U6 | Gene names | KIAA1377 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1377. | |||||
|
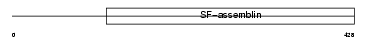
K1C14_MOUSE
|
||||||
| θ value | 8.99809 (rank : 58) | NC score | 0.008458 (rank : 60) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 501 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q61781, Q91VQ4 | Gene names | Krt14, Krt1-14 | |||
|
Domain Architecture |
|
|||||
| Description | Keratin, type I cytoskeletal 14 (Cytokeratin-14) (CK-14) (Keratin-14) (K14). | |||||
|
LZTS2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 59) | NC score | 0.027020 (rank : 26) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 423 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q91YU6, Q569X6 | Gene names | Lzts2, Kiaa1813 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine zipper putative tumor suppressor 2. | |||||
|
MDC1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 60) | NC score | 0.013855 (rank : 56) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
MK07_MOUSE
|
||||||
| θ value | 8.99809 (rank : 61) | NC score | 0.000005 (rank : 66) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1054 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9WVS8 | Gene names | Mapk7, Erk5 | |||
|
Domain Architecture |
|
|||||
| Description | Mitogen-activated protein kinase 7 (EC 2.7.11.24) (Extracellular signal-regulated kinase 5) (ERK-5) (BMK1 kinase). | |||||
|
PGCB_MOUSE
|
||||||
| θ value | 8.99809 (rank : 62) | NC score | 0.007022 (rank : 62) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 459 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61361 | Gene names | Bcan | |||
|
Domain Architecture |

|
|||||
| Description | Brevican core protein precursor. | |||||
|
TSC1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 63) | NC score | 0.022564 (rank : 45) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 765 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q92574 | Gene names | TSC1, KIAA0243, TSC | |||
|
Domain Architecture |

|
|||||
| Description | Hamartin (Tuberous sclerosis 1 protein). | |||||
|
WNK2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 64) | NC score | 0.001503 (rank : 65) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1407 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Y3S1, Q8IY36, Q9C0A3, Q9H3P4 | Gene names | WNK2, KIAA1760, PRKWNK2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK2 (EC 2.7.11.1) (Protein kinase with no lysine 2) (Protein kinase, lysine-deficient 2). | |||||
|
S100R_MOUSE
|
||||||
| θ value | θ > 10 (rank : 65) | NC score | 0.092628 (rank : 12) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99388 | Gene names | D1Lub1, Sp100-rs | |||
|
Domain Architecture |
|
|||||
| Description | Putative Sp100-related protein. | |||||
|
SP100_MOUSE
|
||||||
| θ value | θ > 10 (rank : 66) | NC score | 0.103396 (rank : 11) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O35892, O35897, O88392, O88395 | Gene names | Sp100 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear autoantigen Sp-100 (Speckled 100 kDa) (Nuclear dot-associated Sp100 protein). | |||||
|
GMEB2_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | P58929, Q6PCY0 | Gene names | Gmeb2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glucocorticoid modulatory element-binding protein 2 (GMEB-2). | |||||
|
GMEB2_HUMAN
|
||||||
| NC score | 0.987619 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9UKD1, Q5TDS0, Q9H431, Q9H4X7, Q9H4X8, Q9UF78, Q9ULF1 | Gene names | GMEB2, KIAA1269 | |||
|
Domain Architecture |

|
|||||
| Description | Glucocorticoid modulatory element-binding protein 2 (GMEB-2) (Parvovirus initiation factor p79) (PIF p79) (DNA-binding protein p79PIF). | |||||
|
GMEB1_MOUSE
|
||||||
| NC score | 0.920203 (rank : 3) | θ value | 1.22395e-89 (rank : 3) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9JL60 | Gene names | Gmeb1 | |||
|
Domain Architecture |

|
|||||
| Description | Glucocorticoid modulatory element-binding protein 1 (GMEB-1). | |||||
|
GMEB1_HUMAN
|
||||||
| NC score | 0.919711 (rank : 4) | θ value | 4.65103e-89 (rank : 4) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Y692, Q9NWH1, Q9UKD0 | Gene names | GMEB1 | |||
|
Domain Architecture |

|
|||||
| Description | Glucocorticoid modulatory element-binding protein 1 (GMEB-1) (Parvovirus initiation factor p96) (PIF p96) (DNA-binding protein p96PIF). | |||||
|
IPR1_MOUSE
|
||||||
| NC score | 0.384662 (rank : 5) | θ value | 1.29631e-06 (rank : 5) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8BVK9, Q3UCV7, Q80V00 | Gene names | Ipr1, Ifi75 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intracellular pathogen resistance protein 1. | |||||
|
DEAF1_HUMAN
|
||||||
| NC score | 0.342586 (rank : 6) | θ value | 2.21117e-06 (rank : 6) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O75398, O15152, O75399, O75510, O75511, O75512, O75513, Q9UET1 | Gene names | DEAF1, SPN, ZMYND5 | |||
|
Domain Architecture |

|
|||||
| Description | Deformed epidermal autoregulatory factor 1 homolog (Nuclear DEAF-1- related transcriptional regulator) (NUDR) (Suppressin) (Zinc finger MYND domain-containing protein 5). | |||||
|
DEAF1_MOUSE
|
||||||
| NC score | 0.339768 (rank : 7) | θ value | 2.21117e-06 (rank : 7) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Z1T5 | Gene names | Deaf1 | |||
|
Domain Architecture |

|
|||||
| Description | Deformed epidermal autoregulatory factor 1 homolog (Nuclear DEAF-1- related transcriptional regulator) (NUDR). | |||||
|
SP110_HUMAN
|
||||||
| NC score | 0.273446 (rank : 8) | θ value | 0.000270298 (rank : 9) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9HB58, Q14976, Q14977, Q8WUZ6, Q9HCT8 | Gene names | SP110 | |||
|
Domain Architecture |

|
|||||
| Description | Sp110 nuclear body protein (Speckled 110 kDa) (Transcriptional coactivator Sp110) (Interferon-induced protein 41/75). | |||||
|
LY10_HUMAN
|
||||||
| NC score | 0.252916 (rank : 9) | θ value | 0.00020696 (rank : 8) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q13342, Q13341, Q92881, Q96TG3 | Gene names | SP140, LYSP100 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear body protein SP140 (Nuclear autoantigen Sp-140) (Speckled 140 kDa) (LYSp100 protein) (Lymphoid-restricted homolog of Sp100). | |||||
|
SP100_HUMAN
|
||||||
| NC score | 0.201246 (rank : 10) | θ value | 0.00102713 (rank : 10) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P23497, O75450, Q13343, Q96F70, Q96T24, Q96T95, Q9NP33, Q9UE32 | Gene names | SP100 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear autoantigen Sp-100 (Speckled 100 kDa) (Nuclear dot-associated Sp100 protein) (Lysp100b). | |||||
|
SP100_MOUSE
|
||||||
| NC score | 0.103396 (rank : 11) | θ value | θ > 10 (rank : 66) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O35892, O35897, O88392, O88395 | Gene names | Sp100 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear autoantigen Sp-100 (Speckled 100 kDa) (Nuclear dot-associated Sp100 protein). | |||||
|
S100R_MOUSE
|
||||||
| NC score | 0.092628 (rank : 12) | θ value | θ > 10 (rank : 65) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99388 | Gene names | D1Lub1, Sp100-rs | |||
|
Domain Architecture |

|
|||||
| Description | Putative Sp100-related protein. | |||||
|
NELFA_MOUSE
|
||||||
| NC score | 0.065903 (rank : 13) | θ value | 0.279714 (rank : 14) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 195 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BG30, Q8BVE4, Q8VEI7, Q9CSJ9, Q9Z1V9 | Gene names | Whsc2, Nelfa, Whsc2h | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Negative elongation factor A (NELF-A) (Wolf-Hirschhorn syndrome candidate 2 homolog) (mWHSC2). | |||||
|
FA29A_HUMAN
|
||||||
| NC score | 0.063690 (rank : 14) | θ value | 1.06291 (rank : 21) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7Z4H7, Q6IQ10, Q6NZX5, Q8IZQ4, Q96FN0, Q9H950, Q9H998, Q9HCJ8, Q9NXT8 | Gene names | FAM29A, KIAA1574 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM29A. | |||||
|
NELFA_HUMAN
|
||||||
| NC score | 0.062344 (rank : 15) | θ value | 0.62314 (rank : 17) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H3P2, O95392 | Gene names | WHSC2, NELFA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Negative elongation factor A (NELF-A) (Wolf-Hirschhorn syndrome candidate 2 protein). | |||||
|
NICA_HUMAN
|
||||||
| NC score | 0.050318 (rank : 16) | θ value | 1.81305 (rank : 30) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q92542, Q86VV5 | Gene names | NCSTN, KIAA0253 | |||
|
Domain Architecture |

|
|||||
| Description | Nicastrin precursor. | |||||
|
NICA_MOUSE
|
||||||
| NC score | 0.050156 (rank : 17) | θ value | 1.81305 (rank : 31) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P57716, Q8VE20 | Gene names | Ncstn | |||
|
Domain Architecture |

|
|||||
| Description | Nicastrin precursor. | |||||
|
TM108_MOUSE
|
||||||
| NC score | 0.038446 (rank : 18) | θ value | 1.06291 (rank : 24) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 336 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BHE4, Q80WR9 | Gene names | Tmem108 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 108 precursor. | |||||
|
BRD8_MOUSE
|
||||||
| NC score | 0.036406 (rank : 19) | θ value | 5.27518 (rank : 43) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 422 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8R3B7, Q8C049, Q8R583, Q8VDP0 | Gene names | Brd8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 8. | |||||
|
RHG25_HUMAN
|
||||||
| NC score | 0.035462 (rank : 20) | θ value | 0.279714 (rank : 15) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 550 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P42331, Q8IXQ2 | Gene names | ARHGAP25, KIAA0053 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 25. | |||||
|
FYCO1_HUMAN
|
||||||
| NC score | 0.035004 (rank : 21) | θ value | 0.813845 (rank : 18) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1149 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9BQS8, Q3MJE6, Q86T41, Q86TB1, Q8TEF9, Q96IV5, Q9H8P9 | Gene names | FYCO1, ZFYVE7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE and coiled-coil domain-containing protein 1 (Zinc finger FYVE domain-containing protein 7). | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.034834 (rank : 22) | θ value | 0.163984 (rank : 12) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
KIF5A_HUMAN
|
||||||
| NC score | 0.028104 (rank : 23) | θ value | 0.365318 (rank : 16) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 843 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q12840, Q4LE26 | Gene names | KIF5A, NKHC1 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin heavy chain isoform 5A (Neuronal kinesin heavy chain) (NKHC) (Kinesin heavy chain neuron-specific 1). | |||||
|
ALEX_HUMAN
|
||||||
| NC score | 0.028010 (rank : 24) | θ value | 2.36792 (rank : 33) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P84996 | Gene names | GNAS, GNAS1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ALEX (Alternative gene product encoded by XL-exon). | |||||
|
RB6I2_HUMAN
|
||||||
| NC score | 0.027899 (rank : 25) | θ value | 1.06291 (rank : 23) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1584 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8IUD2, Q6NVK2, Q8IUD3, Q8IUD4, Q8IUD5, Q8NAS1, Q9NXN5, Q9UIK7, Q9UPS1 | Gene names | ERC1, ELKS, KIAA1081, RAB6IP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ELKS/RAB6-interacting/CAST family member 1 (RAB6-interacting protein 2) (ERC protein 1). | |||||
|
LZTS2_MOUSE
|
||||||
| NC score | 0.027020 (rank : 26) | θ value | 8.99809 (rank : 59) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 423 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q91YU6, Q569X6 | Gene names | Lzts2, Kiaa1813 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine zipper putative tumor suppressor 2. | |||||
|
CING_MOUSE
|
||||||
| NC score | 0.026806 (rank : 27) | θ value | 2.36792 (rank : 34) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1432 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P59242 | Gene names | Cgn | |||
|
Domain Architecture |

|
|||||
| Description | Cingulin. | |||||
|
KIF5A_MOUSE
|
||||||
| NC score | 0.026178 (rank : 28) | θ value | 0.813845 (rank : 19) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 859 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P33175, Q5DTP1, Q6PDY7, Q9Z2F9 | Gene names | Kif5a, Kiaa4086, Kif5, Nkhc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin heavy chain isoform 5A (Neuronal kinesin heavy chain) (NKHC) (Kinesin heavy chain neuron-specific 1). | |||||
|
RBL2_MOUSE
|
||||||
| NC score | 0.025788 (rank : 29) | θ value | 5.27518 (rank : 49) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q64700 | Gene names | Rbl2 | |||
|
Domain Architecture |

|
|||||
| Description | Retinoblastoma-like protein 2 (130 kDa retinoblastoma-associated protein) (PRB2) (P130) (RBR-2). | |||||
|
RBL1_HUMAN
|
||||||
| NC score | 0.025250 (rank : 30) | θ value | 4.03905 (rank : 41) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P28749, Q4VXA0, Q8N5K6, Q9H1L5, Q9H1M1 | Gene names | RBL1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinoblastoma-like protein 1 (107 kDa retinoblastoma-associated protein) (PRB1) (P107). | |||||
|
RBL2_HUMAN
|
||||||
| NC score | 0.025198 (rank : 31) | θ value | 5.27518 (rank : 48) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q08999, Q15073, Q16084, Q92812 | Gene names | RBL2, RB2 | |||
|
Domain Architecture |

|
|||||
| Description | Retinoblastoma-like protein 2 (130 kDa retinoblastoma-associated protein) (PRB2) (P130) (RBR-2). | |||||
|
KNTC2_MOUSE
|
||||||
| NC score | 0.025048 (rank : 32) | θ value | 4.03905 (rank : 39) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 692 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9D0F1, Q3TQT6, Q3UWM5, Q99P70 | Gene names | Kntc2, Hec1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinetochore protein Hec1 (Kinetochore-associated protein 2). | |||||
|
RB6I2_MOUSE
|
||||||
| NC score | 0.024987 (rank : 33) | θ value | 3.0926 (rank : 38) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1433 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q99MI1, Q80TK7, Q8BPL1, Q8C7Y1, Q99MI2 | Gene names | Erc1, Cast2, Kiaa1081, Rab6ip2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ELKS/RAB6-interacting/CAST family member 1 (RAB6-interacting protein 2) (ERC protein 1) (ERC1) (CAZ-associated structural protein 2) (CAST2). | |||||
|
CP250_HUMAN
|
||||||
| NC score | 0.024912 (rank : 34) | θ value | 0.279714 (rank : 13) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1805 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9BV73, O14812, O60588, Q9H450 | Gene names | CEP250, CEP2, CNAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1). | |||||
|
ATS12_HUMAN
|
||||||
| NC score | 0.024838 (rank : 35) | θ value | 0.125558 (rank : 11) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P58397, Q6UWL3 | Gene names | ADAMTS12 | |||
|
Domain Architecture |

|
|||||
| Description | ADAMTS-12 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase with thrombospondin motifs 12) (ADAM-TS 12) (ADAM-TS12). | |||||
|
CING_HUMAN
|
||||||
| NC score | 0.024113 (rank : 36) | θ value | 6.88961 (rank : 50) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1275 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9P2M7, Q5T386, Q9NR25 | Gene names | CGN, KIAA1319 | |||
|
Domain Architecture |

|
|||||
| Description | Cingulin. | |||||
|
NIN_HUMAN
|
||||||
| NC score | 0.023520 (rank : 37) | θ value | 5.27518 (rank : 47) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1416 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8N4C6, Q6P0P6, Q9BWU6, Q9C012, Q9C013, Q9C014, Q9H5I6, Q9HAT7, Q9HBY5, Q9HCK7, Q9UH61 | Gene names | NIN, KIAA1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein (hNinein) (Glycogen synthase kinase 3 beta-interacting protein) (GSK3B-interacting protein). | |||||
|
MYH6_HUMAN
|
||||||
| NC score | 0.023304 (rank : 38) | θ value | 1.38821 (rank : 25) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1426 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P13533, Q13943, Q14906, Q14907 | Gene names | MYH6, MYHCA | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
PME17_HUMAN
|
||||||
| NC score | 0.023175 (rank : 39) | θ value | 3.0926 (rank : 37) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P40967, Q12763, Q14448, Q14817, Q16565 | Gene names | SILV, D12S53E, PMEL17 | |||
|
Domain Architecture |

|
|||||
| Description | Melanocyte protein Pmel 17 precursor (Melanocyte lineage-specific antigen GP100) (Melanoma-associated ME20 antigen) (ME20M/ME20S) (ME20- M/ME20-S) (95 kDa melanocyte-specific secreted glycoprotein). | |||||
|
MYH1_HUMAN
|
||||||
| NC score | 0.023108 (rank : 40) | θ value | 5.27518 (rank : 46) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1478 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P12882, Q9Y622 | Gene names | MYH1 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-1 (Myosin heavy chain, skeletal muscle, adult 1) (Myosin heavy chain IIx/d) (MyHC-IIx/d). | |||||
|
MYH7_HUMAN
|
||||||
| NC score | 0.023079 (rank : 41) | θ value | 1.38821 (rank : 26) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P12883, Q14836, Q14837, Q14904, Q16579, Q92679, Q9H1D5, Q9UMM8 | Gene names | MYH7, MYHCB | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle beta isoform (MyHC-beta). | |||||
|
FLIP1_HUMAN
|
||||||
| NC score | 0.022924 (rank : 42) | θ value | 2.36792 (rank : 35) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q7Z7B0, Q5VUL6, Q86TC3, Q8N8B9, Q96SK6, Q9NVI8, Q9ULE5 | Gene names | FILIP1, KIAA1275 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-A-interacting protein 1 (FILIP). | |||||
|
FLIP1_MOUSE
|
||||||
| NC score | 0.022790 (rank : 43) | θ value | 5.27518 (rank : 45) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1176 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9CS72 | Gene names | Filip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-A-interacting protein 1 (FILIP). | |||||
|
AHTF1_HUMAN
|
||||||
| NC score | 0.022737 (rank : 44) | θ value | 2.36792 (rank : 32) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8WYP5, Q7Z4E3, Q8IZA4, Q96EH9, Q9Y4Q6 | Gene names | AHCTF1, ELYS, TMBS62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-hook-containing transcription factor 1 (Embryonic large molecule derived from yolk sac). | |||||
|
TSC1_HUMAN
|
||||||
| NC score | 0.022564 (rank : 45) | θ value | 8.99809 (rank : 63) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 765 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q92574 | Gene names | TSC1, KIAA0243, TSC | |||
|
Domain Architecture |

|
|||||
| Description | Hamartin (Tuberous sclerosis 1 protein). | |||||
|
AZI1_MOUSE
|
||||||
| NC score | 0.021983 (rank : 46) | θ value | 8.99809 (rank : 54) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 990 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q62036 | Gene names | Azi1, Az1, Azi | |||
|
Domain Architecture |

|
|||||
| Description | 5-azacytidine-induced protein 1 (Pre-acrosome localization protein 1). | |||||
|
CO039_MOUSE
|
||||||
| NC score | 0.021584 (rank : 47) | θ value | 8.99809 (rank : 55) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q3TEI4, Q3TMF2, Q3U253, Q3UM75, Q9DA93 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C15orf39 homolog. | |||||
|
MYH6_MOUSE
|
||||||
| NC score | 0.021452 (rank : 48) | θ value | 4.03905 (rank : 40) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1501 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q02566, Q64258, Q64738 | Gene names | Myh6, Myhca | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
MYH13_HUMAN
|
||||||
| NC score | 0.020664 (rank : 49) | θ value | 6.88961 (rank : 52) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1504 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UKX3, O95252 | Gene names | MYH13 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-13 (Myosin heavy chain, skeletal muscle, extraocular) (MyHC- eo). | |||||
|
ZHX1_MOUSE
|
||||||
| NC score | 0.018612 (rank : 50) | θ value | 1.38821 (rank : 27) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 334 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P70121, Q8BQ68, Q8C6T4, Q8CJG3 | Gene names | Zhx1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc fingers and homeoboxes protein 1. | |||||
|
EP400_HUMAN
|
||||||
| NC score | 0.018333 (rank : 51) | θ value | 1.06291 (rank : 20) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96L91, O15411, Q6P2F5, Q8N8Q7, Q8NE05, Q96JK7, Q9P230 | Gene names | EP400, CAGH32, KIAA1498, KIAA1818, TNRC12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E1A-binding protein p400 (EC 3.6.1.-) (p400 kDa SWI2/SNF2-related protein) (Domino homolog) (hDomino) (CAG repeat protein 32) (Trinucleotide repeat-containing gene 12 protein). | |||||
|
WDFY3_HUMAN
|
||||||
| NC score | 0.018115 (rank : 52) | θ value | 2.36792 (rank : 36) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 348 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8IZQ1, Q4W5K5, Q6P0Q5, Q8N1T2, Q96BS7, Q96N85, Q9Y2J7 | Gene names | WDFY3, KIAA0993 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat and FYVE domain-containing protein 3 (Autophagy-linked FYVE protein) (Alfy). | |||||
|
K1377_HUMAN
|
||||||
| NC score | 0.015815 (rank : 53) | θ value | 8.99809 (rank : 57) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9P2H0, Q4G0U6 | Gene names | KIAA1377 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1377. | |||||
|
BTNL1_MOUSE
|
||||||
| NC score | 0.015431 (rank : 54) | θ value | 1.81305 (rank : 28) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7TST0, O70356 | Gene names | Btnl1, Btnl3, Gm316, Ng10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Butyrophilin-like protein 1 precursor. | |||||
|
DEND_HUMAN
|
||||||
| NC score | 0.015161 (rank : 55) | θ value | 6.88961 (rank : 51) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O94850 | Gene names | DDN, KIAA0749 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dendrin. | |||||
|
MDC1_MOUSE
|
||||||
| NC score | 0.013855 (rank : 56) | θ value | 8.99809 (rank : 60) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
TTC28_HUMAN
|
||||||
| NC score | 0.012720 (rank : 57) | θ value | 6.88961 (rank : 53) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96AY4, O95928, O95929, Q9NTE4, Q9UG31, Q9UGG5, Q9UPV8, Q9Y3S5 | Gene names | TTC28, KIAA1043 | |||
|
Domain Architecture |

|
|||||
| Description | Tetratricopeptide repeat protein 28 (TPR repeat protein 28). | |||||
|
PRDM2_HUMAN
|
||||||
| NC score | 0.011703 (rank : 58) | θ value | 1.06291 (rank : 22) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q13029, Q13149, Q14550, Q5VUL9 | Gene names | PRDM2, RIZ | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 2 (PR domain-containing protein 2) (Retinoblastoma protein-interacting zinc-finger protein) (Zinc finger protein RIZ) (MTE-binding protein) (MTB-ZF) (GATA-3-binding protein G3B). | |||||
|
RXRA_HUMAN
|
||||||
| NC score | 0.009486 (rank : 59) | θ value | 4.03905 (rank : 42) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P19793, Q2V504 | Gene names | RXRA, NR2B1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinoic acid receptor RXR-alpha (Retinoid X receptor alpha). | |||||
|
K1C14_MOUSE
|
||||||
| NC score | 0.008458 (rank : 60) | θ value | 8.99809 (rank : 58) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 501 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q61781, Q91VQ4 | Gene names | Krt14, Krt1-14 | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type I cytoskeletal 14 (Cytokeratin-14) (CK-14) (Keratin-14) (K14). | |||||
|
GNRP_MOUSE
|
||||||
| NC score | 0.007304 (rank : 61) | θ value | 8.99809 (rank : 56) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P27671 | Gene names | Rasgrf1, Cdc25, Grf1 | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide-releasing protein (GNRP) (Ras-specific nucleotide exchange factor CDC25) (CDC25Mm). | |||||
|
PGCB_MOUSE
|
||||||
| NC score | 0.007022 (rank : 62) | θ value | 8.99809 (rank : 62) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 459 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61361 | Gene names | Bcan | |||
|
Domain Architecture |

|
|||||
| Description | Brevican core protein precursor. | |||||
|
IRAK1_HUMAN
|
||||||
| NC score | 0.005733 (rank : 63) | θ value | 1.81305 (rank : 29) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 820 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P51617, Q7Z5V4, Q96RL2 | Gene names | IRAK1, IRAK | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin-1 receptor-associated kinase 1 (EC 2.7.11.1) (IRAK-1). | |||||
|
DCC_MOUSE
|
||||||
| NC score | 0.005634 (rank : 64) | θ value | 5.27518 (rank : 44) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P70211 | Gene names | Dcc | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC). | |||||
|
WNK2_HUMAN
|
||||||
| NC score | 0.001503 (rank : 65) | θ value | 8.99809 (rank : 64) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1407 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Y3S1, Q8IY36, Q9C0A3, Q9H3P4 | Gene names | WNK2, KIAA1760, PRKWNK2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK2 (EC 2.7.11.1) (Protein kinase with no lysine 2) (Protein kinase, lysine-deficient 2). | |||||
|
MK07_MOUSE
|
||||||
| NC score | 0.000005 (rank : 66) | θ value | 8.99809 (rank : 61) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1054 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9WVS8 | Gene names | Mapk7, Erk5 | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase 7 (EC 2.7.11.24) (Extracellular signal-regulated kinase 5) (ERK-5) (BMK1 kinase). | |||||