Please be patient as the page loads
|
TRAF1_MOUSE
|
||||||
| SwissProt Accessions | P39428 | Gene names | Traf1 | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 1. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
TRAF1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.993061 (rank : 2) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q13077, Q8NF13 | Gene names | TRAF1, EBI6 | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 1 (Epstein-Barr virus-induced protein 6). | |||||
|
TRAF1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P39428 | Gene names | Traf1 | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 1. | |||||
|
TRAF2_MOUSE
|
||||||
| θ value | 1.22964e-73 (rank : 3) | NC score | 0.861801 (rank : 4) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P39429, O54896 | Gene names | Traf2 | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 2. | |||||
|
TRAF2_HUMAN
|
||||||
| θ value | 2.31901e-72 (rank : 4) | NC score | 0.867370 (rank : 3) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q12933, Q96NT2 | Gene names | TRAF2, TRAP3 | |||
|
Domain Architecture |
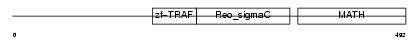
|
|||||
| Description | TNF receptor-associated factor 2 (Tumor necrosis factor type 2 receptor-associated protein 3). | |||||
|
TRAF3_MOUSE
|
||||||
| θ value | 6.99855e-61 (rank : 5) | NC score | 0.809459 (rank : 7) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 476 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q60803, Q62380 | Gene names | Traf3, Craf1, Trafamn | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 3 (CD40 receptor-associated factor 1) (CRAF1) (TRAFAMN). | |||||
|
TRAF3_HUMAN
|
||||||
| θ value | 3.47335e-60 (rank : 6) | NC score | 0.808557 (rank : 8) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 463 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q13114, Q12990, Q13076, Q13947, Q6AZX1, Q9UNL1 | Gene names | TRAF3, CAP1, CRAF1 | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 3 (CD40 receptor-associated factor 1) (CRAF1) (CD40-binding protein) (CD40BP) (LMP1-associated protein) (LAP1) (CAP-1). | |||||
|
TRAF5_MOUSE
|
||||||
| θ value | 8.00737e-57 (rank : 7) | NC score | 0.811958 (rank : 6) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 355 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P70191, Q61480 | Gene names | Traf5 | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 5. | |||||
|
TRAF5_HUMAN
|
||||||
| θ value | 3.974e-56 (rank : 8) | NC score | 0.813615 (rank : 5) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 396 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O00463 | Gene names | TRAF5, RNF84 | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 5 (RING finger protein 84). | |||||
|
TRAF4_HUMAN
|
||||||
| θ value | 9.2256e-29 (rank : 9) | NC score | 0.715634 (rank : 9) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 373 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9BUZ4, O75615, Q14848, Q2PJN8 | Gene names | TRAF4, CART1, MLN62, RNF83 | |||
|
Domain Architecture |
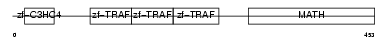
|
|||||
| Description | TNF receptor-associated factor 4 (Cysteine-rich domain associated with RING and Traf domains protein 1) (Malignant 62) (RING finger protein 83). | |||||
|
TRAF4_MOUSE
|
||||||
| θ value | 2.96777e-27 (rank : 10) | NC score | 0.715420 (rank : 10) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 338 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q61382 | Gene names | Traf4, Cart1 | |||
|
Domain Architecture |
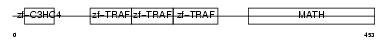
|
|||||
| Description | TNF receptor-associated factor 4 (Cysteine-rich motif associated to RING and Traf domains protein 1). | |||||
|
TRAF6_HUMAN
|
||||||
| θ value | 7.56453e-23 (rank : 11) | NC score | 0.700788 (rank : 11) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9Y4K3, Q8NEH5 | Gene names | TRAF6, RNF85 | |||
|
Domain Architecture |
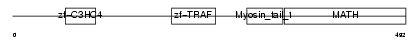
|
|||||
| Description | TNF receptor-associated factor 6 (Interleukin 1 signal transducer) (RING finger protein 85). | |||||
|
TRAF6_MOUSE
|
||||||
| θ value | 5.42112e-21 (rank : 12) | NC score | 0.698528 (rank : 12) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P70196, Q8BLV2 | Gene names | Traf6 | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 6. | |||||
|
MEP1A_MOUSE
|
||||||
| θ value | 0.000158464 (rank : 13) | NC score | 0.216853 (rank : 14) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P28825 | Gene names | Mep1a | |||
|
Domain Architecture |

|
|||||
| Description | Meprin A subunit alpha precursor (EC 3.4.24.18) (Endopeptidase-2) (MEP-1). | |||||
|
TACC3_MOUSE
|
||||||
| θ value | 0.0148317 (rank : 14) | NC score | 0.065176 (rank : 51) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 607 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9JJ11, Q9WVK9 | Gene names | Tacc3, Aint | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 3 (ARNT-interacting protein). | |||||
|
MEP1A_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 15) | NC score | 0.183769 (rank : 17) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q16819 | Gene names | MEP1A | |||
|
Domain Architecture |

|
|||||
| Description | Meprin A subunit alpha precursor (EC 3.4.24.18) (Endopeptidase-2) (N- benzoyl-L-tyrosyl-P-amino-benzoic acid hydrolase subunit alpha) (PABA peptide hydrolase) (PPH alpha). | |||||
|
MEP1B_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 16) | NC score | 0.218789 (rank : 13) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q61847 | Gene names | Mep1b, Mep-1b | |||
|
Domain Architecture |

|
|||||
| Description | Meprin A subunit beta precursor (EC 3.4.24.18) (Endopeptidase-2). | |||||
|
CP250_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 17) | NC score | 0.032405 (rank : 82) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1805 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9BV73, O14812, O60588, Q9H450 | Gene names | CEP250, CEP2, CNAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1). | |||||
|
CKAP4_MOUSE
|
||||||
| θ value | 0.279714 (rank : 18) | NC score | 0.041284 (rank : 78) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 596 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8BMK4, Q8BTK8, Q8R3F2 | Gene names | Ckap4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-associated protein 4 (63 kDa membrane protein) (p63). | |||||
|
CE290_HUMAN
|
||||||
| θ value | 0.365318 (rank : 19) | NC score | 0.043614 (rank : 76) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1553 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O15078, Q1PSK5, Q66GS8, Q9H2G6, Q9H6Q7, Q9H8I0 | Gene names | CEP290, KIAA0373, NPHP6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein Cep290 (Nephrocystin-6) (Tumor antigen se2-2). | |||||
|
RUFY2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 20) | NC score | 0.026629 (rank : 89) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1208 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8WXA3, Q96P51, Q9P1Z1 | Gene names | RUFY2, KIAA1537, RABIP4R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and FYVE domain-containing protein 2 (Rab4-interacting protein related). | |||||
|
PLEC1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 21) | NC score | 0.026537 (rank : 90) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1326 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q15149, Q15148, Q16640 | Gene names | PLEC1 | |||
|
Domain Architecture |

|
|||||
| Description | Plectin-1 (PLTN) (PCN) (Hemidesmosomal protein 1) (HD1). | |||||
|
TRAF7_MOUSE
|
||||||
| θ value | 0.813845 (rank : 22) | NC score | 0.150452 (rank : 23) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 470 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q922B6, Q99JV3 | Gene names | Traf7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin protein ligase TRAF7 (EC 6.3.2.-) (TNF receptor- associated factor 7). | |||||
|
TRAF7_HUMAN
|
||||||
| θ value | 1.06291 (rank : 23) | NC score | 0.148445 (rank : 24) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 482 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q6Q0C0, Q9H073 | Gene names | TRAF7, RFWD1, RNF119 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin protein ligase TRAF7 (EC 6.3.2.-) (TNF receptor- associated factor 7) (RING finger and WD repeat domain protein 1) (RING finger protein 119). | |||||
|
TTC17_HUMAN
|
||||||
| θ value | 1.06291 (rank : 24) | NC score | 0.034660 (rank : 80) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96AE7 | Gene names | TTC17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tetratricopeptide repeat protein 17 (TPR repeat protein 17). | |||||
|
LAMB1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 25) | NC score | 0.016955 (rank : 101) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1018 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P07942 | Gene names | LAMB1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-1 chain precursor (Laminin B1 chain). | |||||
|
MYH6_MOUSE
|
||||||
| θ value | 1.38821 (rank : 26) | NC score | 0.028923 (rank : 86) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1501 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q02566, Q64258, Q64738 | Gene names | Myh6, Myhca | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
MYH7_HUMAN
|
||||||
| θ value | 1.38821 (rank : 27) | NC score | 0.030926 (rank : 83) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P12883, Q14836, Q14837, Q14904, Q16579, Q92679, Q9H1D5, Q9UMM8 | Gene names | MYH7, MYHCB | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle beta isoform (MyHC-beta). | |||||
|
RAB6C_HUMAN
|
||||||
| θ value | 1.81305 (rank : 28) | NC score | 0.006483 (rank : 106) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H0N0, Q9P128 | Gene names | RAB6C, WTH3 | |||
|
Domain Architecture |
|
|||||
| Description | Ras-related protein Rab-6C (Rab6-like protein WTH3). | |||||
|
CRIM1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 29) | NC score | 0.017556 (rank : 99) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NZV1, Q59GH0, Q7LCQ5, Q9H318 | Gene names | CRIM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cysteine-rich motor neuron 1 protein precursor (CRIM-1) (Cysteine-rich repeat-containing protein S52). | |||||
|
CYLN2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 30) | NC score | 0.028404 (rank : 88) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1004 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UDT6, O14527, O43611 | Gene names | CYLN2, KIAA0291, WBSCR4, WSCR4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoplasmic linker protein 2 (Cytoplasmic linker protein 115) (CLIP- 115) (Williams-Beuren syndrome chromosome region 4 protein). | |||||
|
LAMB1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 31) | NC score | 0.019203 (rank : 96) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P02469 | Gene names | Lamb1-1, Lamb-1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-1 chain precursor (Laminin B1 chain). | |||||
|
BPA1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 32) | NC score | 0.017448 (rank : 100) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q91ZU6, Q91ZU7 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 1/2/3/4 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
CYLN2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 33) | NC score | 0.024704 (rank : 91) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1002 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9Z0H8, Q7TSI9, Q8CHU1, Q9EP81 | Gene names | Cyln2, Kiaa0291 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoplasmic linker protein 2 (Cytoplasmic linker protein 115) (CLIP- 115). | |||||
|
KIF17_MOUSE
|
||||||
| θ value | 3.0926 (rank : 34) | NC score | 0.010850 (rank : 105) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q99PW8 | Gene names | Kif17 | |||
|
Domain Architecture |
|
|||||
| Description | Kinesin-like protein KIF17 (MmKIF17). | |||||
|
LRC45_HUMAN
|
||||||
| θ value | 3.0926 (rank : 35) | NC score | 0.021999 (rank : 92) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 933 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q96CN5 | Gene names | LRRC45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 45. | |||||
|
AMNLS_MOUSE
|
||||||
| θ value | 4.03905 (rank : 36) | NC score | 0.018835 (rank : 97) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99JB7, Q5I0U1, Q99MP9 | Gene names | Amn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amnionless protein precursor. | |||||
|
CT174_HUMAN
|
||||||
| θ value | 4.03905 (rank : 37) | NC score | 0.019342 (rank : 95) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5JPB2, Q5TDR4, Q8TCP0 | Gene names | C20orf174 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C20orf174. | |||||
|
KTN1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 38) | NC score | 0.032538 (rank : 81) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1324 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q61595, Q8BG49, Q8BHF4, Q8BHM8, Q8C9Y5, Q8CG51, Q8CG52, Q8CG53, Q8CG54, Q8CG55, Q8CG56, Q8CG57, Q8CG58, Q8CG59, Q8CG60, Q8CG61, Q8CG62, Q8CG63 | Gene names | Ktn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinectin. | |||||
|
SDCG8_HUMAN
|
||||||
| θ value | 4.03905 (rank : 39) | NC score | 0.020861 (rank : 93) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1002 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q86SQ7, O60527, Q3ZCR6, Q8N5F2, Q9P0F1 | Gene names | SDCCAG8, CCCAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serologically defined colon cancer antigen 8 (Centrosomal colon cancer autoantigen protein) (hCCCAP) (Antigen NY-CO-8). | |||||
|
TSP2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 40) | NC score | 0.005023 (rank : 107) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P35442 | Gene names | THBS2, TSP2 | |||
|
Domain Architecture |

|
|||||
| Description | Thrombospondin-2 precursor. | |||||
|
CRDL2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 41) | NC score | 0.013150 (rank : 102) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8VEA6, Q925I3 | Gene names | Chrdl2, Chl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chordin-like protein 2 precursor. | |||||
|
GOGA5_MOUSE
|
||||||
| θ value | 5.27518 (rank : 42) | NC score | 0.018024 (rank : 98) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1434 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9QYE6, O88317, Q3TGE7, Q3U6S5, Q3UUF9 | Gene names | Golga5, Retii | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 5 (Golgin-84) (Sumiko protein) (Ret-II protein). | |||||
|
CEP57_MOUSE
|
||||||
| θ value | 6.88961 (rank : 43) | NC score | 0.051049 (rank : 72) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 580 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8CEE0, Q6ZQJ3, Q7TN18, Q80X65, Q810F2, Q9D4J4, Q9D5S4, Q9D5W5 | Gene names | Cep57, Kiaa0092, Tsp57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 57 kDa (Cep57 protein) (Testis-specific protein 57) (Translokin). | |||||
|
MYH1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 44) | NC score | 0.028917 (rank : 87) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1478 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P12882, Q9Y622 | Gene names | MYH1 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-1 (Myosin heavy chain, skeletal muscle, adult 1) (Myosin heavy chain IIx/d) (MyHC-IIx/d). | |||||
|
CCD62_HUMAN
|
||||||
| θ value | 8.99809 (rank : 45) | NC score | 0.020365 (rank : 94) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 631 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q6P9F0, Q6ZVF2, Q86VJ0, Q9BYZ5 | Gene names | CCDC62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 60 (Aaa-protein) (Protein TSP- NY). | |||||
|
CEP57_HUMAN
|
||||||
| θ value | 8.99809 (rank : 46) | NC score | 0.045349 (rank : 75) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 576 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q86XR8, Q14704, Q8IXP0, Q9BVF9 | Gene names | CEP57, KIAA0092, TSP57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 57 kDa (Cep57 protein) (Testis-specific protein 57) (Translokin) (FGF2-interacting protein). | |||||
|
CNOT4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 47) | NC score | 0.012983 (rank : 103) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O95628, O95339, O95627, Q8IYM7, Q8NCL0, Q9NPQ1, Q9NZN6 | Gene names | CNOT4, NOT4 | |||
|
Domain Architecture |
|
|||||
| Description | CCR4-NOT transcription complex subunit 4 (EC 6.3.2.-) (E3 ubiquitin protein ligase CNOT4) (CCR4-associated factor 4) (Potential transcriptional repressor NOT4Hp). | |||||
|
CNOT4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 48) | NC score | 0.012195 (rank : 104) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BT14, Q8CCR4, Q9CV74, Q9Z1D0 | Gene names | Cnot4, Not4 | |||
|
Domain Architecture |
|
|||||
| Description | CCR4-NOT transcription complex subunit 4 (EC 6.3.2.-) (E3 ubiquitin protein ligase CNOT4) (CCR4-associated factor 4) (Potential transcriptional repressor NOT4Hp). | |||||
|
KTN1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 49) | NC score | 0.041337 (rank : 77) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1197 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q86UP2, Q13999, Q14707, Q15387, Q86W57 | Gene names | KTN1, CG1, KIAA0004 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinectin (Kinesin receptor) (CG-1 antigen). | |||||
|
LRRF2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 50) | NC score | 0.037500 (rank : 79) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 791 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9Y608, Q68CV3, Q9NXH5 | Gene names | LRRFIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat flightless-interacting protein 2 (LRR FLII- interacting protein 2). | |||||
|
MYH2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 51) | NC score | 0.029483 (rank : 84) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1491 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9UKX2, Q14322, Q16229 | Gene names | MYH2, MYHSA2 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, skeletal muscle, adult 2 (Myosin heavy chain IIa) (MyHC-IIa). | |||||
|
TACC3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 52) | NC score | 0.029393 (rank : 85) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9Y6A5, Q9UMQ1 | Gene names | TACC3, ERIC1 | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 3 (ERIC-1). | |||||
|
BFAR_HUMAN
|
||||||
| θ value | θ > 10 (rank : 53) | NC score | 0.079063 (rank : 40) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NZS9 | Gene names | BFAR, BAR, RNF47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bifunctional apoptosis regulator (RING finger protein 47). | |||||
|
BFAR_MOUSE
|
||||||
| θ value | θ > 10 (rank : 54) | NC score | 0.075596 (rank : 43) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8R079, Q8C1A7, Q9CXY3 | Gene names | Bfar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bifunctional apoptosis regulator. | |||||
|
BRCA1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 55) | NC score | 0.072736 (rank : 45) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P38398 | Gene names | BRCA1, RNF53 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 1 susceptibility protein (RING finger protein 53). | |||||
|
BRCA1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 56) | NC score | 0.070454 (rank : 47) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 436 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P48754, Q60957, Q60983 | Gene names | Brca1 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 1 susceptibility protein homolog. | |||||
|
FBX28_HUMAN
|
||||||
| θ value | θ > 10 (rank : 57) | NC score | 0.054582 (rank : 64) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NVF7, O75070 | Gene names | FBXO28, KIAA0483 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 28. | |||||
|
LNX1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 58) | NC score | 0.085755 (rank : 36) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 379 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8TBB1, Q8N4C2, Q96MJ7, Q9BY20 | Gene names | LNX1, LNX | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin ligase LNX (EC 6.3.2.-) (Numb-binding protein 1) (Ligand of Numb-protein X 1). | |||||
|
LNX1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 59) | NC score | 0.086081 (rank : 34) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 377 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O70263, O70264, Q8BRI8, Q8CFR3 | Gene names | Lnx1, Lnx | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin ligase LNX (EC 6.3.2.-) (Numb-binding protein 1) (Ligand of Numb-binding protein 1) (Ligand of Numb-protein X 1). | |||||
|
LNX2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 60) | NC score | 0.067093 (rank : 50) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 389 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8N448, Q5W0P0, Q6ZMH2, Q96SH4 | Gene names | LNX2, PDZRN1 | |||
|
Domain Architecture |

|
|||||
| Description | Ligand of Numb-protein X 2 (Numb-binding protein 2) (PDZ domain- containing RING finger protein 1). | |||||
|
LNX2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 61) | NC score | 0.071081 (rank : 46) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 407 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q91XL2, Q8CBE1 | Gene names | Lnx2 | |||
|
Domain Architecture |

|
|||||
| Description | Ligand of Numb-protein X 2 (Numb-binding protein 2). | |||||
|
MAT1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 62) | NC score | 0.052164 (rank : 69) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P51949, Q3TZP0, Q9D8A0, Q9D8D2 | Gene names | Mnat1, Mat1 | |||
|
Domain Architecture |
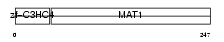
|
|||||
| Description | CDK-activating kinase assembly factor MAT1 (RING finger protein MAT1) (Menage a trois) (CDK7/cyclin H assembly factor) (p36) (p35). | |||||
|
MEP1B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 63) | NC score | 0.177911 (rank : 18) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q16820, Q670J1 | Gene names | MEP1B | |||
|
Domain Architecture |

|
|||||
| Description | Meprin A subunit beta precursor (EC 3.4.24.18) (Endopeptidase-2) (N- benzoyl-L-tyrosyl-P-amino-benzoic acid hydrolase subunit beta) (PABA peptide hydrolase) (PPH beta). | |||||
|
PEX10_HUMAN
|
||||||
| θ value | θ > 10 (rank : 64) | NC score | 0.067398 (rank : 49) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O60683, Q5T095, Q9BW90 | Gene names | PEX10, RNF69 | |||
|
Domain Architecture |
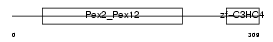
|
|||||
| Description | Peroxisome assembly protein 10 (Peroxin-10) (Peroxisome biogenesis factor 10) (RING finger protein 69). | |||||
|
PZRN3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 65) | NC score | 0.142229 (rank : 27) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 461 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UPQ7, Q8N2N7, Q96CC2, Q9NSQ2 | Gene names | PDZRN3, KIAA1095, LNX3, SEMCAP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing RING finger protein 3 (Ligand of Numb-protein X 3) (Semaphorin cytoplasmic domain-associated protein 3) (SEMACAP3 protein). | |||||
|
PZRN3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 66) | NC score | 0.145768 (rank : 26) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q69ZS0, Q91Z03, Q9QY54, Q9QY55 | Gene names | Pdzrn3, Kiaa1095, Semcap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing RING finger protein 3 (Semaphorin cytoplasmic domain-associated protein 3) (SEMACAP3 protein). | |||||
|
PZRN4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 67) | NC score | 0.109057 (rank : 29) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 438 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q6ZMN7, Q6N052, Q8IUU1, Q9NTP7 | Gene names | PDZRN4, LNX4, SEMCAP3L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing RING finger protein 4 (Ligand of Numb-protein X 4) (SEMACAP3-like protein). | |||||
|
R113A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 68) | NC score | 0.085896 (rank : 35) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O15541 | Gene names | RNF113A, RNF113, ZNF183 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 113A (Zinc finger protein 183). | |||||
|
R113B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 69) | NC score | 0.076382 (rank : 42) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8IZP6, Q8WWF9, Q96QY9 | Gene names | RNF113B, RNF161, ZNF183L1 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 113B (Zinc finger protein 183-like 1). | |||||
|
RAG1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 70) | NC score | 0.106917 (rank : 31) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P15918, Q8NER2 | Gene names | RAG1, RNF74 | |||
|
Domain Architecture |

|
|||||
| Description | V(D)J recombination-activating protein 1 (RAG-1) (RING finger protein 74). | |||||
|
RAG1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 71) | NC score | 0.107481 (rank : 30) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P15919 | Gene names | Rag1, Rag-1 | |||
|
Domain Architecture |

|
|||||
| Description | V(D)J recombination-activating protein 1 (RAG-1). | |||||
|
RFWD2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 72) | NC score | 0.051264 (rank : 71) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 518 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8NHY2, Q6H103, Q9H6L7 | Gene names | RFWD2, COP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger and WD repeat domain protein 2 (EC 6.3.2.-) (Ubiquitin- protein ligase COP1) (Constitutive photomorphogenesis protein 1 homolog) (hCOP1). | |||||
|
RFWD2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 73) | NC score | 0.050708 (rank : 73) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 535 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9R1A8 | Gene names | Rfwd2, Cop1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger and WD repeat domain protein 2 (EC 6.3.2.-) (Ubiquitin- protein ligase COP1) (Constitutive photomorphogenesis protein 1 homolog) (mCOP1). | |||||
|
RING1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 74) | NC score | 0.094625 (rank : 32) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q06587, Q5JP96, Q5SQW2, Q86V19 | Gene names | RING1, RNF1 | |||
|
Domain Architecture |
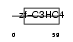
|
|||||
| Description | Polycomb complex protein RING1 (RING finger protein 1). | |||||
|
RING1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 75) | NC score | 0.092794 (rank : 33) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O35730, Q3U242, Q3U333, Q4FK33, Q63ZX8, Q921Z8 | Gene names | Ring1, Ring1A, Rnf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polycomb complex protein RING1 (RING finger protein 1) (Transcription repressor Ring1A). | |||||
|
RING2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 76) | NC score | 0.085243 (rank : 37) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q99496, Q5TEN1, Q5TEN2 | Gene names | RNF2, BAP1, DING, HIPI3, RING1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein RING2 (EC 6.3.2.-) (RING finger protein 2) (RING finger protein 1B) (RING1b) (RING finger protein BAP-1) (DinG protein) (Huntingtin-interacting protein 2-interacting protein 3) (HIP2-interacting protein 3). | |||||
|
RING2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 77) | NC score | 0.085234 (rank : 38) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9CQJ4, O35699, O35729, Q4FJV5, Q8C1X8 | Gene names | Rnf2, DinG, Ring1b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein RING2 (EC 6.3.2.-) (RING finger protein 2) (RING finger protein 1B) (RING1b). | |||||
|
RN125_HUMAN
|
||||||
| θ value | θ > 10 (rank : 78) | NC score | 0.168248 (rank : 19) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q96EQ8, Q9NX39 | Gene names | RNF125 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 125 (EC 6.3.2.-) (T-cell RING activation protein 1) (TRAC-1). | |||||
|
RN125_MOUSE
|
||||||
| θ value | θ > 10 (rank : 79) | NC score | 0.141470 (rank : 28) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9D9R0, Q52KL4, Q8C7F2 | Gene names | Rnf125 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 125 (EC 6.3.2.-). | |||||
|
RN166_HUMAN
|
||||||
| θ value | θ > 10 (rank : 80) | NC score | 0.193012 (rank : 15) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q96A37, Q96DM0 | Gene names | RNF166 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 166. | |||||
|
RN166_MOUSE
|
||||||
| θ value | θ > 10 (rank : 81) | NC score | 0.187379 (rank : 16) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q3U9F6 | Gene names | Rnf166 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 166. | |||||
|
RNF41_HUMAN
|
||||||
| θ value | θ > 10 (rank : 82) | NC score | 0.163969 (rank : 20) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 226 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9H4P4, O75598 | Gene names | RNF41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 41. | |||||
|
RNF41_MOUSE
|
||||||
| θ value | θ > 10 (rank : 83) | NC score | 0.163047 (rank : 21) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 224 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8BH75, Q8BGJ2, Q8BMC9, Q8VEA2, Q9CRK6, Q9D568 | Gene names | Rnf41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 41. | |||||
|
RNF4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 84) | NC score | 0.059834 (rank : 57) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P78317 | Gene names | RNF4 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 4. | |||||
|
RNF4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 85) | NC score | 0.062658 (rank : 53) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9QZS2, O35941, Q541Z6 | Gene names | Rnf4 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 4. | |||||
|
TRI12_MOUSE
|
||||||
| θ value | θ > 10 (rank : 86) | NC score | 0.059849 (rank : 56) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q99PQ1, Q9D704 | Gene names | Trim12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 12. | |||||
|
TRI22_HUMAN
|
||||||
| θ value | θ > 10 (rank : 87) | NC score | 0.052772 (rank : 68) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 396 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8IYM9, Q15521 | Gene names | TRIM22, RNF94, STAF50 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 22 (RING finger protein 94) (50 kDa-stimulated trans-acting factor) (Staf-50). | |||||
|
TRI25_HUMAN
|
||||||
| θ value | θ > 10 (rank : 88) | NC score | 0.056450 (rank : 61) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q14258 | Gene names | TRIM25, EFP, RNF147, ZNF147 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 25 (Zinc finger protein 147) (Estrogen-responsive finger protein) (Efp) (RING finger protein 147). | |||||
|
TRI25_MOUSE
|
||||||
| θ value | θ > 10 (rank : 89) | NC score | 0.064635 (rank : 52) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 446 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q61510 | Gene names | Trim25, Efp, Zfp147, Znf147 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 25 (Zinc finger protein 147) (Estrogen-responsive finger protein) (Efp). | |||||
|
TRI26_HUMAN
|
||||||
| θ value | θ > 10 (rank : 90) | NC score | 0.054977 (rank : 62) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q12899 | Gene names | TRIM26, RNF95, ZNF173 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 26 (Zinc finger protein 173) (Acid finger protein) (AFP) (RING finger protein 95). | |||||
|
TRI26_MOUSE
|
||||||
| θ value | θ > 10 (rank : 91) | NC score | 0.053732 (rank : 67) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 339 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q99PN3, Q8C9E9, Q8JZT7, Q99PN2 | Gene names | Trim26, Znf173 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 26 (Zinc finger protein 173). | |||||
|
TRI31_HUMAN
|
||||||
| θ value | θ > 10 (rank : 92) | NC score | 0.073184 (rank : 44) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 299 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9BZY9, Q53H52, Q5RI37, Q5SRJ7, Q5SRJ8, Q5SS28, Q96AK4, Q96AP8, Q99579, Q9BZY8 | Gene names | TRIM31, C6orf13, RNF | |||
|
Domain Architecture |
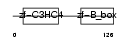
|
|||||
| Description | Tripartite motif-containing protein 31. | |||||
|
TRI47_HUMAN
|
||||||
| θ value | θ > 10 (rank : 93) | NC score | 0.059139 (rank : 58) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q96LD4, Q96GU5, Q9BRN7 | Gene names | TRIM47, GOA, RNF100 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 47 (Gene overexpressed in astrocytoma protein) (RING finger protein 100). | |||||
|
TRI47_MOUSE
|
||||||
| θ value | θ > 10 (rank : 94) | NC score | 0.068561 (rank : 48) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 295 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8C0E3, Q6P249, Q811J7, Q8BVZ8, Q8R1K0, Q8R3Y1 | Gene names | Trim47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 47. | |||||
|
TRI52_HUMAN
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.051511 (rank : 70) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96A61 | Gene names | TRIM52, RNF102 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 52 (RING finger protein 102). | |||||
|
TRI56_HUMAN
|
||||||
| θ value | θ > 10 (rank : 96) | NC score | 0.053763 (rank : 66) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 339 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9BRZ2, Q86VT6, Q8N2H8, Q8NAC0, Q9H031 | Gene names | TRIM56, RNF109 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 56 (RING finger protein 109). | |||||
|
TRI65_HUMAN
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.056644 (rank : 60) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6PJ69, Q4G0F0, Q6DKJ6, Q9BRP6 | Gene names | TRIM65 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 65. | |||||
|
TRI73_HUMAN
|
||||||
| θ value | θ > 10 (rank : 98) | NC score | 0.057161 (rank : 59) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q86UV7, Q8N0S3 | Gene names | TRIM73, TRIM50B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 73 (Tripartite motif-containing protein 50B). | |||||
|
TRIM8_HUMAN
|
||||||
| θ value | θ > 10 (rank : 99) | NC score | 0.054196 (rank : 65) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9BZR9, Q9C028 | Gene names | TRIM8, GERP, RNF27 | |||
|
Domain Architecture |
|
|||||
| Description | Tripartite motif-containing protein 8 (RING finger protein 27) (Glioblastoma-expressed RING finger protein). | |||||
|
TRIM8_MOUSE
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.054956 (rank : 63) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q99PJ2, Q8C508, Q8C700, Q8CGI2, Q99PQ4 | Gene names | Trim8, Gerp, Rnf27 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 8 (RING finger protein 27) (Glioblastoma-expressed RING finger protein). | |||||
|
UHRF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.061102 (rank : 54) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 272 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96T88, Q8J022, Q9H6S6, Q9P115, Q9P1U7 | Gene names | UHRF1, ICBP90, NP95, RNF106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 1 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 1) (Inverted CCAAT box-binding protein of 90 kDa) (Transcription factor ICBP90) (Nuclear zinc finger protein Np95) (Nuclear protein 95) (HuNp95) (RING finger protein 106). | |||||
|
UHRF1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.060379 (rank : 55) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8VDF2, Q8C6F1, Q8VIA1, Q9Z1H6 | Gene names | Uhrf1, Np95 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 1 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 1) (Nuclear zinc finger protein Np95) (Nuclear protein 95). | |||||
|
UHRF2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.079759 (rank : 39) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96PU4, Q5VYR1, Q5VYR3, Q659C8, Q8TAG7 | Gene names | UHRF2, NIRF, RNF107 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 2 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 2) (Np95/ICBP90-like RING finger protein) (Np95-like RING finger protein) (Nuclear zinc finger protein Np97) (RING finger protein 107). | |||||
|
UHRF2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.078002 (rank : 41) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q7TMI3, Q8BG56, Q8BJP6, Q8BY30, Q8K1I5 | Gene names | Uhrf2, Nirf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 2 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 2) (Np95-like ring finger protein) (Nuclear zinc finger protein Np97) (NIRF). | |||||
|
ZN179_HUMAN
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.050037 (rank : 74) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9ULX5, O60633 | Gene names | ZNF179, BFP, RNF112 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 179 (Brain finger protein) (RING finger protein 112). | |||||
|
ZN313_HUMAN
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.146543 (rank : 25) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y508, Q6N0B0 | Gene names | ZNF313, RNF114, ZNF228 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger protein 313 (RING finger protein 114). | |||||
|
ZN313_MOUSE
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.153558 (rank : 22) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9ET26, Q3UFU8, Q8K5A2 | Gene names | Znf313, Zfp228, Zfp313, Znf228 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger protein 313. | |||||
|
TRAF1_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P39428 | Gene names | Traf1 | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 1. | |||||
|
TRAF1_HUMAN
|
||||||
| NC score | 0.993061 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q13077, Q8NF13 | Gene names | TRAF1, EBI6 | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 1 (Epstein-Barr virus-induced protein 6). | |||||
|
TRAF2_HUMAN
|
||||||
| NC score | 0.867370 (rank : 3) | θ value | 2.31901e-72 (rank : 4) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q12933, Q96NT2 | Gene names | TRAF2, TRAP3 | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 2 (Tumor necrosis factor type 2 receptor-associated protein 3). | |||||
|
TRAF2_MOUSE
|
||||||
| NC score | 0.861801 (rank : 4) | θ value | 1.22964e-73 (rank : 3) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P39429, O54896 | Gene names | Traf2 | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 2. | |||||
|
TRAF5_HUMAN
|
||||||
| NC score | 0.813615 (rank : 5) | θ value | 3.974e-56 (rank : 8) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 396 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O00463 | Gene names | TRAF5, RNF84 | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 5 (RING finger protein 84). | |||||
|
TRAF5_MOUSE
|
||||||
| NC score | 0.811958 (rank : 6) | θ value | 8.00737e-57 (rank : 7) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 355 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P70191, Q61480 | Gene names | Traf5 | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 5. | |||||
|
TRAF3_MOUSE
|
||||||
| NC score | 0.809459 (rank : 7) | θ value | 6.99855e-61 (rank : 5) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 476 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q60803, Q62380 | Gene names | Traf3, Craf1, Trafamn | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 3 (CD40 receptor-associated factor 1) (CRAF1) (TRAFAMN). | |||||
|
TRAF3_HUMAN
|
||||||
| NC score | 0.808557 (rank : 8) | θ value | 3.47335e-60 (rank : 6) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 463 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q13114, Q12990, Q13076, Q13947, Q6AZX1, Q9UNL1 | Gene names | TRAF3, CAP1, CRAF1 | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 3 (CD40 receptor-associated factor 1) (CRAF1) (CD40-binding protein) (CD40BP) (LMP1-associated protein) (LAP1) (CAP-1). | |||||
|
TRAF4_HUMAN
|
||||||
| NC score | 0.715634 (rank : 9) | θ value | 9.2256e-29 (rank : 9) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 373 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9BUZ4, O75615, Q14848, Q2PJN8 | Gene names | TRAF4, CART1, MLN62, RNF83 | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 4 (Cysteine-rich domain associated with RING and Traf domains protein 1) (Malignant 62) (RING finger protein 83). | |||||
|
TRAF4_MOUSE
|
||||||
| NC score | 0.715420 (rank : 10) | θ value | 2.96777e-27 (rank : 10) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 338 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q61382 | Gene names | Traf4, Cart1 | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 4 (Cysteine-rich motif associated to RING and Traf domains protein 1). | |||||
|
TRAF6_HUMAN
|
||||||
| NC score | 0.700788 (rank : 11) | θ value | 7.56453e-23 (rank : 11) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9Y4K3, Q8NEH5 | Gene names | TRAF6, RNF85 | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 6 (Interleukin 1 signal transducer) (RING finger protein 85). | |||||
|
TRAF6_MOUSE
|
||||||
| NC score | 0.698528 (rank : 12) | θ value | 5.42112e-21 (rank : 12) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P70196, Q8BLV2 | Gene names | Traf6 | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 6. | |||||
|
MEP1B_MOUSE
|
||||||
| NC score | 0.218789 (rank : 13) | θ value | 0.0330416 (rank : 16) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q61847 | Gene names | Mep1b, Mep-1b | |||
|
Domain Architecture |

|
|||||
| Description | Meprin A subunit beta precursor (EC 3.4.24.18) (Endopeptidase-2). | |||||
|
MEP1A_MOUSE
|
||||||
| NC score | 0.216853 (rank : 14) | θ value | 0.000158464 (rank : 13) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P28825 | Gene names | Mep1a | |||
|
Domain Architecture |

|
|||||
| Description | Meprin A subunit alpha precursor (EC 3.4.24.18) (Endopeptidase-2) (MEP-1). | |||||
|
RN166_HUMAN
|
||||||
| NC score | 0.193012 (rank : 15) | θ value | θ > 10 (rank : 80) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q96A37, Q96DM0 | Gene names | RNF166 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 166. | |||||
|
RN166_MOUSE
|
||||||
| NC score | 0.187379 (rank : 16) | θ value | θ > 10 (rank : 81) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q3U9F6 | Gene names | Rnf166 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 166. | |||||
|
MEP1A_HUMAN
|
||||||
| NC score | 0.183769 (rank : 17) | θ value | 0.0193708 (rank : 15) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q16819 | Gene names | MEP1A | |||
|
Domain Architecture |

|
|||||
| Description | Meprin A subunit alpha precursor (EC 3.4.24.18) (Endopeptidase-2) (N- benzoyl-L-tyrosyl-P-amino-benzoic acid hydrolase subunit alpha) (PABA peptide hydrolase) (PPH alpha). | |||||
|
MEP1B_HUMAN
|
||||||
| NC score | 0.177911 (rank : 18) | θ value | θ > 10 (rank : 63) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q16820, Q670J1 | Gene names | MEP1B | |||
|
Domain Architecture |

|
|||||
| Description | Meprin A subunit beta precursor (EC 3.4.24.18) (Endopeptidase-2) (N- benzoyl-L-tyrosyl-P-amino-benzoic acid hydrolase subunit beta) (PABA peptide hydrolase) (PPH beta). | |||||
|
RN125_HUMAN
|
||||||
| NC score | 0.168248 (rank : 19) | θ value | θ > 10 (rank : 78) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q96EQ8, Q9NX39 | Gene names | RNF125 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 125 (EC 6.3.2.-) (T-cell RING activation protein 1) (TRAC-1). | |||||
|
RNF41_HUMAN
|
||||||
| NC score | 0.163969 (rank : 20) | θ value | θ > 10 (rank : 82) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 226 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9H4P4, O75598 | Gene names | RNF41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 41. | |||||
|
RNF41_MOUSE
|
||||||
| NC score | 0.163047 (rank : 21) | θ value | θ > 10 (rank : 83) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 224 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8BH75, Q8BGJ2, Q8BMC9, Q8VEA2, Q9CRK6, Q9D568 | Gene names | Rnf41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 41. | |||||
|
ZN313_MOUSE
|
||||||
| NC score | 0.153558 (rank : 22) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9ET26, Q3UFU8, Q8K5A2 | Gene names | Znf313, Zfp228, Zfp313, Znf228 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 313. | |||||
|
TRAF7_MOUSE
|
||||||
| NC score | 0.150452 (rank : 23) | θ value | 0.813845 (rank : 22) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 470 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q922B6, Q99JV3 | Gene names | Traf7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin protein ligase TRAF7 (EC 6.3.2.-) (TNF receptor- associated factor 7). | |||||
|
TRAF7_HUMAN
|
||||||
| NC score | 0.148445 (rank : 24) | θ value | 1.06291 (rank : 23) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 482 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q6Q0C0, Q9H073 | Gene names | TRAF7, RFWD1, RNF119 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin protein ligase TRAF7 (EC 6.3.2.-) (TNF receptor- associated factor 7) (RING finger and WD repeat domain protein 1) (RING finger protein 119). | |||||
|
ZN313_HUMAN
|
||||||
| NC score | 0.146543 (rank : 25) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y508, Q6N0B0 | Gene names | ZNF313, RNF114, ZNF228 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 313 (RING finger protein 114). | |||||
|
PZRN3_MOUSE
|
||||||
| NC score | 0.145768 (rank : 26) | θ value | θ > 10 (rank : 66) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q69ZS0, Q91Z03, Q9QY54, Q9QY55 | Gene names | Pdzrn3, Kiaa1095, Semcap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing RING finger protein 3 (Semaphorin cytoplasmic domain-associated protein 3) (SEMACAP3 protein). | |||||
|
PZRN3_HUMAN
|
||||||
| NC score | 0.142229 (rank : 27) | θ value | θ > 10 (rank : 65) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 461 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UPQ7, Q8N2N7, Q96CC2, Q9NSQ2 | Gene names | PDZRN3, KIAA1095, LNX3, SEMCAP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing RING finger protein 3 (Ligand of Numb-protein X 3) (Semaphorin cytoplasmic domain-associated protein 3) (SEMACAP3 protein). | |||||
|
RN125_MOUSE
|
||||||
| NC score | 0.141470 (rank : 28) | θ value | θ > 10 (rank : 79) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9D9R0, Q52KL4, Q8C7F2 | Gene names | Rnf125 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 125 (EC 6.3.2.-). | |||||
|
PZRN4_HUMAN
|
||||||
| NC score | 0.109057 (rank : 29) | θ value | θ > 10 (rank : 67) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 438 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q6ZMN7, Q6N052, Q8IUU1, Q9NTP7 | Gene names | PDZRN4, LNX4, SEMCAP3L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing RING finger protein 4 (Ligand of Numb-protein X 4) (SEMACAP3-like protein). | |||||
|
RAG1_MOUSE
|
||||||
| NC score | 0.107481 (rank : 30) | θ value | θ > 10 (rank : 71) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P15919 | Gene names | Rag1, Rag-1 | |||
|
Domain Architecture |

|
|||||
| Description | V(D)J recombination-activating protein 1 (RAG-1). | |||||
|
RAG1_HUMAN
|
||||||
| NC score | 0.106917 (rank : 31) | θ value | θ > 10 (rank : 70) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P15918, Q8NER2 | Gene names | RAG1, RNF74 | |||
|
Domain Architecture |

|
|||||
| Description | V(D)J recombination-activating protein 1 (RAG-1) (RING finger protein 74). | |||||
|
RING1_HUMAN
|
||||||
| NC score | 0.094625 (rank : 32) | θ value | θ > 10 (rank : 74) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q06587, Q5JP96, Q5SQW2, Q86V19 | Gene names | RING1, RNF1 | |||
|
Domain Architecture |

|
|||||
| Description | Polycomb complex protein RING1 (RING finger protein 1). | |||||
|
RING1_MOUSE
|
||||||
| NC score | 0.092794 (rank : 33) | θ value | θ > 10 (rank : 75) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O35730, Q3U242, Q3U333, Q4FK33, Q63ZX8, Q921Z8 | Gene names | Ring1, Ring1A, Rnf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polycomb complex protein RING1 (RING finger protein 1) (Transcription repressor Ring1A). | |||||
|
LNX1_MOUSE
|
||||||
| NC score | 0.086081 (rank : 34) | θ value | θ > 10 (rank : 59) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 377 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O70263, O70264, Q8BRI8, Q8CFR3 | Gene names | Lnx1, Lnx | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin ligase LNX (EC 6.3.2.-) (Numb-binding protein 1) (Ligand of Numb-binding protein 1) (Ligand of Numb-protein X 1). | |||||
|
R113A_HUMAN
|
||||||
| NC score | 0.085896 (rank : 35) | θ value | θ > 10 (rank : 68) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O15541 | Gene names | RNF113A, RNF113, ZNF183 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 113A (Zinc finger protein 183). | |||||
|
LNX1_HUMAN
|
||||||
| NC score | 0.085755 (rank : 36) | θ value | θ > 10 (rank : 58) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 379 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8TBB1, Q8N4C2, Q96MJ7, Q9BY20 | Gene names | LNX1, LNX | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin ligase LNX (EC 6.3.2.-) (Numb-binding protein 1) (Ligand of Numb-protein X 1). | |||||
|
RING2_HUMAN
|
||||||
| NC score | 0.085243 (rank : 37) | θ value | θ > 10 (rank : 76) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q99496, Q5TEN1, Q5TEN2 | Gene names | RNF2, BAP1, DING, HIPI3, RING1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein RING2 (EC 6.3.2.-) (RING finger protein 2) (RING finger protein 1B) (RING1b) (RING finger protein BAP-1) (DinG protein) (Huntingtin-interacting protein 2-interacting protein 3) (HIP2-interacting protein 3). | |||||
|
RING2_MOUSE
|
||||||
| NC score | 0.085234 (rank : 38) | θ value | θ > 10 (rank : 77) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9CQJ4, O35699, O35729, Q4FJV5, Q8C1X8 | Gene names | Rnf2, DinG, Ring1b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein RING2 (EC 6.3.2.-) (RING finger protein 2) (RING finger protein 1B) (RING1b). | |||||
|
UHRF2_HUMAN
|
||||||
| NC score | 0.079759 (rank : 39) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96PU4, Q5VYR1, Q5VYR3, Q659C8, Q8TAG7 | Gene names | UHRF2, NIRF, RNF107 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 2 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 2) (Np95/ICBP90-like RING finger protein) (Np95-like RING finger protein) (Nuclear zinc finger protein Np97) (RING finger protein 107). | |||||
|
BFAR_HUMAN
|
||||||
| NC score | 0.079063 (rank : 40) | θ value | θ > 10 (rank : 53) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NZS9 | Gene names | BFAR, BAR, RNF47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bifunctional apoptosis regulator (RING finger protein 47). | |||||
|
UHRF2_MOUSE
|
||||||
| NC score | 0.078002 (rank : 41) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q7TMI3, Q8BG56, Q8BJP6, Q8BY30, Q8K1I5 | Gene names | Uhrf2, Nirf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 2 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 2) (Np95-like ring finger protein) (Nuclear zinc finger protein Np97) (NIRF). | |||||
|
R113B_HUMAN
|
||||||
| NC score | 0.076382 (rank : 42) | θ value | θ > 10 (rank : 69) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8IZP6, Q8WWF9, Q96QY9 | Gene names | RNF113B, RNF161, ZNF183L1 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 113B (Zinc finger protein 183-like 1). | |||||
|
BFAR_MOUSE
|
||||||
| NC score | 0.075596 (rank : 43) | θ value | θ > 10 (rank : 54) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8R079, Q8C1A7, Q9CXY3 | Gene names | Bfar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bifunctional apoptosis regulator. | |||||
|
TRI31_HUMAN
|
||||||
| NC score | 0.073184 (rank : 44) | θ value | θ > 10 (rank : 92) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 299 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9BZY9, Q53H52, Q5RI37, Q5SRJ7, Q5SRJ8, Q5SS28, Q96AK4, Q96AP8, Q99579, Q9BZY8 | Gene names | TRIM31, C6orf13, RNF | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 31. | |||||
|
BRCA1_HUMAN
|
||||||
| NC score | 0.072736 (rank : 45) | θ value | θ > 10 (rank : 55) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P38398 | Gene names | BRCA1, RNF53 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 1 susceptibility protein (RING finger protein 53). | |||||
|
LNX2_MOUSE
|
||||||
| NC score | 0.071081 (rank : 46) | θ value | θ > 10 (rank : 61) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 407 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q91XL2, Q8CBE1 | Gene names | Lnx2 | |||
|
Domain Architecture |

|
|||||
| Description | Ligand of Numb-protein X 2 (Numb-binding protein 2). | |||||
|
BRCA1_MOUSE
|
||||||
| NC score | 0.070454 (rank : 47) | θ value | θ > 10 (rank : 56) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 436 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P48754, Q60957, Q60983 | Gene names | Brca1 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 1 susceptibility protein homolog. | |||||
|
TRI47_MOUSE
|
||||||
| NC score | 0.068561 (rank : 48) | θ value | θ > 10 (rank : 94) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 295 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8C0E3, Q6P249, Q811J7, Q8BVZ8, Q8R1K0, Q8R3Y1 | Gene names | Trim47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 47. | |||||
|
PEX10_HUMAN
|
||||||
| NC score | 0.067398 (rank : 49) | θ value | θ > 10 (rank : 64) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O60683, Q5T095, Q9BW90 | Gene names | PEX10, RNF69 | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisome assembly protein 10 (Peroxin-10) (Peroxisome biogenesis factor 10) (RING finger protein 69). | |||||
|
LNX2_HUMAN
|
||||||
| NC score | 0.067093 (rank : 50) | θ value | θ > 10 (rank : 60) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 389 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8N448, Q5W0P0, Q6ZMH2, Q96SH4 | Gene names | LNX2, PDZRN1 | |||
|
Domain Architecture |

|
|||||
| Description | Ligand of Numb-protein X 2 (Numb-binding protein 2) (PDZ domain- containing RING finger protein 1). | |||||
|
TACC3_MOUSE
|
||||||
| NC score | 0.065176 (rank : 51) | θ value | 0.0148317 (rank : 14) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 607 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9JJ11, Q9WVK9 | Gene names | Tacc3, Aint | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 3 (ARNT-interacting protein). | |||||
|
TRI25_MOUSE
|
||||||
| NC score | 0.064635 (rank : 52) | θ value | θ > 10 (rank : 89) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 446 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q61510 | Gene names | Trim25, Efp, Zfp147, Znf147 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 25 (Zinc finger protein 147) (Estrogen-responsive finger protein) (Efp). | |||||
|
RNF4_MOUSE
|
||||||
| NC score | 0.062658 (rank : 53) | θ value | θ > 10 (rank : 85) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9QZS2, O35941, Q541Z6 | Gene names | Rnf4 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 4. | |||||
|
UHRF1_HUMAN
|
||||||
| NC score | 0.061102 (rank : 54) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 272 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96T88, Q8J022, Q9H6S6, Q9P115, Q9P1U7 | Gene names | UHRF1, ICBP90, NP95, RNF106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 1 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 1) (Inverted CCAAT box-binding protein of 90 kDa) (Transcription factor ICBP90) (Nuclear zinc finger protein Np95) (Nuclear protein 95) (HuNp95) (RING finger protein 106). | |||||
|
UHRF1_MOUSE
|
||||||
| NC score | 0.060379 (rank : 55) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8VDF2, Q8C6F1, Q8VIA1, Q9Z1H6 | Gene names | Uhrf1, Np95 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 1 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 1) (Nuclear zinc finger protein Np95) (Nuclear protein 95). | |||||
|
TRI12_MOUSE
|
||||||
| NC score | 0.059849 (rank : 56) | θ value | θ > 10 (rank : 86) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q99PQ1, Q9D704 | Gene names | Trim12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 12. | |||||
|
RNF4_HUMAN
|
||||||
| NC score | 0.059834 (rank : 57) | θ value | θ > 10 (rank : 84) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P78317 | Gene names | RNF4 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 4. | |||||
|
TRI47_HUMAN
|
||||||
| NC score | 0.059139 (rank : 58) | θ value | θ > 10 (rank : 93) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q96LD4, Q96GU5, Q9BRN7 | Gene names | TRIM47, GOA, RNF100 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 47 (Gene overexpressed in astrocytoma protein) (RING finger protein 100). | |||||
|
TRI73_HUMAN
|
||||||
| NC score | 0.057161 (rank : 59) | θ value | θ > 10 (rank : 98) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q86UV7, Q8N0S3 | Gene names | TRIM73, TRIM50B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 73 (Tripartite motif-containing protein 50B). | |||||
|
TRI65_HUMAN
|
||||||
| NC score | 0.056644 (rank : 60) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6PJ69, Q4G0F0, Q6DKJ6, Q9BRP6 | Gene names | TRIM65 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 65. | |||||
|
TRI25_HUMAN
|
||||||
| NC score | 0.056450 (rank : 61) | θ value | θ > 10 (rank : 88) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q14258 | Gene names | TRIM25, EFP, RNF147, ZNF147 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 25 (Zinc finger protein 147) (Estrogen-responsive finger protein) (Efp) (RING finger protein 147). | |||||
|
TRI26_HUMAN
|
||||||
| NC score | 0.054977 (rank : 62) | θ value | θ > 10 (rank : 90) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q12899 | Gene names | TRIM26, RNF95, ZNF173 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 26 (Zinc finger protein 173) (Acid finger protein) (AFP) (RING finger protein 95). | |||||
|
TRIM8_MOUSE
|
||||||
| NC score | 0.054956 (rank : 63) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q99PJ2, Q8C508, Q8C700, Q8CGI2, Q99PQ4 | Gene names | Trim8, Gerp, Rnf27 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 8 (RING finger protein 27) (Glioblastoma-expressed RING finger protein). | |||||
|
FBX28_HUMAN
|
||||||
| NC score | 0.054582 (rank : 64) | θ value | θ > 10 (rank : 57) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NVF7, O75070 | Gene names | FBXO28, KIAA0483 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 28. | |||||
|
TRIM8_HUMAN
|
||||||
| NC score | 0.054196 (rank : 65) | θ value | θ > 10 (rank : 99) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9BZR9, Q9C028 | Gene names | TRIM8, GERP, RNF27 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 8 (RING finger protein 27) (Glioblastoma-expressed RING finger protein). | |||||
|
TRI56_HUMAN
|
||||||
| NC score | 0.053763 (rank : 66) | θ value | θ > 10 (rank : 96) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 339 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9BRZ2, Q86VT6, Q8N2H8, Q8NAC0, Q9H031 | Gene names | TRIM56, RNF109 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 56 (RING finger protein 109). | |||||
|
TRI26_MOUSE
|
||||||
| NC score | 0.053732 (rank : 67) | θ value | θ > 10 (rank : 91) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 339 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q99PN3, Q8C9E9, Q8JZT7, Q99PN2 | Gene names | Trim26, Znf173 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 26 (Zinc finger protein 173). | |||||
|
TRI22_HUMAN
|
||||||
| NC score | 0.052772 (rank : 68) | θ value | θ > 10 (rank : 87) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 396 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8IYM9, Q15521 | Gene names | TRIM22, RNF94, STAF50 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 22 (RING finger protein 94) (50 kDa-stimulated trans-acting factor) (Staf-50). | |||||
|
MAT1_MOUSE
|
||||||
| NC score | 0.052164 (rank : 69) | θ value | θ > 10 (rank : 62) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P51949, Q3TZP0, Q9D8A0, Q9D8D2 | Gene names | Mnat1, Mat1 | |||
|
Domain Architecture |

|
|||||
| Description | CDK-activating kinase assembly factor MAT1 (RING finger protein MAT1) (Menage a trois) (CDK7/cyclin H assembly factor) (p36) (p35). | |||||
|
TRI52_HUMAN
|
||||||
| NC score | 0.051511 (rank : 70) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96A61 | Gene names | TRIM52, RNF102 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 52 (RING finger protein 102). | |||||
|
RFWD2_HUMAN
|
||||||
| NC score | 0.051264 (rank : 71) | θ value | θ > 10 (rank : 72) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 518 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8NHY2, Q6H103, Q9H6L7 | Gene names | RFWD2, COP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger and WD repeat domain protein 2 (EC 6.3.2.-) (Ubiquitin- protein ligase COP1) (Constitutive photomorphogenesis protein 1 homolog) (hCOP1). | |||||
|
CEP57_MOUSE
|
||||||
| NC score | 0.051049 (rank : 72) | θ value | 6.88961 (rank : 43) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 580 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8CEE0, Q6ZQJ3, Q7TN18, Q80X65, Q810F2, Q9D4J4, Q9D5S4, Q9D5W5 | Gene names | Cep57, Kiaa0092, Tsp57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 57 kDa (Cep57 protein) (Testis-specific protein 57) (Translokin). | |||||
|
RFWD2_MOUSE
|
||||||
| NC score | 0.050708 (rank : 73) | θ value | θ > 10 (rank : 73) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 535 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9R1A8 | Gene names | Rfwd2, Cop1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger and WD repeat domain protein 2 (EC 6.3.2.-) (Ubiquitin- protein ligase COP1) (Constitutive photomorphogenesis protein 1 homolog) (mCOP1). | |||||
|
ZN179_HUMAN
|
||||||
| NC score | 0.050037 (rank : 74) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9ULX5, O60633 | Gene names | ZNF179, BFP, RNF112 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 179 (Brain finger protein) (RING finger protein 112). | |||||
|
CEP57_HUMAN
|
||||||
| NC score | 0.045349 (rank : 75) | θ value | 8.99809 (rank : 46) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 576 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q86XR8, Q14704, Q8IXP0, Q9BVF9 | Gene names | CEP57, KIAA0092, TSP57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 57 kDa (Cep57 protein) (Testis-specific protein 57) (Translokin) (FGF2-interacting protein). | |||||
|
CE290_HUMAN
|
||||||
| NC score | 0.043614 (rank : 76) | θ value | 0.365318 (rank : 19) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1553 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O15078, Q1PSK5, Q66GS8, Q9H2G6, Q9H6Q7, Q9H8I0 | Gene names | CEP290, KIAA0373, NPHP6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein Cep290 (Nephrocystin-6) (Tumor antigen se2-2). | |||||
|
KTN1_HUMAN
|
||||||
| NC score | 0.041337 (rank : 77) | θ value | 8.99809 (rank : 49) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1197 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q86UP2, Q13999, Q14707, Q15387, Q86W57 | Gene names | KTN1, CG1, KIAA0004 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinectin (Kinesin receptor) (CG-1 antigen). | |||||
|
CKAP4_MOUSE
|
||||||
| NC score | 0.041284 (rank : 78) | θ value | 0.279714 (rank : 18) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 596 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8BMK4, Q8BTK8, Q8R3F2 | Gene names | Ckap4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-associated protein 4 (63 kDa membrane protein) (p63). | |||||
|
LRRF2_HUMAN
|
||||||
| NC score | 0.037500 (rank : 79) | θ value | 8.99809 (rank : 50) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 791 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9Y608, Q68CV3, Q9NXH5 | Gene names | LRRFIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat flightless-interacting protein 2 (LRR FLII- interacting protein 2). | |||||
|
TTC17_HUMAN
|
||||||
| NC score | 0.034660 (rank : 80) | θ value | 1.06291 (rank : 24) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96AE7 | Gene names | TTC17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tetratricopeptide repeat protein 17 (TPR repeat protein 17). | |||||
|
KTN1_MOUSE
|
||||||
| NC score | 0.032538 (rank : 81) | θ value | 4.03905 (rank : 38) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1324 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q61595, Q8BG49, Q8BHF4, Q8BHM8, Q8C9Y5, Q8CG51, Q8CG52, Q8CG53, Q8CG54, Q8CG55, Q8CG56, Q8CG57, Q8CG58, Q8CG59, Q8CG60, Q8CG61, Q8CG62, Q8CG63 | Gene names | Ktn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinectin. | |||||
|
CP250_HUMAN
|
||||||
| NC score | 0.032405 (rank : 82) | θ value | 0.0961366 (rank : 17) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1805 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9BV73, O14812, O60588, Q9H450 | Gene names | CEP250, CEP2, CNAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1). | |||||
|
MYH7_HUMAN
|
||||||
| NC score | 0.030926 (rank : 83) | θ value | 1.38821 (rank : 27) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P12883, Q14836, Q14837, Q14904, Q16579, Q92679, Q9H1D5, Q9UMM8 | Gene names | MYH7, MYHCB | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle beta isoform (MyHC-beta). | |||||
|
MYH2_HUMAN
|
||||||
| NC score | 0.029483 (rank : 84) | θ value | 8.99809 (rank : 51) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1491 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9UKX2, Q14322, Q16229 | Gene names | MYH2, MYHSA2 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, skeletal muscle, adult 2 (Myosin heavy chain IIa) (MyHC-IIa). | |||||
|
TACC3_HUMAN
|
||||||
| NC score | 0.029393 (rank : 85) | θ value | 8.99809 (rank : 52) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9Y6A5, Q9UMQ1 | Gene names | TACC3, ERIC1 | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 3 (ERIC-1). | |||||
|
MYH6_MOUSE
|
||||||
| NC score | 0.028923 (rank : 86) | θ value | 1.38821 (rank : 26) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1501 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q02566, Q64258, Q64738 | Gene names | Myh6, Myhca | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
MYH1_HUMAN
|
||||||
| NC score | 0.028917 (rank : 87) | θ value | 6.88961 (rank : 44) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1478 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P12882, Q9Y622 | Gene names | MYH1 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-1 (Myosin heavy chain, skeletal muscle, adult 1) (Myosin heavy chain IIx/d) (MyHC-IIx/d). | |||||
|
CYLN2_HUMAN
|
||||||
| NC score | 0.028404 (rank : 88) | θ value | 2.36792 (rank : 30) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1004 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UDT6, O14527, O43611 | Gene names | CYLN2, KIAA0291, WBSCR4, WSCR4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoplasmic linker protein 2 (Cytoplasmic linker protein 115) (CLIP- 115) (Williams-Beuren syndrome chromosome region 4 protein). | |||||
|
RUFY2_HUMAN
|
||||||
| NC score | 0.026629 (rank : 89) | θ value | 0.365318 (rank : 20) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1208 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8WXA3, Q96P51, Q9P1Z1 | Gene names | RUFY2, KIAA1537, RABIP4R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and FYVE domain-containing protein 2 (Rab4-interacting protein related). | |||||
|
PLEC1_HUMAN
|
||||||
| NC score | 0.026537 (rank : 90) | θ value | 0.47712 (rank : 21) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1326 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q15149, Q15148, Q16640 | Gene names | PLEC1 | |||
|
Domain Architecture |

|
|||||
| Description | Plectin-1 (PLTN) (PCN) (Hemidesmosomal protein 1) (HD1). | |||||
|
CYLN2_MOUSE
|
||||||
| NC score | 0.024704 (rank : 91) | θ value | 3.0926 (rank : 33) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1002 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9Z0H8, Q7TSI9, Q8CHU1, Q9EP81 | Gene names | Cyln2, Kiaa0291 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoplasmic linker protein 2 (Cytoplasmic linker protein 115) (CLIP- 115). | |||||
|
LRC45_HUMAN
|
||||||
| NC score | 0.021999 (rank : 92) | θ value | 3.0926 (rank : 35) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 933 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q96CN5 | Gene names | LRRC45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 45. | |||||
|
SDCG8_HUMAN
|
||||||
| NC score | 0.020861 (rank : 93) | θ value | 4.03905 (rank : 39) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1002 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q86SQ7, O60527, Q3ZCR6, Q8N5F2, Q9P0F1 | Gene names | SDCCAG8, CCCAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serologically defined colon cancer antigen 8 (Centrosomal colon cancer autoantigen protein) (hCCCAP) (Antigen NY-CO-8). | |||||
|
CCD62_HUMAN
|
||||||
| NC score | 0.020365 (rank : 94) | θ value | 8.99809 (rank : 45) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 631 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q6P9F0, Q6ZVF2, Q86VJ0, Q9BYZ5 | Gene names | CCDC62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 60 (Aaa-protein) (Protein TSP- NY). | |||||
|
CT174_HUMAN
|
||||||
| NC score | 0.019342 (rank : 95) | θ value | 4.03905 (rank : 37) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5JPB2, Q5TDR4, Q8TCP0 | Gene names | C20orf174 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C20orf174. | |||||
|
LAMB1_MOUSE
|
||||||
| NC score | 0.019203 (rank : 96) | θ value | 2.36792 (rank : 31) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P02469 | Gene names | Lamb1-1, Lamb-1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-1 chain precursor (Laminin B1 chain). | |||||
|
AMNLS_MOUSE
|
||||||
| NC score | 0.018835 (rank : 97) | θ value | 4.03905 (rank : 36) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99JB7, Q5I0U1, Q99MP9 | Gene names | Amn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amnionless protein precursor. | |||||
|
GOGA5_MOUSE
|
||||||
| NC score | 0.018024 (rank : 98) | θ value | 5.27518 (rank : 42) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1434 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9QYE6, O88317, Q3TGE7, Q3U6S5, Q3UUF9 | Gene names | Golga5, Retii | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 5 (Golgin-84) (Sumiko protein) (Ret-II protein). | |||||
|
CRIM1_HUMAN
|
||||||
| NC score | 0.017556 (rank : 99) | θ value | 2.36792 (rank : 29) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NZV1, Q59GH0, Q7LCQ5, Q9H318 | Gene names | CRIM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cysteine-rich motor neuron 1 protein precursor (CRIM-1) (Cysteine-rich repeat-containing protein S52). | |||||
|
BPA1_MOUSE
|
||||||
| NC score | 0.017448 (rank : 100) | θ value | 3.0926 (rank : 32) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q91ZU6, Q91ZU7 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 1/2/3/4 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
LAMB1_HUMAN
|
||||||
| NC score | 0.016955 (rank : 101) | θ value | 1.38821 (rank : 25) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1018 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P07942 | Gene names | LAMB1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-1 chain precursor (Laminin B1 chain). | |||||
|
CRDL2_MOUSE
|
||||||
| NC score | 0.013150 (rank : 102) | θ value | 5.27518 (rank : 41) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8VEA6, Q925I3 | Gene names | Chrdl2, Chl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chordin-like protein 2 precursor. | |||||
|
CNOT4_HUMAN
|
||||||
| NC score | 0.012983 (rank : 103) | θ value | 8.99809 (rank : 47) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O95628, O95339, O95627, Q8IYM7, Q8NCL0, Q9NPQ1, Q9NZN6 | Gene names | CNOT4, NOT4 | |||
|
Domain Architecture |

|
|||||
| Description | CCR4-NOT transcription complex subunit 4 (EC 6.3.2.-) (E3 ubiquitin protein ligase CNOT4) (CCR4-associated factor 4) (Potential transcriptional repressor NOT4Hp). | |||||
|
CNOT4_MOUSE
|
||||||
| NC score | 0.012195 (rank : 104) | θ value | 8.99809 (rank : 48) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BT14, Q8CCR4, Q9CV74, Q9Z1D0 | Gene names | Cnot4, Not4 | |||
|
Domain Architecture |

|
|||||
| Description | CCR4-NOT transcription complex subunit 4 (EC 6.3.2.-) (E3 ubiquitin protein ligase CNOT4) (CCR4-associated factor 4) (Potential transcriptional repressor NOT4Hp). | |||||
|
KIF17_MOUSE
|
||||||
| NC score | 0.010850 (rank : 105) | θ value | 3.0926 (rank : 34) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q99PW8 | Gene names | Kif17 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF17 (MmKIF17). | |||||
|
RAB6C_HUMAN
|
||||||
| NC score | 0.006483 (rank : 106) | θ value | 1.81305 (rank : 28) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H0N0, Q9P128 | Gene names | RAB6C, WTH3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras-related protein Rab-6C (Rab6-like protein WTH3). | |||||
|
TSP2_HUMAN
|
||||||
| NC score | 0.005023 (rank : 107) | θ value | 4.03905 (rank : 40) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P35442 | Gene names | THBS2, TSP2 | |||
|
Domain Architecture |

|
|||||
| Description | Thrombospondin-2 precursor. | |||||