Please be patient as the page loads
|
UBAP1_HUMAN
|
||||||
| SwissProt Accessions | Q9NZ09, Q4V759, Q5T7B3, Q6FI75, Q8NC52, Q8NCG6, Q8NCH9 | Gene names | UBAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-associated protein 1 (UBAP). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
UBAP1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9NZ09, Q4V759, Q5T7B3, Q6FI75, Q8NC52, Q8NCG6, Q8NCH9 | Gene names | UBAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-associated protein 1 (UBAP). | |||||
|
UBAP1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.992585 (rank : 2) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8BH48, Q8BQ80, Q8BSW6, Q9D749, Q9ERV5 | Gene names | Ubap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-associated protein 1 (UBAP) (Ubiquitin-associated protein NAG20). | |||||
|
UBP5_HUMAN
|
||||||
| θ value | 0.00665767 (rank : 3) | NC score | 0.150712 (rank : 3) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P45974, Q96J22 | Gene names | USP5, ISOT | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 5 (EC 3.1.2.15) (Ubiquitin thioesterase 5) (Ubiquitin-specific-processing protease 5) (Deubiquitinating enzyme 5) (Isopeptidase T). | |||||
|
UBP5_MOUSE
|
||||||
| θ value | 0.00665767 (rank : 4) | NC score | 0.150691 (rank : 4) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P56399 | Gene names | Usp5, Isot | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 5 (EC 3.1.2.15) (Ubiquitin thioesterase 5) (Ubiquitin-specific-processing protease 5) (Deubiquitinating enzyme 5) (Isopeptidase T). | |||||
|
COA1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 5) | NC score | 0.042276 (rank : 9) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13085 | Gene names | ACACA, ACAC, ACC1, ACCA | |||
|
Domain Architecture |

|
|||||
| Description | Acetyl-CoA carboxylase 1 (EC 6.4.1.2) (ACC-alpha) [Includes: Biotin carboxylase (EC 6.3.4.14)]. | |||||
|
NOTC2_MOUSE
|
||||||
| θ value | 0.813845 (rank : 6) | NC score | 0.008807 (rank : 29) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1069 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O35516, Q06008, Q60941 | Gene names | Notch2 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 2 precursor (Notch 2) (Motch B) [Contains: Notch 2 extracellular truncation; Notch 2 intracellular domain]. | |||||
|
CHD9_HUMAN
|
||||||
| θ value | 1.06291 (rank : 7) | NC score | 0.021245 (rank : 15) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q3L8U1, O15025, Q1WF12, Q6DTK9, Q9H9V7 | Gene names | CHD9, KIAA0308, KISH2, PRIC320 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain-helicase-DNA-binding protein 9 (EC 3.6.1.-) (ATP- dependent helicase CHD9) (CHD-9) (Chromatin-related mesenchymal modulator) (CReMM) (Chromatin remodeling factor CHROM1) (Peroxisomal proliferator-activated receptor A-interacting complex 320 kDa protein) (PPAR-alpha-interacting complex protein 320 kDa) (Kismet homolog 2). | |||||
|
NEIL3_HUMAN
|
||||||
| θ value | 1.06291 (rank : 8) | NC score | 0.041177 (rank : 10) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8TAT5, Q2PPJ3, Q8NG51, Q9NV95 | Gene names | NEIL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endonuclease VIII-like 3 (Nei-like 3) (DNA glycosylase FPG2). | |||||
|
DLEC1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 9) | NC score | 0.040284 (rank : 12) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y238, Q9NSW0, Q9NTG5 | Gene names | DLEC1, DLC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Deleted in lung and esophageal cancer protein 1 (Deleted in lung cancer protein 1) (DLC-1). | |||||
|
GP176_MOUSE
|
||||||
| θ value | 1.38821 (rank : 10) | NC score | 0.013724 (rank : 24) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 539 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q80WT4, Q80UC4 | Gene names | Gpr176, Agr9, Gm1012 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 176 (G-protein coupled receptor AGR9). | |||||
|
VMDL3_HUMAN
|
||||||
| θ value | 1.81305 (rank : 11) | NC score | 0.023064 (rank : 14) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N1M1, Q8N356, Q8NFT9, Q9BR80 | Gene names | VMD2L3 | |||
|
Domain Architecture |

|
|||||
| Description | Bestrophin-4 (Vitelliform macular dystrophy 2-like protein 3). | |||||
|
CCHL_MOUSE
|
||||||
| θ value | 2.36792 (rank : 12) | NC score | 0.064260 (rank : 7) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P53702 | Gene names | Hccs, Cchl | |||
|
Domain Architecture |

|
|||||
| Description | Cytochrome c-type heme lyase (EC 4.4.1.17) (CCHL) (Holocytochrome c- type synthase). | |||||
|
UBP13_HUMAN
|
||||||
| θ value | 2.36792 (rank : 13) | NC score | 0.104002 (rank : 5) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q92995 | Gene names | USP13, ISOT3 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 13 (EC 3.1.2.15) (Ubiquitin thioesterase 13) (Ubiquitin-specific-processing protease 13) (Deubiquitinating enzyme 13) (Isopeptidase T-3) (ISOT-3). | |||||
|
V2RX_MOUSE
|
||||||
| θ value | 2.36792 (rank : 14) | NC score | 0.021099 (rank : 16) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O70410 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Vomeronasal type-2 receptor X precursor (Putative pheromone receptor V2RX). | |||||
|
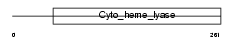
CCHL_HUMAN
|
||||||
| θ value | 3.0926 (rank : 15) | NC score | 0.060634 (rank : 8) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P53701, Q502X8 | Gene names | HCCS, CCHL | |||
|
Domain Architecture |
|
|||||
| Description | Cytochrome c-type heme lyase (EC 4.4.1.17) (CCHL) (Holocytochrome c- type synthase). | |||||
|
CEP41_MOUSE
|
||||||
| θ value | 3.0926 (rank : 16) | NC score | 0.031789 (rank : 13) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99NF3, Q8BT53 | Gene names | Tsga14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 41 kDa (Cep41 protein) (Testis-specific gene A14 protein). | |||||
|
KNTC2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 17) | NC score | 0.019039 (rank : 18) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 673 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O14777, Q6PJX2 | Gene names | KNTC2, HEC, HEC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinetochore protein Hec1 (HsHec1) (Kinetochore-associated protein 2) (Highly expressed in cancer protein) (Retinoblastoma-associated protein HEC). | |||||
|
LGR5_HUMAN
|
||||||
| θ value | 3.0926 (rank : 18) | NC score | 0.015155 (rank : 22) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75473, Q9UP75 | Gene names | LGR5, GPR49, GPR67 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeat-containing G-protein coupled receptor 5 precursor (Orphan G-protein coupled receptor HG38) (G-protein coupled receptor 49) (G-protein coupled receptor 67). | |||||
|
LGR5_MOUSE
|
||||||
| θ value | 3.0926 (rank : 19) | NC score | 0.015271 (rank : 21) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 484 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z1P4 | Gene names | Lgr5, Fex, Gpr49 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeat-containing G-protein coupled receptor 5 precursor (G-protein coupled receptor 49) (Orphan G-protein coupled receptor FEX). | |||||
|
MUC13_HUMAN
|
||||||
| θ value | 3.0926 (rank : 20) | NC score | 0.040413 (rank : 11) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H3R2, Q6UWD9, Q9NXT5 | Gene names | MUC13, DRCC1, RECC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-13 precursor (Down-regulated in colon cancer 1). | |||||
|
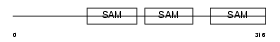
LIPB1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 21) | NC score | 0.010844 (rank : 28) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 727 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8C8U0, Q69ZN5, Q6GQV3, Q80VB4, Q9CUT7 | Gene names | Ppfibp1, Kiaa1230 | |||
|
Domain Architecture |
|
|||||
| Description | Liprin-beta-1 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein-binding protein 1) (PTPRF-interacting protein-binding protein 1). | |||||
|
CT141_HUMAN
|
||||||
| θ value | 5.27518 (rank : 22) | NC score | 0.079296 (rank : 6) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NUB4 | Gene names | C20orf141 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C20orf141. | |||||
|
MORC1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 23) | NC score | 0.014900 (rank : 23) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9WVL5 | Gene names | Morc1, Morc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORC family CW-type zinc finger protein 1 (Protein microrchidia). | |||||
|
OSBL9_HUMAN
|
||||||
| θ value | 5.27518 (rank : 24) | NC score | 0.013666 (rank : 25) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96SU4, Q9H9X2 | Gene names | OSBPL9, ORP9, OSBP4 | |||
|
Domain Architecture |

|
|||||
| Description | Oxysterol-binding protein-related protein 9 (OSBP-related protein 9) (ORP-9). | |||||
|
SPTN4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 25) | NC score | 0.006684 (rank : 30) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 570 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9H254, Q9H1K7, Q9H1K8, Q9H1K9, Q9H3G8, Q9HCD0 | Gene names | SPTBN4, KIAA1642, SPTBN3 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin beta chain, brain 3 (Spectrin, non-erythroid beta chain 3) (Beta-IV spectrin). | |||||
|
SYGP1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 26) | NC score | 0.011832 (rank : 26) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96PV0, Q8TCS2, Q9UGE2 | Gene names | SYNGAP1, KIAA1938 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein SynGAP (Synaptic Ras-GTPase-activating protein 1) (Synaptic Ras-GAP 1) (Neuronal RasGAP). | |||||
|
TAB2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 27) | NC score | 0.017960 (rank : 19) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NYJ8, O94838, Q6I9W8, Q76N06, Q9UFP7 | Gene names | MAP3K7IP2, KIAA0733, TAB2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase 7-interacting protein 2 (TAK1-binding protein 2). | |||||
|
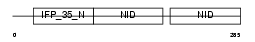
NMI_HUMAN
|
||||||
| θ value | 6.88961 (rank : 28) | NC score | 0.019142 (rank : 17) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q13287, Q53TI8, Q9BVE5 | Gene names | NMI | |||
|
Domain Architecture |
|
|||||
| Description | N-myc-interactor (Nmi) (N-myc and STAT interactor). | |||||
|
TLX1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 29) | NC score | 0.005271 (rank : 31) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P31314, O75699, Q9HCA0 | Gene names | TLX1, HOX11, TCL3 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell leukemia homeobox protein 1 (Homeobox protein Hox-11) (TCL-3 proto-oncogene). | |||||
|
CXXC6_HUMAN
|
||||||
| θ value | 8.99809 (rank : 30) | NC score | 0.017425 (rank : 20) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NFU7, Q5VUP7, Q7Z6B6, Q8TCR1, Q9C0I7 | Gene names | CXXC6, KIAA1676, LCX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CXXC-type zinc finger protein 6 (Leukemia-associated protein with a CXXC domain). | |||||
|
TENS1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 31) | NC score | 0.010977 (rank : 27) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 400 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9HBL0 | Gene names | TNS1, TNS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tensin-1. | |||||
|
UBAP1_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9NZ09, Q4V759, Q5T7B3, Q6FI75, Q8NC52, Q8NCG6, Q8NCH9 | Gene names | UBAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-associated protein 1 (UBAP). | |||||
|
UBAP1_MOUSE
|
||||||
| NC score | 0.992585 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8BH48, Q8BQ80, Q8BSW6, Q9D749, Q9ERV5 | Gene names | Ubap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-associated protein 1 (UBAP) (Ubiquitin-associated protein NAG20). | |||||
|
UBP5_HUMAN
|
||||||
| NC score | 0.150712 (rank : 3) | θ value | 0.00665767 (rank : 3) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P45974, Q96J22 | Gene names | USP5, ISOT | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 5 (EC 3.1.2.15) (Ubiquitin thioesterase 5) (Ubiquitin-specific-processing protease 5) (Deubiquitinating enzyme 5) (Isopeptidase T). | |||||
|
UBP5_MOUSE
|
||||||
| NC score | 0.150691 (rank : 4) | θ value | 0.00665767 (rank : 4) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P56399 | Gene names | Usp5, Isot | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 5 (EC 3.1.2.15) (Ubiquitin thioesterase 5) (Ubiquitin-specific-processing protease 5) (Deubiquitinating enzyme 5) (Isopeptidase T). | |||||
|
UBP13_HUMAN
|
||||||
| NC score | 0.104002 (rank : 5) | θ value | 2.36792 (rank : 13) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q92995 | Gene names | USP13, ISOT3 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 13 (EC 3.1.2.15) (Ubiquitin thioesterase 13) (Ubiquitin-specific-processing protease 13) (Deubiquitinating enzyme 13) (Isopeptidase T-3) (ISOT-3). | |||||
|
CT141_HUMAN
|
||||||
| NC score | 0.079296 (rank : 6) | θ value | 5.27518 (rank : 22) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NUB4 | Gene names | C20orf141 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C20orf141. | |||||
|
CCHL_MOUSE
|
||||||
| NC score | 0.064260 (rank : 7) | θ value | 2.36792 (rank : 12) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P53702 | Gene names | Hccs, Cchl | |||
|
Domain Architecture |

|
|||||
| Description | Cytochrome c-type heme lyase (EC 4.4.1.17) (CCHL) (Holocytochrome c- type synthase). | |||||
|
CCHL_HUMAN
|
||||||
| NC score | 0.060634 (rank : 8) | θ value | 3.0926 (rank : 15) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P53701, Q502X8 | Gene names | HCCS, CCHL | |||
|
Domain Architecture |

|
|||||
| Description | Cytochrome c-type heme lyase (EC 4.4.1.17) (CCHL) (Holocytochrome c- type synthase). | |||||
|
COA1_HUMAN
|
||||||
| NC score | 0.042276 (rank : 9) | θ value | 0.813845 (rank : 5) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13085 | Gene names | ACACA, ACAC, ACC1, ACCA | |||
|
Domain Architecture |

|
|||||
| Description | Acetyl-CoA carboxylase 1 (EC 6.4.1.2) (ACC-alpha) [Includes: Biotin carboxylase (EC 6.3.4.14)]. | |||||
|
NEIL3_HUMAN
|
||||||
| NC score | 0.041177 (rank : 10) | θ value | 1.06291 (rank : 8) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8TAT5, Q2PPJ3, Q8NG51, Q9NV95 | Gene names | NEIL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endonuclease VIII-like 3 (Nei-like 3) (DNA glycosylase FPG2). | |||||
|
MUC13_HUMAN
|
||||||
| NC score | 0.040413 (rank : 11) | θ value | 3.0926 (rank : 20) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H3R2, Q6UWD9, Q9NXT5 | Gene names | MUC13, DRCC1, RECC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-13 precursor (Down-regulated in colon cancer 1). | |||||
|
DLEC1_HUMAN
|
||||||
| NC score | 0.040284 (rank : 12) | θ value | 1.38821 (rank : 9) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y238, Q9NSW0, Q9NTG5 | Gene names | DLEC1, DLC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Deleted in lung and esophageal cancer protein 1 (Deleted in lung cancer protein 1) (DLC-1). | |||||
|
CEP41_MOUSE
|
||||||
| NC score | 0.031789 (rank : 13) | θ value | 3.0926 (rank : 16) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99NF3, Q8BT53 | Gene names | Tsga14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 41 kDa (Cep41 protein) (Testis-specific gene A14 protein). | |||||
|
VMDL3_HUMAN
|
||||||
| NC score | 0.023064 (rank : 14) | θ value | 1.81305 (rank : 11) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N1M1, Q8N356, Q8NFT9, Q9BR80 | Gene names | VMD2L3 | |||
|
Domain Architecture |

|
|||||
| Description | Bestrophin-4 (Vitelliform macular dystrophy 2-like protein 3). | |||||
|
CHD9_HUMAN
|
||||||
| NC score | 0.021245 (rank : 15) | θ value | 1.06291 (rank : 7) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q3L8U1, O15025, Q1WF12, Q6DTK9, Q9H9V7 | Gene names | CHD9, KIAA0308, KISH2, PRIC320 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain-helicase-DNA-binding protein 9 (EC 3.6.1.-) (ATP- dependent helicase CHD9) (CHD-9) (Chromatin-related mesenchymal modulator) (CReMM) (Chromatin remodeling factor CHROM1) (Peroxisomal proliferator-activated receptor A-interacting complex 320 kDa protein) (PPAR-alpha-interacting complex protein 320 kDa) (Kismet homolog 2). | |||||
|
V2RX_MOUSE
|
||||||
| NC score | 0.021099 (rank : 16) | θ value | 2.36792 (rank : 14) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O70410 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Vomeronasal type-2 receptor X precursor (Putative pheromone receptor V2RX). | |||||
|
NMI_HUMAN
|
||||||
| NC score | 0.019142 (rank : 17) | θ value | 6.88961 (rank : 28) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q13287, Q53TI8, Q9BVE5 | Gene names | NMI | |||
|
Domain Architecture |

|
|||||
| Description | N-myc-interactor (Nmi) (N-myc and STAT interactor). | |||||
|
KNTC2_HUMAN
|
||||||
| NC score | 0.019039 (rank : 18) | θ value | 3.0926 (rank : 17) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 673 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O14777, Q6PJX2 | Gene names | KNTC2, HEC, HEC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinetochore protein Hec1 (HsHec1) (Kinetochore-associated protein 2) (Highly expressed in cancer protein) (Retinoblastoma-associated protein HEC). | |||||
|
TAB2_HUMAN
|
||||||
| NC score | 0.017960 (rank : 19) | θ value | 5.27518 (rank : 27) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NYJ8, O94838, Q6I9W8, Q76N06, Q9UFP7 | Gene names | MAP3K7IP2, KIAA0733, TAB2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase 7-interacting protein 2 (TAK1-binding protein 2). | |||||
|
CXXC6_HUMAN
|
||||||
| NC score | 0.017425 (rank : 20) | θ value | 8.99809 (rank : 30) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NFU7, Q5VUP7, Q7Z6B6, Q8TCR1, Q9C0I7 | Gene names | CXXC6, KIAA1676, LCX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CXXC-type zinc finger protein 6 (Leukemia-associated protein with a CXXC domain). | |||||
|
LGR5_MOUSE
|
||||||
| NC score | 0.015271 (rank : 21) | θ value | 3.0926 (rank : 19) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 484 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z1P4 | Gene names | Lgr5, Fex, Gpr49 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeat-containing G-protein coupled receptor 5 precursor (G-protein coupled receptor 49) (Orphan G-protein coupled receptor FEX). | |||||
|
LGR5_HUMAN
|
||||||
| NC score | 0.015155 (rank : 22) | θ value | 3.0926 (rank : 18) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75473, Q9UP75 | Gene names | LGR5, GPR49, GPR67 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeat-containing G-protein coupled receptor 5 precursor (Orphan G-protein coupled receptor HG38) (G-protein coupled receptor 49) (G-protein coupled receptor 67). | |||||
|
MORC1_MOUSE
|
||||||
| NC score | 0.014900 (rank : 23) | θ value | 5.27518 (rank : 23) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9WVL5 | Gene names | Morc1, Morc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORC family CW-type zinc finger protein 1 (Protein microrchidia). | |||||
|
GP176_MOUSE
|
||||||
| NC score | 0.013724 (rank : 24) | θ value | 1.38821 (rank : 10) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 539 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q80WT4, Q80UC4 | Gene names | Gpr176, Agr9, Gm1012 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 176 (G-protein coupled receptor AGR9). | |||||
|
OSBL9_HUMAN
|
||||||
| NC score | 0.013666 (rank : 25) | θ value | 5.27518 (rank : 24) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96SU4, Q9H9X2 | Gene names | OSBPL9, ORP9, OSBP4 | |||
|
Domain Architecture |

|
|||||
| Description | Oxysterol-binding protein-related protein 9 (OSBP-related protein 9) (ORP-9). | |||||
|
SYGP1_HUMAN
|
||||||
| NC score | 0.011832 (rank : 26) | θ value | 5.27518 (rank : 26) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96PV0, Q8TCS2, Q9UGE2 | Gene names | SYNGAP1, KIAA1938 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein SynGAP (Synaptic Ras-GTPase-activating protein 1) (Synaptic Ras-GAP 1) (Neuronal RasGAP). | |||||
|
TENS1_HUMAN
|
||||||
| NC score | 0.010977 (rank : 27) | θ value | 8.99809 (rank : 31) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 400 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9HBL0 | Gene names | TNS1, TNS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tensin-1. | |||||
|
LIPB1_MOUSE
|
||||||
| NC score | 0.010844 (rank : 28) | θ value | 4.03905 (rank : 21) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 727 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8C8U0, Q69ZN5, Q6GQV3, Q80VB4, Q9CUT7 | Gene names | Ppfibp1, Kiaa1230 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-beta-1 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein-binding protein 1) (PTPRF-interacting protein-binding protein 1). | |||||
|
NOTC2_MOUSE
|
||||||
| NC score | 0.008807 (rank : 29) | θ value | 0.813845 (rank : 6) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1069 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O35516, Q06008, Q60941 | Gene names | Notch2 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 2 precursor (Notch 2) (Motch B) [Contains: Notch 2 extracellular truncation; Notch 2 intracellular domain]. | |||||
|
SPTN4_HUMAN
|
||||||
| NC score | 0.006684 (rank : 30) | θ value | 5.27518 (rank : 25) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 570 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9H254, Q9H1K7, Q9H1K8, Q9H1K9, Q9H3G8, Q9HCD0 | Gene names | SPTBN4, KIAA1642, SPTBN3 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin beta chain, brain 3 (Spectrin, non-erythroid beta chain 3) (Beta-IV spectrin). | |||||
|
TLX1_HUMAN
|
||||||
| NC score | 0.005271 (rank : 31) | θ value | 6.88961 (rank : 29) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P31314, O75699, Q9HCA0 | Gene names | TLX1, HOX11, TCL3 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell leukemia homeobox protein 1 (Homeobox protein Hox-11) (TCL-3 proto-oncogene). | |||||