Please be patient as the page loads
|
SV2C_HUMAN
|
||||||
| SwissProt Accessions | Q496J9, Q496K1, Q9UPU8 | Gene names | SV2C, KIAA1054 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptic vesicle glycoprotein 2C. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
SV2A_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.978076 (rank : 5) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q7L0J3, O94841, Q5QNX8, Q7Z3L6, Q8NBJ6, Q9BVZ9 | Gene names | SV2A, KIAA0736 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptic vesicle glycoprotein 2A. | |||||
|
SV2A_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.977920 (rank : 6) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9JIS5, Q80TT0, Q8R0R5 | Gene names | Sv2a, Kiaa0736, Sv2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptic vesicle glycoprotein 2A (Synaptic vesicle protein 2A) (Synaptic vesicle protein 2) (Calcium regulator SV2A). | |||||
|
SV2B_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.986002 (rank : 3) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q7L1I2, O94840, Q6IAR8 | Gene names | SV2B, KIAA0735 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptic vesicle glycoprotein 2B. | |||||
|
SV2B_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.984073 (rank : 4) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8BG39, Q80TT1, Q9ES95 | Gene names | Sv2b, Kiaa0735 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptic vesicle glycoprotein 2B (Synaptic vesicle protein 2B). | |||||
|
SV2C_HUMAN
|
||||||
| θ value | 0 (rank : 5) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 124 | |
| SwissProt Accessions | Q496J9, Q496K1, Q9UPU8 | Gene names | SV2C, KIAA1054 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptic vesicle glycoprotein 2C. | |||||
|
SV2C_MOUSE
|
||||||
| θ value | 0 (rank : 6) | NC score | 0.994150 (rank : 2) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q69ZS6 | Gene names | Sv2c, Kiaa1054 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptic vesicle glycoprotein 2C (Synaptic vesicle protein 2C). | |||||
|
S22A5_HUMAN
|
||||||
| θ value | 5.60996e-18 (rank : 7) | NC score | 0.582638 (rank : 7) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O76082, Q6ZQZ8, Q96EH6 | Gene names | SLC22A5, OCTN2 | |||
|
Domain Architecture |

|
|||||
| Description | Organic cation/carnitine transporter 2 (Solute carrier family 22 member 5) (High-affinity sodium-dependent carnitine cotransporter). | |||||
|
S22A5_MOUSE
|
||||||
| θ value | 1.80466e-16 (rank : 8) | NC score | 0.567247 (rank : 9) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9Z0E8 | Gene names | Slc22a5, Octn2 | |||
|
Domain Architecture |

|
|||||
| Description | Organic cation/carnitine transporter 2 (Solute carrier family 22 member 5) (High-affinity sodium-dependent carnitine cotransporter). | |||||
|
OCTN3_MOUSE
|
||||||
| θ value | 5.8054e-15 (rank : 9) | NC score | 0.567410 (rank : 8) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9WTN6, Q5SWV0 | Gene names | Slc22a21, Octn3, Slc22a9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Organic cation/carnitine transporter 3 (Solute carrier family 22 member 21) (Solute carrier family 22 member 9). | |||||
|
S22A4_MOUSE
|
||||||
| θ value | 7.09661e-13 (rank : 10) | NC score | 0.550843 (rank : 10) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9Z306 | Gene names | Slc22a4, Octn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Organic cation/carnitine transporter 1 (Solute carrier family 22 member 4). | |||||
|
GTR8_HUMAN
|
||||||
| θ value | 1.74796e-11 (rank : 11) | NC score | 0.486586 (rank : 18) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9NY64, Q9NSC4 | Gene names | SLC2A8, GLUT8, GLUTX1 | |||
|
Domain Architecture |
|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 8 (Glucose transporter type 8) (Glucose transporter type X1). | |||||
|
S22A4_HUMAN
|
||||||
| θ value | 1.74796e-11 (rank : 12) | NC score | 0.545393 (rank : 11) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9H015, O14546 | Gene names | SLC22A4, ETT, OCTN1, UT2H | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Organic cation/carnitine transporter 1 (Solute carrier family 22 member 4) (Ergothioneine transporter) (ET transporter). | |||||
|
GTR8_MOUSE
|
||||||
| θ value | 6.64225e-11 (rank : 13) | NC score | 0.485434 (rank : 19) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9JIF3, Q9JJP4, Q9JJZ0 | Gene names | Slc2a8, Glut8, GlutX1 | |||
|
Domain Architecture |
|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 8 (Glucose transporter type 8) (Glucose transporter type X1). | |||||
|
ORCT3_HUMAN
|
||||||
| θ value | 1.133e-10 (rank : 14) | NC score | 0.533823 (rank : 12) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9Y226, Q8IYG1 | Gene names | SLC22A13, ORCTL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Organic-cation transporter-like 3 (Solute carrier family 22 member 13). | |||||
|
ORCT3_MOUSE
|
||||||
| θ value | 1.25267e-09 (rank : 15) | NC score | 0.530375 (rank : 13) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q6A4L0, Q8VC89 | Gene names | Slc22a13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Organic cation transporter-like 3 (Solute carrier family 22 member 13). | |||||
|
GTR6_HUMAN
|
||||||
| θ value | 2.13673e-09 (rank : 16) | NC score | 0.467924 (rank : 21) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9UGQ3, Q5SXD7 | Gene names | SLC2A6, GLUT9 | |||
|
Domain Architecture |

|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 6 (Glucose transporter type 6) (Glucose transporter type 9). | |||||
|
S22A3_MOUSE
|
||||||
| θ value | 3.64472e-09 (rank : 17) | NC score | 0.527220 (rank : 14) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9WTW5, Q9R209 | Gene names | Slc22a3, Oct3 | |||
|
Domain Architecture |

|
|||||
| Description | Organic cation transporter 3 (Solute carrier family 22 member 3). | |||||
|
GTR2_MOUSE
|
||||||
| θ value | 8.11959e-09 (rank : 18) | NC score | 0.380809 (rank : 32) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P14246, Q3TNA5, Q9DBA7 | Gene names | Slc2a2, Glut-2, Glut2 | |||
|
Domain Architecture |
|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 2 (Glucose transporter type 2, liver). | |||||
|
ORCT4_HUMAN
|
||||||
| θ value | 1.80886e-08 (rank : 19) | NC score | 0.487238 (rank : 17) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9Y267, Q6DJT3 | Gene names | SLC22A14, ORCTL4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Organic-cation transporter-like 4 (Solute carrier family 22 member 14). | |||||
|
OAT6_MOUSE
|
||||||
| θ value | 2.36244e-08 (rank : 20) | NC score | 0.501943 (rank : 16) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q80UJ1 | Gene names | Slc22a20, Gm963, Oat6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Solute carrier family 22 member 20 (Organic anion transporter 6). | |||||
|
S22A3_HUMAN
|
||||||
| θ value | 3.08544e-08 (rank : 21) | NC score | 0.504229 (rank : 15) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O75751, Q9UP02 | Gene names | SLC22A3, EMTH | |||
|
Domain Architecture |

|
|||||
| Description | Organic cation transporter 3 (Extraneuronal monoamine transporter) (EMT) (Solute carrier family 22 member 3). | |||||
|
OAT4_HUMAN
|
||||||
| θ value | 4.45536e-07 (rank : 22) | NC score | 0.470331 (rank : 20) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9NSA0, Q53GR2, Q6ZP72, Q8NBU4 | Gene names | SLC22A11, OAT4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Solute carrier family 22 member 11 (Organic anion transporter 4). | |||||
|
MYCT_HUMAN
|
||||||
| θ value | 5.81887e-07 (rank : 23) | NC score | 0.464363 (rank : 22) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q96QE2 | Gene names | SLC2A13 | |||
|
Domain Architecture |

|
|||||
| Description | Proton myo-inositol cotransporter (H(+)-myo-inositol cotransporter) (Hmit). | |||||
|
GTR3_HUMAN
|
||||||
| θ value | 2.88788e-06 (rank : 24) | NC score | 0.392237 (rank : 26) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P11169, Q6I9U2, Q9UG15 | Gene names | SLC2A3, GLUT3 | |||
|
Domain Architecture |
|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 3 (Glucose transporter type 3, brain). | |||||
|
GTR11_HUMAN
|
||||||
| θ value | 1.43324e-05 (rank : 25) | NC score | 0.392623 (rank : 25) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9BYW1 | Gene names | SLC2A11, GLUT10, GLUT11 | |||
|
Domain Architecture |
|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 11 (Glucose transporter type 11) (Glucose transporter type 10). | |||||
|
GTR3_MOUSE
|
||||||
| θ value | 1.87187e-05 (rank : 26) | NC score | 0.387711 (rank : 30) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P32037 | Gene names | Slc2a3, Glut-3, Glut3 | |||
|
Domain Architecture |
|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 3 (Glucose transporter type 3, brain). | |||||
|
GTR1_MOUSE
|
||||||
| θ value | 2.44474e-05 (rank : 27) | NC score | 0.388126 (rank : 29) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P17809, Q61608 | Gene names | Slc2a1, Glut-1, Glut1 | |||
|
Domain Architecture |
|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 1 (Glucose transporter type 1, erythrocyte/brain) (GT1). | |||||
|
GTR1_HUMAN
|
||||||
| θ value | 4.1701e-05 (rank : 28) | NC score | 0.389181 (rank : 28) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P11166, O75535 | Gene names | SLC2A1, GLUT1 | |||
|
Domain Architecture |
|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 1 (Glucose transporter type 1, erythrocyte/brain) (HepG2 glucose transporter). | |||||
|
GTR5_MOUSE
|
||||||
| θ value | 7.1131e-05 (rank : 29) | NC score | 0.390961 (rank : 27) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9WV38 | Gene names | Slc2a5, Glut5 | |||
|
Domain Architecture |
|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 5 (Glucose transporter type 5, small intestine) (Fructose transporter). | |||||
|
GTR5_HUMAN
|
||||||
| θ value | 9.29e-05 (rank : 30) | NC score | 0.359425 (rank : 34) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P22732, Q14770 | Gene names | SLC2A5, GLUT5 | |||
|
Domain Architecture |
|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 5 (Glucose transporter type 5, small intestine) (Fructose transporter). | |||||
|
GTR14_HUMAN
|
||||||
| θ value | 0.000121331 (rank : 31) | NC score | 0.371508 (rank : 33) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8TDB8, Q6UY84, Q8TDB9 | Gene names | SLC2A14, GLUT14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 14 (Glucose transporter type 14). | |||||
|
GTR9_HUMAN
|
||||||
| θ value | 0.00020696 (rank : 32) | NC score | 0.384747 (rank : 31) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9NRM0 | Gene names | SLC2A9, GLUT9 | |||
|
Domain Architecture |

|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 9 (Glucose transporter type 9). | |||||
|
GTR4_MOUSE
|
||||||
| θ value | 0.000270298 (rank : 33) | NC score | 0.321606 (rank : 35) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P14142, Q9JJN9 | Gene names | Slc2a4, Glut-4, Glut4 | |||
|
Domain Architecture |
|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 4 (Glucose transporter type 4, insulin-responsive) (GT2). | |||||
|
GTR2_HUMAN
|
||||||
| θ value | 0.00134147 (rank : 34) | NC score | 0.316866 (rank : 37) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P11168 | Gene names | SLC2A2, GLUT2 | |||
|
Domain Architecture |

|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 2 (Glucose transporter type 2, liver). | |||||
|
GTR4_HUMAN
|
||||||
| θ value | 0.00175202 (rank : 35) | NC score | 0.318092 (rank : 36) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P14672 | Gene names | SLC2A4, GLUT4 | |||
|
Domain Architecture |
|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 4 (Glucose transporter type 4, insulin-responsive). | |||||
|
OAT5_HUMAN
|
||||||
| θ value | 0.00228821 (rank : 36) | NC score | 0.419208 (rank : 23) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q63ZE4 | Gene names | SLC22A10, OAT5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Solute carrier family 22 member 10 (Organic anion transporter 5). | |||||
|
CT059_MOUSE
|
||||||
| θ value | 0.00869519 (rank : 37) | NC score | 0.062909 (rank : 48) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8VCL5 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative transporter C20orf59 homolog. | |||||
|
S22A9_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 38) | NC score | 0.399767 (rank : 24) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8IVM8, Q8TEC0, Q8WYN7 | Gene names | SLC22A9, OAT4, UST3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Solute carrier family 22 member 9 (Organic anion transporter 4). | |||||
|
AN32A_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 39) | NC score | 0.066880 (rank : 44) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 254 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O35381, P97437 | Gene names | Anp32a, Anp32, Lanp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member A (Potent heat-stable protein phosphatase 2A inhibitor I1PP2A) (Acidic nuclear phosphoprotein pp32) (Leucine-rich acidic nuclear protein). | |||||
|
AN32C_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 40) | NC score | 0.066880 (rank : 45) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 254 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q64G17 | Gene names | Anp32c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member C. | |||||
|
HNRPC_HUMAN
|
||||||
| θ value | 0.163984 (rank : 41) | NC score | 0.039847 (rank : 56) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P07910, P22628, Q53EX2, Q59FD3, Q5FWE8, Q86SF8, Q86U45, Q96HK7, Q96HM4, Q96IY5, Q9BTS3 | Gene names | HNRPC | |||
|
Domain Architecture |
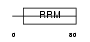
|
|||||
| Description | Heterogeneous nuclear ribonucleoproteins C1/C2 (hnRNP C1 / hnRNP C2). | |||||
|
HNRCL_HUMAN
|
||||||
| θ value | 0.21417 (rank : 42) | NC score | 0.049197 (rank : 55) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O60812 | Gene names | HNRPCL1 | |||
|
Domain Architecture |
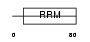
|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein C-like 1 (hnRNP core protein C-like 1). | |||||
|
MYT1_MOUSE
|
||||||
| θ value | 0.21417 (rank : 43) | NC score | 0.026612 (rank : 63) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8CFC2, O08995, Q8CFH1 | Gene names | Myt1, Kiaa0835, Nzf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1 (MyT1) (Neural zinc finger factor 2) (NZF-2). | |||||
|
AN32B_HUMAN
|
||||||
| θ value | 0.279714 (rank : 44) | NC score | 0.071391 (rank : 40) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q92688, O00655, P78458, P78459 | Gene names | ANP32B, APRIL, PHAPI2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member B (PHAPI2 protein) (Silver-stainable protein SSP29) (Acidic protein rich in leucines). | |||||
|
AN32E_MOUSE
|
||||||
| θ value | 0.365318 (rank : 45) | NC score | 0.063769 (rank : 47) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P97822, Q3TH89, Q8BPF8, Q8C2L4, Q8C7Q8, Q9CZD2 | Gene names | Anp32e, Cpd1 | |||
|
Domain Architecture |
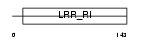
|
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member E (LANP- like protein) (LANP-L) (Cerebellar postnatal development protein 1). | |||||
|
CCAR1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 46) | NC score | 0.027934 (rank : 60) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 444 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8IX12, Q32NE3, Q5VUP6, Q6PIZ0, Q6X935, Q9H8N4, Q9NVA7, Q9NVQ0, Q9NWM6 | Gene names | CCAR1, CARP1, DIS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle and apoptosis regulator protein 1 (Cell cycle and apoptosis regulatory protein 1) (CARP-1) (Death inducer with SAP domain). | |||||
|
VMAT2_MOUSE
|
||||||
| θ value | 0.47712 (rank : 47) | NC score | 0.060181 (rank : 50) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8BRU6, Q8CC55 | Gene names | Slc18a2, Vmat2 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptic vesicular amine transporter (Monoamine transporter) (Vesicular amine transporter 2) (VAT2) (Solute carrier family 18 member 2). | |||||
|
AN32B_MOUSE
|
||||||
| θ value | 0.62314 (rank : 48) | NC score | 0.069515 (rank : 41) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9EST5 | Gene names | Anp32b, Pal31 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member B (Proliferation-related acidic leucine-rich protein PAL31). | |||||
|
KI21B_MOUSE
|
||||||
| θ value | 0.813845 (rank : 49) | NC score | 0.011876 (rank : 91) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 997 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9QXL1, P97424 | Gene names | Kif21b, Kif6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 21B (Kinesin-like protein KIF6). | |||||
|
NFM_MOUSE
|
||||||
| θ value | 0.813845 (rank : 50) | NC score | 0.007094 (rank : 113) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1159 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P08553, Q61961 | Gene names | Nef3, Nefm, Nfm | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M). | |||||
|
TPR_HUMAN
|
||||||
| θ value | 0.813845 (rank : 51) | NC score | 0.007880 (rank : 109) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
DAXX_MOUSE
|
||||||
| θ value | 1.06291 (rank : 52) | NC score | 0.026997 (rank : 61) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O35613, Q9QWT8, Q9QWV3 | Gene names | Daxx | |||
|
Domain Architecture |

|
|||||
| Description | Death domain-associated protein 6 (Daxx). | |||||
|
GTR10_HUMAN
|
||||||
| θ value | 1.06291 (rank : 53) | NC score | 0.267450 (rank : 39) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O95528, Q9H4I6 | Gene names | SLC2A10, GLUT10 | |||
|
Domain Architecture |

|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 10 (Glucose transporter type 10). | |||||
|
SET_MOUSE
|
||||||
| θ value | 1.06291 (rank : 54) | NC score | 0.024881 (rank : 65) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9EQU5, Q9CY82, Q9D0A9, Q9Z181 | Gene names | Set | |||
|
Domain Architecture |
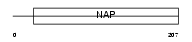
|
|||||
| Description | Protein SET (Phosphatase 2A inhibitor I2PP2A) (I-2PP2A) (Template- activating factor I) (TAF-I). | |||||
|
CENPB_MOUSE
|
||||||
| θ value | 1.38821 (rank : 55) | NC score | 0.053452 (rank : 51) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P27790 | Gene names | Cenpb, Cenp-b | |||
|
Domain Architecture |

|
|||||
| Description | Major centromere autoantigen B (Centromere protein B) (CENP-B). | |||||
|
IF2P_HUMAN
|
||||||
| θ value | 1.38821 (rank : 56) | NC score | 0.013048 (rank : 90) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 889 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O60841, O95805, Q9UF81, Q9UMN7 | Gene names | EIF5B, IF2, KIAA0741 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 5B (eIF-5B) (Translation initiation factor IF-2). | |||||
|
NFM_HUMAN
|
||||||
| θ value | 1.38821 (rank : 57) | NC score | 0.005194 (rank : 119) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1328 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P07197 | Gene names | NEF3, NEFM, NFM | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M) (Neurofilament 3). | |||||
|
UBF1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 58) | NC score | 0.029650 (rank : 59) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P25976 | Gene names | Ubtf, Tcfubf, Ubf-1, Ubf1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar transcription factor 1 (Upstream-binding factor 1) (UBF-1). | |||||
|
VMAT2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 59) | NC score | 0.052731 (rank : 52) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q05940, Q15876, Q9H3P6 | Gene names | SLC18A2, SVMT, VMAT2 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptic vesicular amine transporter (Monoamine transporter) (Vesicular amine transporter 2) (VAT2) (Solute carrier family 18 member 2). | |||||
|
CCAR1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 60) | NC score | 0.024325 (rank : 66) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8CH18, Q6AXC9, Q6PAR2, Q80XE4, Q8BJY0, Q8BVN2, Q8CGG1, Q9CSR5 | Gene names | Ccar1, Carp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle and apoptosis regulator protein 1 (Cell cycle and apoptosis regulatory protein 1) (CARP-1). | |||||
|
CYLC2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 61) | NC score | 0.025620 (rank : 64) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q14093 | Gene names | CYLC2, CYL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-2 (Cylicin II) (Multiple-band polypeptide II). | |||||
|
KI21A_MOUSE
|
||||||
| θ value | 1.81305 (rank : 62) | NC score | 0.010767 (rank : 98) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1317 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9QXL2, Q6P5H1, Q6ZPJ8, Q8BWZ9, Q8BXF1 | Gene names | Kif21a, Kiaa1708 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 21A. | |||||
|
NRDC_MOUSE
|
||||||
| θ value | 1.81305 (rank : 63) | NC score | 0.015703 (rank : 81) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BHG1 | Gene names | Nrd1 | |||
|
Domain Architecture |

|
|||||
| Description | Nardilysin precursor (EC 3.4.24.61) (N-arginine dibasic convertase) (NRD convertase) (NRD-C). | |||||
|
PHF14_HUMAN
|
||||||
| θ value | 1.81305 (rank : 64) | NC score | 0.013999 (rank : 85) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O94880 | Gene names | PHF14, KIAA0783 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 14. | |||||
|
PTN13_MOUSE
|
||||||
| θ value | 1.81305 (rank : 65) | NC score | 0.005121 (rank : 120) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 426 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q64512, Q61494, Q62135, Q64499 | Gene names | Ptpn13, Ptp14 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 13 (EC 3.1.3.48) (Protein tyrosine phosphatase PTP-BL) (Protein-tyrosine phosphatase RIP) (protein tyrosine phosphatase DPZPTP) (PTP36). | |||||
|
SRCH_HUMAN
|
||||||
| θ value | 1.81305 (rank : 66) | NC score | 0.021757 (rank : 71) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P23327, Q504Y6 | Gene names | HRC, HCP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcoplasmic reticulum histidine-rich calcium-binding protein precursor. | |||||
|
UBF1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 67) | NC score | 0.021138 (rank : 74) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 415 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P17480 | Gene names | UBTF, UBF, UBF1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar transcription factor 1 (Upstream-binding factor 1) (UBF-1) (Autoantigen NOR-90). | |||||
|
AN32A_HUMAN
|
||||||
| θ value | 2.36792 (rank : 68) | NC score | 0.064900 (rank : 46) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P39687 | Gene names | ANP32A, C15orf1, LANP, MAPM, PHAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member A (Potent heat-stable protein phosphatase 2A inhibitor I1PP2A) (Acidic nuclear phosphoprotein pp32) (Leucine-rich acidic nuclear protein) (Lanp) (Putative HLA-DR-associated protein I) (PHAPI) (Mapmodulin). | |||||
|
BAZ2B_HUMAN
|
||||||
| θ value | 2.36792 (rank : 69) | NC score | 0.010901 (rank : 94) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 803 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9UIF8, Q96EA1, Q96SQ8, Q9P252, Q9Y4N8 | Gene names | BAZ2B, KIAA1476 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2B (hWALp4). | |||||
|
CC47_MOUSE
|
||||||
| θ value | 2.36792 (rank : 70) | NC score | 0.052318 (rank : 53) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9D024, Q3TSB5, Q3V1S7, Q8C5D9, Q920S6 | Gene names | Ccdc47, Asp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 47 precursor (Adipocyte-specific protein 4). | |||||
|
KIF3B_HUMAN
|
||||||
| θ value | 2.36792 (rank : 71) | NC score | 0.010660 (rank : 99) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 540 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O15066 | Gene names | KIF3B, KIAA0359 | |||
|
Domain Architecture |
|
|||||
| Description | Kinesin-like protein KIF3B (Microtubule plus end-directed kinesin motor 3B) (HH0048). | |||||
|
MYT1L_MOUSE
|
||||||
| θ value | 2.36792 (rank : 72) | NC score | 0.021231 (rank : 73) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 461 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P97500, O08996, Q8C643, Q8C7L4, Q8CHB4 | Gene names | Myt1l, Kiaa1106, Nzf1, Png1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1-like protein (MyT1L protein) (Postmitotic neural gene 1 protein) (Zinc finger protein Png-1) (Neural zinc finger factor 1) (NZF-1). | |||||
|
AN32C_HUMAN
|
||||||
| θ value | 3.0926 (rank : 73) | NC score | 0.061882 (rank : 49) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O43423 | Gene names | ANP32C, PP32R1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member C (Phosphoprotein 32-related protein 1) (Tumorigenic protein pp32r1). | |||||
|
AN32E_HUMAN
|
||||||
| θ value | 3.0926 (rank : 74) | NC score | 0.051974 (rank : 54) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9BTT0, Q8N1S4, Q8WWW9 | Gene names | ANP32E | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member E (LANP- like protein) (LANP-L). | |||||
|
AP3B1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 75) | NC score | 0.008946 (rank : 104) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O00203, O00580, Q7Z393, Q9HD66 | Gene names | AP3B1, ADTB3A | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-1 (Adapter-related protein complex 3 beta-1 subunit) (Beta3A-adaptin) (Adaptor protein complex AP-3 beta-1 subunit) (Clathrin assembly protein complex 3 beta-1 large chain). | |||||
|
HUWE1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 76) | NC score | 0.010774 (rank : 97) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7Z6Z7, O15029, Q4G2Z2, Q5H961, Q6P4D0, Q8NG67, Q9BUI0, Q9HCJ4, Q9NSL6, Q9P0A9 | Gene names | HUWE1, KIAA0312, KIAA1578, UREB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (Mcl-1 ubiquitin ligase E3) (Mule) (ARF-binding protein 1) (ARF-BP1). | |||||
|
HUWE1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 77) | NC score | 0.010860 (rank : 95) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7TMY8, Q4G2Z1, Q5BMM7, Q6NS61, Q8BNJ7, Q8CFH2, Q8VD14, Q921M5, Q9R0P2 | Gene names | Huwe1, Kiaa0312, Ureb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (E3Histone). | |||||
|
PP2CG_MOUSE
|
||||||
| θ value | 3.0926 (rank : 78) | NC score | 0.015377 (rank : 83) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q61074 | Gene names | Ppm1g, Fin13, Ppm1c | |||
|
Domain Architecture |

|
|||||
| Description | Protein phosphatase 2C isoform gamma (EC 3.1.3.16) (PP2C-gamma) (Protein phosphatase magnesium-dependent 1 gamma) (Protein phosphatase 1C) (Fibroblast growth factor-inducible protein 13) (FIN13). | |||||
|
SET_HUMAN
|
||||||
| θ value | 3.0926 (rank : 79) | NC score | 0.021514 (rank : 72) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q01105, Q15541, Q5VXV1 | Gene names | SET | |||
|
Domain Architecture |
|
|||||
| Description | Protein SET (Phosphatase 2A inhibitor I2PP2A) (I-2PP2A) (Template- activating factor I) (TAF-I) (HLA-DR-associated protein II) (PHAPII) (Inhibitor of granzyme A-activated DNase) (IGAAD). | |||||
|
SSXT_MOUSE
|
||||||
| θ value | 3.0926 (rank : 80) | NC score | 0.021887 (rank : 69) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q62280 | Gene names | Ss18, Ssxt, Syt | |||
|
Domain Architecture |
|
|||||
| Description | SSXT protein (SYT protein) (Synovial sarcoma-associated Ss18-alpha). | |||||
|
TRI26_MOUSE
|
||||||
| θ value | 3.0926 (rank : 81) | NC score | 0.003784 (rank : 121) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 339 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q99PN3, Q8C9E9, Q8JZT7, Q99PN2 | Gene names | Trim26, Znf173 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 26 (Zinc finger protein 173). | |||||
|
VPS72_HUMAN
|
||||||
| θ value | 3.0926 (rank : 82) | NC score | 0.068115 (rank : 43) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q15906, Q53GJ2, Q5U0R4 | Gene names | VPS72, TCFL1, YL1 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 72 homolog (Transcription factor-like 1) (Protein YL-1). | |||||
|
VPS72_MOUSE
|
||||||
| θ value | 3.0926 (rank : 83) | NC score | 0.068699 (rank : 42) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q62481, Q3U2N4, Q810A9, Q99K81 | Gene names | Vps72, Tcfl1, Yl1 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 72 homolog (Transcription factor-like 1) (Protein YL-1). | |||||
|
ATAD2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 84) | NC score | 0.007397 (rank : 110) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 322 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6PL18, Q658P2, Q68CQ0, Q6PJV6, Q8N890, Q9UHS5 | Gene names | ATAD2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATPase family AAA domain-containing protein 2. | |||||
|
CLMN_HUMAN
|
||||||
| θ value | 4.03905 (rank : 85) | NC score | 0.006997 (rank : 114) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q96JQ2, Q9H713, Q9HA23, Q9HA57, Q9UFP4, Q9ULN2 | Gene names | CLMN, KIAA1188 | |||
|
Domain Architecture |
|
|||||
| Description | Calmin. | |||||
|
CYLC1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 86) | NC score | 0.013100 (rank : 89) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P35663, Q5JQQ9 | Gene names | CYLC1, CYL, CYL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-1 (Cylicin I) (Multiple-band polypeptide I). | |||||
|
KI21B_HUMAN
|
||||||
| θ value | 4.03905 (rank : 87) | NC score | 0.010529 (rank : 100) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1017 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O75037, Q5T4J3 | Gene names | KIF21B, KIAA0449 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 21B. | |||||
|
PHF14_MOUSE
|
||||||
| θ value | 4.03905 (rank : 88) | NC score | 0.013211 (rank : 88) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9D4H9, Q810Z6, Q8C284, Q8CGF8, Q9D1Q8 | Gene names | Phf14 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 14. | |||||
|
RAD50_HUMAN
|
||||||
| θ value | 4.03905 (rank : 89) | NC score | 0.006755 (rank : 115) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1317 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q92878, O43254, Q6GMT7, Q6P5X3, Q9UP86 | Gene names | RAD50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (hRAD50). | |||||
|
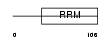
RBMX2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 90) | NC score | 0.006173 (rank : 116) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y388, Q5JY82, Q9Y3I8 | Gene names | RBMX2 | |||
|
Domain Architecture |
|
|||||
| Description | RNA-binding motif protein, X-linked 2. | |||||
|
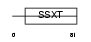
S18L1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 91) | NC score | 0.022613 (rank : 67) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75177, Q5JXJ3, Q8NE69, Q9BR55, Q9H4K6 | Gene names | SS18L1, KIAA0693 | |||
|
Domain Architecture |
|
|||||
| Description | SS18-like protein 1 (SYT homolog 1). | |||||
|
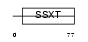
SSXT_HUMAN
|
||||||
| θ value | 4.03905 (rank : 92) | NC score | 0.021036 (rank : 75) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15532, Q16404, Q9BXC6 | Gene names | SS18, SSXT, SYT | |||
|
Domain Architecture |
|
|||||
| Description | SSXT protein (Synovial sarcoma, translocated to X chromosome) (SYT protein). | |||||
|
ABTAP_MOUSE
|
||||||
| θ value | 5.27518 (rank : 93) | NC score | 0.011305 (rank : 93) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q3V1V3 | Gene names | Abtap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ABT1-associated protein. | |||||
|
CCD39_MOUSE
|
||||||
| θ value | 5.27518 (rank : 94) | NC score | 0.007369 (rank : 111) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1026 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9D5Y1, Q8CDG6 | Gene names | Ccdc39, D3Ertd789e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 39. | |||||
|
CK060_HUMAN
|
||||||
| θ value | 5.27518 (rank : 95) | NC score | 0.014003 (rank : 84) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NQC8 | Gene names | C11orf60, C11orf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UPF0360 protein C11orf60. | |||||
|
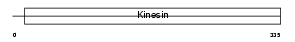
KIF3B_MOUSE
|
||||||
| θ value | 5.27518 (rank : 96) | NC score | 0.009730 (rank : 103) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 533 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q61771 | Gene names | Kif3b | |||
|
Domain Architecture |
|
|||||
| Description | Kinesin-like protein KIF3B (Microtubule plus end-directed kinesin motor 3B). | |||||
|
OSTP_HUMAN
|
||||||
| θ value | 5.27518 (rank : 97) | NC score | 0.016125 (rank : 79) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P10451, Q15681, Q15682, Q15683, Q8NBK2, Q96IZ1 | Gene names | SPP1, OPN | |||
|
Domain Architecture |

|
|||||
| Description | Osteopontin precursor (Bone sialoprotein-1) (Secreted phosphoprotein 1) (SPP-1) (Urinary stone protein) (Nephropontin) (Uropontin). | |||||
|
PP2CG_HUMAN
|
||||||
| θ value | 5.27518 (rank : 98) | NC score | 0.015396 (rank : 82) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O15355 | Gene names | PPM1G, PPM1C | |||
|
Domain Architecture |

|
|||||
| Description | Protein phosphatase 2C isoform gamma (EC 3.1.3.16) (PP2C-gamma) (Protein phosphatase magnesium-dependent 1 gamma) (Protein phosphatase 1C). | |||||
|
SEC63_MOUSE
|
||||||
| θ value | 5.27518 (rank : 99) | NC score | 0.026700 (rank : 62) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 601 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q8VHE0, Q8VEB9 | Gene names | Sec63, Sec63l | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
CALR_MOUSE
|
||||||
| θ value | 6.88961 (rank : 100) | NC score | 0.007138 (rank : 112) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P14211 | Gene names | Calr | |||
|
Domain Architecture |

|
|||||
| Description | Calreticulin precursor (CRP55) (Calregulin) (HACBP) (ERp60). | |||||
|
CI004_HUMAN
|
||||||
| θ value | 6.88961 (rank : 101) | NC score | 0.016407 (rank : 78) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 5 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9P0K9, Q5T4G4 | Gene names | C9orf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C9orf4 (Brain protein CG-6). | |||||
|
CK051_HUMAN
|
||||||
| θ value | 6.88961 (rank : 102) | NC score | 0.022204 (rank : 68) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P60006, Q9CXK2, Q9Y269 | Gene names | C11orf51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C11orf51. | |||||
|
CK051_MOUSE
|
||||||
| θ value | 6.88961 (rank : 103) | NC score | 0.021782 (rank : 70) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P60007, Q3UXV5, Q9CXK2, Q9Y269 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C11orf51 homolog. | |||||
|
LEO1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 104) | NC score | 0.036143 (rank : 58) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
NP1L1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 105) | NC score | 0.013425 (rank : 87) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P55209 | Gene names | NAP1L1, NRP | |||
|
Domain Architecture |
|
|||||
| Description | Nucleosome assembly protein 1-like 1 (NAP-1-related protein) (hNRP). | |||||
|
NP1L1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 106) | NC score | 0.013687 (rank : 86) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P28656, Q3UL14 | Gene names | Nap1l1, Nrp | |||
|
Domain Architecture |
|
|||||
| Description | Nucleosome assembly protein 1-like 1 (NAP-1-related protein) (Brain protein DN38). | |||||
|
RPGR1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 107) | NC score | 0.016108 (rank : 80) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 713 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9EPQ2, Q8CAC2, Q8CDJ9, Q91WE0, Q9CUK6, Q9D5Q1 | Gene names | Rpgrip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator-interacting protein 1 (RPGR-interacting protein 1). | |||||
|
WDR43_HUMAN
|
||||||
| θ value | 6.88961 (rank : 108) | NC score | 0.008247 (rank : 108) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q15061, Q15395, Q92577 | Gene names | WDR43, KIAA0007 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 43. | |||||
|
BOCT_HUMAN
|
||||||
| θ value | 8.99809 (rank : 109) | NC score | 0.271147 (rank : 38) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8WUG5, Q86U04, Q9H1D3, Q9NQD5 | Gene names | SLC22A17, BOCT, BOIT | |||
|
Domain Architecture |
|
|||||
| Description | Brain-type organic cation transporter (Solute carrier family 22 member 17). | |||||
|
CAC1D_HUMAN
|
||||||
| θ value | 8.99809 (rank : 110) | NC score | 0.001875 (rank : 123) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q01668, Q13916, Q13931 | Gene names | CACNA1D, CACH3, CACN4, CACNL1A2, CCHL1A2 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1D (Voltage- gated calcium channel subunit alpha Cav1.3) (Calcium channel, L type, alpha-1 polypeptide, isoform 2). | |||||
|
CENPB_HUMAN
|
||||||
| θ value | 8.99809 (rank : 111) | NC score | 0.017725 (rank : 77) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P07199 | Gene names | CENPB | |||
|
Domain Architecture |

|
|||||
| Description | Major centromere autoantigen B (Centromere protein B) (CENP-B). | |||||
|
CFDP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 112) | NC score | 0.011309 (rank : 92) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O88271, O70565, Q9JMA5 | Gene names | Cfdp1, Bcnt, Cfdp, Cp27 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Craniofacial development protein 1 (Bucentaur) (27 kDa craniofacial protein) (Protein Cp27). | |||||
|
CN037_HUMAN
|
||||||
| θ value | 8.99809 (rank : 113) | NC score | 0.008843 (rank : 105) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q86TY3, Q6P5Q1, Q86TY1 | Gene names | C14orf37 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf37 precursor. | |||||
|
FAM9A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 114) | NC score | 0.010306 (rank : 101) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8IZU1 | Gene names | FAM9A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM9A. | |||||
|
HIAL1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 115) | NC score | 0.008571 (rank : 107) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5SR56, Q3KQT4, Q53GU5, Q8WU95, Q96SM4 | Gene names | HIATL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hippocampus abundant transcript-like protein 1. | |||||
|
HIAL1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 116) | NC score | 0.008640 (rank : 106) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CIA9, Q8VCW0 | Gene names | Hiatl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hippocampus abundant transcript-like protein 1. | |||||
|
MIA2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 117) | NC score | 0.010265 (rank : 102) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q91ZV0 | Gene names | Mia2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma inhibitory activity protein 2 precursor. | |||||
|
MYT1L_HUMAN
|
||||||
| θ value | 8.99809 (rank : 118) | NC score | 0.018495 (rank : 76) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UL68, Q6IQ17, Q9UPP6 | Gene names | MYT1L, KIAA1106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1-like protein (MyT1L protein) (MyT1-L). | |||||
|
NUCL_MOUSE
|
||||||
| θ value | 8.99809 (rank : 119) | NC score | 0.010838 (rank : 96) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P09405, Q548M9, Q61991, Q99K50 | Gene names | Ncl, Nuc | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolin (Protein C23). | |||||
|
PELP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 120) | NC score | 0.005229 (rank : 118) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9DBD5, Q5F2E2, Q6PEM0, Q91YM9 | Gene names | Pelp1, Mnar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor). | |||||
|
PEPL_HUMAN
|
||||||
| θ value | 8.99809 (rank : 121) | NC score | 0.003745 (rank : 122) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1141 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O60437, O60314, O60454 | Gene names | PPL, KIAA0568 | |||
|
Domain Architecture |
|
|||||
| Description | Periplakin (195 kDa cornified envelope precursor protein) (190 kDa paraneoplastic pemphigus antigen). | |||||
|
PSYR_MOUSE
|
||||||
| θ value | 8.99809 (rank : 122) | NC score | -0.001377 (rank : 124) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 780 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61038, Q921N3 | Gene names | Gpr65, Gpcr25, Tdag8 | |||
|
Domain Architecture |
|
|||||
| Description | Psychosine receptor (G-protein coupled receptor 65) (T cell death- associated protein 8). | |||||
|
S17A5_HUMAN
|
||||||
| θ value | 8.99809 (rank : 123) | NC score | 0.039239 (rank : 57) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NRA2, Q8NBR5, Q9UGH0 | Gene names | SLC17A5 | |||
|
Domain Architecture |
|
|||||
| Description | Sialin (Solute carrier family 17 member 5) (Sodium/sialic acid cotransporter) (AST) (Membrane glycoprotein HP59). | |||||
|
TRI44_HUMAN
|
||||||
| θ value | 8.99809 (rank : 124) | NC score | 0.005672 (rank : 117) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 393 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q96DX7, Q96QY2, Q9UGK0 | Gene names | TRIM44, DIPB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 44 (Protein DIPB). | |||||
|
SV2C_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 5) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 124 | |
| SwissProt Accessions | Q496J9, Q496K1, Q9UPU8 | Gene names | SV2C, KIAA1054 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptic vesicle glycoprotein 2C. | |||||
|
SV2C_MOUSE
|
||||||
| NC score | 0.994150 (rank : 2) | θ value | 0 (rank : 6) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q69ZS6 | Gene names | Sv2c, Kiaa1054 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptic vesicle glycoprotein 2C (Synaptic vesicle protein 2C). | |||||
|
SV2B_HUMAN
|
||||||
| NC score | 0.986002 (rank : 3) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q7L1I2, O94840, Q6IAR8 | Gene names | SV2B, KIAA0735 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptic vesicle glycoprotein 2B. | |||||
|
SV2B_MOUSE
|
||||||
| NC score | 0.984073 (rank : 4) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8BG39, Q80TT1, Q9ES95 | Gene names | Sv2b, Kiaa0735 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptic vesicle glycoprotein 2B (Synaptic vesicle protein 2B). | |||||
|
SV2A_HUMAN
|
||||||
| NC score | 0.978076 (rank : 5) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q7L0J3, O94841, Q5QNX8, Q7Z3L6, Q8NBJ6, Q9BVZ9 | Gene names | SV2A, KIAA0736 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptic vesicle glycoprotein 2A. | |||||
|
SV2A_MOUSE
|
||||||
| NC score | 0.977920 (rank : 6) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9JIS5, Q80TT0, Q8R0R5 | Gene names | Sv2a, Kiaa0736, Sv2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptic vesicle glycoprotein 2A (Synaptic vesicle protein 2A) (Synaptic vesicle protein 2) (Calcium regulator SV2A). | |||||
|
S22A5_HUMAN
|
||||||
| NC score | 0.582638 (rank : 7) | θ value | 5.60996e-18 (rank : 7) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O76082, Q6ZQZ8, Q96EH6 | Gene names | SLC22A5, OCTN2 | |||
|
Domain Architecture |

|
|||||
| Description | Organic cation/carnitine transporter 2 (Solute carrier family 22 member 5) (High-affinity sodium-dependent carnitine cotransporter). | |||||
|
OCTN3_MOUSE
|
||||||
| NC score | 0.567410 (rank : 8) | θ value | 5.8054e-15 (rank : 9) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9WTN6, Q5SWV0 | Gene names | Slc22a21, Octn3, Slc22a9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Organic cation/carnitine transporter 3 (Solute carrier family 22 member 21) (Solute carrier family 22 member 9). | |||||
|
S22A5_MOUSE
|
||||||
| NC score | 0.567247 (rank : 9) | θ value | 1.80466e-16 (rank : 8) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9Z0E8 | Gene names | Slc22a5, Octn2 | |||
|
Domain Architecture |

|
|||||
| Description | Organic cation/carnitine transporter 2 (Solute carrier family 22 member 5) (High-affinity sodium-dependent carnitine cotransporter). | |||||
|
S22A4_MOUSE
|
||||||
| NC score | 0.550843 (rank : 10) | θ value | 7.09661e-13 (rank : 10) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9Z306 | Gene names | Slc22a4, Octn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Organic cation/carnitine transporter 1 (Solute carrier family 22 member 4). | |||||
|
S22A4_HUMAN
|
||||||
| NC score | 0.545393 (rank : 11) | θ value | 1.74796e-11 (rank : 12) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9H015, O14546 | Gene names | SLC22A4, ETT, OCTN1, UT2H | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Organic cation/carnitine transporter 1 (Solute carrier family 22 member 4) (Ergothioneine transporter) (ET transporter). | |||||
|
ORCT3_HUMAN
|
||||||
| NC score | 0.533823 (rank : 12) | θ value | 1.133e-10 (rank : 14) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9Y226, Q8IYG1 | Gene names | SLC22A13, ORCTL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Organic-cation transporter-like 3 (Solute carrier family 22 member 13). | |||||
|
ORCT3_MOUSE
|
||||||
| NC score | 0.530375 (rank : 13) | θ value | 1.25267e-09 (rank : 15) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q6A4L0, Q8VC89 | Gene names | Slc22a13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Organic cation transporter-like 3 (Solute carrier family 22 member 13). | |||||
|
S22A3_MOUSE
|
||||||
| NC score | 0.527220 (rank : 14) | θ value | 3.64472e-09 (rank : 17) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9WTW5, Q9R209 | Gene names | Slc22a3, Oct3 | |||
|
Domain Architecture |

|
|||||
| Description | Organic cation transporter 3 (Solute carrier family 22 member 3). | |||||
|
S22A3_HUMAN
|
||||||
| NC score | 0.504229 (rank : 15) | θ value | 3.08544e-08 (rank : 21) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O75751, Q9UP02 | Gene names | SLC22A3, EMTH | |||
|
Domain Architecture |

|
|||||
| Description | Organic cation transporter 3 (Extraneuronal monoamine transporter) (EMT) (Solute carrier family 22 member 3). | |||||
|
OAT6_MOUSE
|
||||||
| NC score | 0.501943 (rank : 16) | θ value | 2.36244e-08 (rank : 20) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q80UJ1 | Gene names | Slc22a20, Gm963, Oat6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Solute carrier family 22 member 20 (Organic anion transporter 6). | |||||
|
ORCT4_HUMAN
|
||||||
| NC score | 0.487238 (rank : 17) | θ value | 1.80886e-08 (rank : 19) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9Y267, Q6DJT3 | Gene names | SLC22A14, ORCTL4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Organic-cation transporter-like 4 (Solute carrier family 22 member 14). | |||||
|
GTR8_HUMAN
|
||||||
| NC score | 0.486586 (rank : 18) | θ value | 1.74796e-11 (rank : 11) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9NY64, Q9NSC4 | Gene names | SLC2A8, GLUT8, GLUTX1 | |||
|
Domain Architecture |

|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 8 (Glucose transporter type 8) (Glucose transporter type X1). | |||||
|
GTR8_MOUSE
|
||||||
| NC score | 0.485434 (rank : 19) | θ value | 6.64225e-11 (rank : 13) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9JIF3, Q9JJP4, Q9JJZ0 | Gene names | Slc2a8, Glut8, GlutX1 | |||
|
Domain Architecture |

|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 8 (Glucose transporter type 8) (Glucose transporter type X1). | |||||
|
OAT4_HUMAN
|
||||||
| NC score | 0.470331 (rank : 20) | θ value | 4.45536e-07 (rank : 22) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9NSA0, Q53GR2, Q6ZP72, Q8NBU4 | Gene names | SLC22A11, OAT4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Solute carrier family 22 member 11 (Organic anion transporter 4). | |||||
|
GTR6_HUMAN
|
||||||
| NC score | 0.467924 (rank : 21) | θ value | 2.13673e-09 (rank : 16) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9UGQ3, Q5SXD7 | Gene names | SLC2A6, GLUT9 | |||
|
Domain Architecture |

|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 6 (Glucose transporter type 6) (Glucose transporter type 9). | |||||
|
MYCT_HUMAN
|
||||||
| NC score | 0.464363 (rank : 22) | θ value | 5.81887e-07 (rank : 23) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q96QE2 | Gene names | SLC2A13 | |||
|
Domain Architecture |

|
|||||
| Description | Proton myo-inositol cotransporter (H(+)-myo-inositol cotransporter) (Hmit). | |||||
|
OAT5_HUMAN
|
||||||
| NC score | 0.419208 (rank : 23) | θ value | 0.00228821 (rank : 36) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q63ZE4 | Gene names | SLC22A10, OAT5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Solute carrier family 22 member 10 (Organic anion transporter 5). | |||||
|
S22A9_HUMAN
|
||||||
| NC score | 0.399767 (rank : 24) | θ value | 0.0148317 (rank : 38) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8IVM8, Q8TEC0, Q8WYN7 | Gene names | SLC22A9, OAT4, UST3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Solute carrier family 22 member 9 (Organic anion transporter 4). | |||||
|
GTR11_HUMAN
|
||||||
| NC score | 0.392623 (rank : 25) | θ value | 1.43324e-05 (rank : 25) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9BYW1 | Gene names | SLC2A11, GLUT10, GLUT11 | |||
|
Domain Architecture |

|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 11 (Glucose transporter type 11) (Glucose transporter type 10). | |||||
|
GTR3_HUMAN
|
||||||
| NC score | 0.392237 (rank : 26) | θ value | 2.88788e-06 (rank : 24) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P11169, Q6I9U2, Q9UG15 | Gene names | SLC2A3, GLUT3 | |||
|
Domain Architecture |
|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 3 (Glucose transporter type 3, brain). | |||||
|
GTR5_MOUSE
|
||||||
| NC score | 0.390961 (rank : 27) | θ value | 7.1131e-05 (rank : 29) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9WV38 | Gene names | Slc2a5, Glut5 | |||
|
Domain Architecture |

|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 5 (Glucose transporter type 5, small intestine) (Fructose transporter). | |||||
|
GTR1_HUMAN
|
||||||
| NC score | 0.389181 (rank : 28) | θ value | 4.1701e-05 (rank : 28) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P11166, O75535 | Gene names | SLC2A1, GLUT1 | |||
|
Domain Architecture |
|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 1 (Glucose transporter type 1, erythrocyte/brain) (HepG2 glucose transporter). | |||||
|
GTR1_MOUSE
|
||||||
| NC score | 0.388126 (rank : 29) | θ value | 2.44474e-05 (rank : 27) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P17809, Q61608 | Gene names | Slc2a1, Glut-1, Glut1 | |||
|
Domain Architecture |
|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 1 (Glucose transporter type 1, erythrocyte/brain) (GT1). | |||||
|
GTR3_MOUSE
|
||||||
| NC score | 0.387711 (rank : 30) | θ value | 1.87187e-05 (rank : 26) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P32037 | Gene names | Slc2a3, Glut-3, Glut3 | |||
|
Domain Architecture |

|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 3 (Glucose transporter type 3, brain). | |||||
|
GTR9_HUMAN
|
||||||
| NC score | 0.384747 (rank : 31) | θ value | 0.00020696 (rank : 32) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9NRM0 | Gene names | SLC2A9, GLUT9 | |||
|
Domain Architecture |

|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 9 (Glucose transporter type 9). | |||||
|
GTR2_MOUSE
|
||||||
| NC score | 0.380809 (rank : 32) | θ value | 8.11959e-09 (rank : 18) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P14246, Q3TNA5, Q9DBA7 | Gene names | Slc2a2, Glut-2, Glut2 | |||
|
Domain Architecture |
|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 2 (Glucose transporter type 2, liver). | |||||
|
GTR14_HUMAN
|
||||||
| NC score | 0.371508 (rank : 33) | θ value | 0.000121331 (rank : 31) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8TDB8, Q6UY84, Q8TDB9 | Gene names | SLC2A14, GLUT14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 14 (Glucose transporter type 14). | |||||
|
GTR5_HUMAN
|
||||||
| NC score | 0.359425 (rank : 34) | θ value | 9.29e-05 (rank : 30) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P22732, Q14770 | Gene names | SLC2A5, GLUT5 | |||
|
Domain Architecture |

|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 5 (Glucose transporter type 5, small intestine) (Fructose transporter). | |||||
|
GTR4_MOUSE
|
||||||
| NC score | 0.321606 (rank : 35) | θ value | 0.000270298 (rank : 33) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P14142, Q9JJN9 | Gene names | Slc2a4, Glut-4, Glut4 | |||
|
Domain Architecture |

|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 4 (Glucose transporter type 4, insulin-responsive) (GT2). | |||||
|
GTR4_HUMAN
|
||||||
| NC score | 0.318092 (rank : 36) | θ value | 0.00175202 (rank : 35) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P14672 | Gene names | SLC2A4, GLUT4 | |||
|
Domain Architecture |

|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 4 (Glucose transporter type 4, insulin-responsive). | |||||
|
GTR2_HUMAN
|
||||||
| NC score | 0.316866 (rank : 37) | θ value | 0.00134147 (rank : 34) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P11168 | Gene names | SLC2A2, GLUT2 | |||
|
Domain Architecture |

|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 2 (Glucose transporter type 2, liver). | |||||
|
BOCT_HUMAN
|
||||||
| NC score | 0.271147 (rank : 38) | θ value | 8.99809 (rank : 109) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8WUG5, Q86U04, Q9H1D3, Q9NQD5 | Gene names | SLC22A17, BOCT, BOIT | |||
|
Domain Architecture |

|
|||||
| Description | Brain-type organic cation transporter (Solute carrier family 22 member 17). | |||||
|
GTR10_HUMAN
|
||||||
| NC score | 0.267450 (rank : 39) | θ value | 1.06291 (rank : 53) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O95528, Q9H4I6 | Gene names | SLC2A10, GLUT10 | |||
|
Domain Architecture |

|
|||||
| Description | Solute carrier family 2, facilitated glucose transporter member 10 (Glucose transporter type 10). | |||||
|
AN32B_HUMAN
|
||||||
| NC score | 0.071391 (rank : 40) | θ value | 0.279714 (rank : 44) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q92688, O00655, P78458, P78459 | Gene names | ANP32B, APRIL, PHAPI2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member B (PHAPI2 protein) (Silver-stainable protein SSP29) (Acidic protein rich in leucines). | |||||
|
AN32B_MOUSE
|
||||||
| NC score | 0.069515 (rank : 41) | θ value | 0.62314 (rank : 48) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9EST5 | Gene names | Anp32b, Pal31 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member B (Proliferation-related acidic leucine-rich protein PAL31). | |||||
|
VPS72_MOUSE
|
||||||
| NC score | 0.068699 (rank : 42) | θ value | 3.0926 (rank : 83) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q62481, Q3U2N4, Q810A9, Q99K81 | Gene names | Vps72, Tcfl1, Yl1 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 72 homolog (Transcription factor-like 1) (Protein YL-1). | |||||
|
VPS72_HUMAN
|
||||||
| NC score | 0.068115 (rank : 43) | θ value | 3.0926 (rank : 82) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q15906, Q53GJ2, Q5U0R4 | Gene names | VPS72, TCFL1, YL1 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 72 homolog (Transcription factor-like 1) (Protein YL-1). | |||||
|
AN32A_MOUSE
|
||||||
| NC score | 0.066880 (rank : 44) | θ value | 0.0961366 (rank : 39) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 254 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O35381, P97437 | Gene names | Anp32a, Anp32, Lanp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member A (Potent heat-stable protein phosphatase 2A inhibitor I1PP2A) (Acidic nuclear phosphoprotein pp32) (Leucine-rich acidic nuclear protein). | |||||
|
AN32C_MOUSE
|
||||||
| NC score | 0.066880 (rank : 45) | θ value | 0.0961366 (rank : 40) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 254 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q64G17 | Gene names | Anp32c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member C. | |||||
|
AN32A_HUMAN
|
||||||
| NC score | 0.064900 (rank : 46) | θ value | 2.36792 (rank : 68) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P39687 | Gene names | ANP32A, C15orf1, LANP, MAPM, PHAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member A (Potent heat-stable protein phosphatase 2A inhibitor I1PP2A) (Acidic nuclear phosphoprotein pp32) (Leucine-rich acidic nuclear protein) (Lanp) (Putative HLA-DR-associated protein I) (PHAPI) (Mapmodulin). | |||||
|
AN32E_MOUSE
|
||||||
| NC score | 0.063769 (rank : 47) | θ value | 0.365318 (rank : 45) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P97822, Q3TH89, Q8BPF8, Q8C2L4, Q8C7Q8, Q9CZD2 | Gene names | Anp32e, Cpd1 | |||
|
Domain Architecture |

|
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member E (LANP- like protein) (LANP-L) (Cerebellar postnatal development protein 1). | |||||
|
CT059_MOUSE
|
||||||
| NC score | 0.062909 (rank : 48) | θ value | 0.00869519 (rank : 37) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8VCL5 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative transporter C20orf59 homolog. | |||||
|
AN32C_HUMAN
|
||||||
| NC score | 0.061882 (rank : 49) | θ value | 3.0926 (rank : 73) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O43423 | Gene names | ANP32C, PP32R1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member C (Phosphoprotein 32-related protein 1) (Tumorigenic protein pp32r1). | |||||
|
VMAT2_MOUSE
|
||||||
| NC score | 0.060181 (rank : 50) | θ value | 0.47712 (rank : 47) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8BRU6, Q8CC55 | Gene names | Slc18a2, Vmat2 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptic vesicular amine transporter (Monoamine transporter) (Vesicular amine transporter 2) (VAT2) (Solute carrier family 18 member 2). | |||||
|
CENPB_MOUSE
|
||||||
| NC score | 0.053452 (rank : 51) | θ value | 1.38821 (rank : 55) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P27790 | Gene names | Cenpb, Cenp-b | |||
|
Domain Architecture |

|
|||||
| Description | Major centromere autoantigen B (Centromere protein B) (CENP-B). | |||||
|
VMAT2_HUMAN
|
||||||
| NC score | 0.052731 (rank : 52) | θ value | 1.38821 (rank : 59) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q05940, Q15876, Q9H3P6 | Gene names | SLC18A2, SVMT, VMAT2 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptic vesicular amine transporter (Monoamine transporter) (Vesicular amine transporter 2) (VAT2) (Solute carrier family 18 member 2). | |||||
|
CC47_MOUSE
|
||||||
| NC score | 0.052318 (rank : 53) | θ value | 2.36792 (rank : 70) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9D024, Q3TSB5, Q3V1S7, Q8C5D9, Q920S6 | Gene names | Ccdc47, Asp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 47 precursor (Adipocyte-specific protein 4). | |||||
|
AN32E_HUMAN
|
||||||
| NC score | 0.051974 (rank : 54) | θ value | 3.0926 (rank : 74) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9BTT0, Q8N1S4, Q8WWW9 | Gene names | ANP32E | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member E (LANP- like protein) (LANP-L). | |||||
|
HNRCL_HUMAN
|
||||||
| NC score | 0.049197 (rank : 55) | θ value | 0.21417 (rank : 42) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O60812 | Gene names | HNRPCL1 | |||
|
Domain Architecture |

|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein C-like 1 (hnRNP core protein C-like 1). | |||||
|
HNRPC_HUMAN
|
||||||
| NC score | 0.039847 (rank : 56) | θ value | 0.163984 (rank : 41) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P07910, P22628, Q53EX2, Q59FD3, Q5FWE8, Q86SF8, Q86U45, Q96HK7, Q96HM4, Q96IY5, Q9BTS3 | Gene names | HNRPC | |||
|
Domain Architecture |

|
|||||
| Description | Heterogeneous nuclear ribonucleoproteins C1/C2 (hnRNP C1 / hnRNP C2). | |||||
|
S17A5_HUMAN
|
||||||
| NC score | 0.039239 (rank : 57) | θ value | 8.99809 (rank : 123) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NRA2, Q8NBR5, Q9UGH0 | Gene names | SLC17A5 | |||
|
Domain Architecture |

|
|||||
| Description | Sialin (Solute carrier family 17 member 5) (Sodium/sialic acid cotransporter) (AST) (Membrane glycoprotein HP59). | |||||
|
LEO1_MOUSE
|
||||||
| NC score | 0.036143 (rank : 58) | θ value | 6.88961 (rank : 104) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
UBF1_MOUSE
|
||||||
| NC score | 0.029650 (rank : 59) | θ value | 1.38821 (rank : 58) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P25976 | Gene names | Ubtf, Tcfubf, Ubf-1, Ubf1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar transcription factor 1 (Upstream-binding factor 1) (UBF-1). | |||||
|
CCAR1_HUMAN
|
||||||
| NC score | 0.027934 (rank : 60) | θ value | 0.365318 (rank : 46) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 444 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8IX12, Q32NE3, Q5VUP6, Q6PIZ0, Q6X935, Q9H8N4, Q9NVA7, Q9NVQ0, Q9NWM6 | Gene names | CCAR1, CARP1, DIS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle and apoptosis regulator protein 1 (Cell cycle and apoptosis regulatory protein 1) (CARP-1) (Death inducer with SAP domain). | |||||
|
DAXX_MOUSE
|
||||||
| NC score | 0.026997 (rank : 61) | θ value | 1.06291 (rank : 52) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O35613, Q9QWT8, Q9QWV3 | Gene names | Daxx | |||
|
Domain Architecture |

|
|||||
| Description | Death domain-associated protein 6 (Daxx). | |||||
|
SEC63_MOUSE
|
||||||
| NC score | 0.026700 (rank : 62) | θ value | 5.27518 (rank : 99) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 601 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q8VHE0, Q8VEB9 | Gene names | Sec63, Sec63l | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
MYT1_MOUSE
|
||||||
| NC score | 0.026612 (rank : 63) | θ value | 0.21417 (rank : 43) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8CFC2, O08995, Q8CFH1 | Gene names | Myt1, Kiaa0835, Nzf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1 (MyT1) (Neural zinc finger factor 2) (NZF-2). | |||||
|
CYLC2_HUMAN
|
||||||
| NC score | 0.025620 (rank : 64) | θ value | 1.81305 (rank : 61) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q14093 | Gene names | CYLC2, CYL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-2 (Cylicin II) (Multiple-band polypeptide II). | |||||
|
SET_MOUSE
|
||||||
| NC score | 0.024881 (rank : 65) | θ value | 1.06291 (rank : 54) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9EQU5, Q9CY82, Q9D0A9, Q9Z181 | Gene names | Set | |||
|
Domain Architecture |

|
|||||
| Description | Protein SET (Phosphatase 2A inhibitor I2PP2A) (I-2PP2A) (Template- activating factor I) (TAF-I). | |||||
|
CCAR1_MOUSE
|
||||||
| NC score | 0.024325 (rank : 66) | θ value | 1.81305 (rank : 60) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8CH18, Q6AXC9, Q6PAR2, Q80XE4, Q8BJY0, Q8BVN2, Q8CGG1, Q9CSR5 | Gene names | Ccar1, Carp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle and apoptosis regulator protein 1 (Cell cycle and apoptosis regulatory protein 1) (CARP-1). | |||||
|
S18L1_HUMAN
|
||||||
| NC score | 0.022613 (rank : 67) | θ value | 4.03905 (rank : 91) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75177, Q5JXJ3, Q8NE69, Q9BR55, Q9H4K6 | Gene names | SS18L1, KIAA0693 | |||
|
Domain Architecture |

|
|||||
| Description | SS18-like protein 1 (SYT homolog 1). | |||||
|
CK051_HUMAN
|
||||||
| NC score | 0.022204 (rank : 68) | θ value | 6.88961 (rank : 102) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P60006, Q9CXK2, Q9Y269 | Gene names | C11orf51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C11orf51. | |||||
|
SSXT_MOUSE
|
||||||
| NC score | 0.021887 (rank : 69) | θ value | 3.0926 (rank : 80) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q62280 | Gene names | Ss18, Ssxt, Syt | |||
|
Domain Architecture |

|
|||||
| Description | SSXT protein (SYT protein) (Synovial sarcoma-associated Ss18-alpha). | |||||
|
CK051_MOUSE
|
||||||
| NC score | 0.021782 (rank : 70) | θ value | 6.88961 (rank : 103) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P60007, Q3UXV5, Q9CXK2, Q9Y269 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C11orf51 homolog. | |||||
|
SRCH_HUMAN
|
||||||
| NC score | 0.021757 (rank : 71) | θ value | 1.81305 (rank : 66) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P23327, Q504Y6 | Gene names | HRC, HCP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcoplasmic reticulum histidine-rich calcium-binding protein precursor. | |||||
|
SET_HUMAN
|
||||||
| NC score | 0.021514 (rank : 72) | θ value | 3.0926 (rank : 79) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q01105, Q15541, Q5VXV1 | Gene names | SET | |||
|
Domain Architecture |

|
|||||
| Description | Protein SET (Phosphatase 2A inhibitor I2PP2A) (I-2PP2A) (Template- activating factor I) (TAF-I) (HLA-DR-associated protein II) (PHAPII) (Inhibitor of granzyme A-activated DNase) (IGAAD). | |||||
|
MYT1L_MOUSE
|
||||||
| NC score | 0.021231 (rank : 73) | θ value | 2.36792 (rank : 72) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 461 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P97500, O08996, Q8C643, Q8C7L4, Q8CHB4 | Gene names | Myt1l, Kiaa1106, Nzf1, Png1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1-like protein (MyT1L protein) (Postmitotic neural gene 1 protein) (Zinc finger protein Png-1) (Neural zinc finger factor 1) (NZF-1). | |||||
|
UBF1_HUMAN
|
||||||
| NC score | 0.021138 (rank : 74) | θ value | 1.81305 (rank : 67) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 415 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P17480 | Gene names | UBTF, UBF, UBF1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar transcription factor 1 (Upstream-binding factor 1) (UBF-1) (Autoantigen NOR-90). | |||||
|
SSXT_HUMAN
|
||||||
| NC score | 0.021036 (rank : 75) | θ value | 4.03905 (rank : 92) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15532, Q16404, Q9BXC6 | Gene names | SS18, SSXT, SYT | |||
|
Domain Architecture |

|
|||||
| Description | SSXT protein (Synovial sarcoma, translocated to X chromosome) (SYT protein). | |||||
|
MYT1L_HUMAN
|
||||||
| NC score | 0.018495 (rank : 76) | θ value | 8.99809 (rank : 118) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UL68, Q6IQ17, Q9UPP6 | Gene names | MYT1L, KIAA1106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1-like protein (MyT1L protein) (MyT1-L). | |||||
|
CENPB_HUMAN
|
||||||
| NC score | 0.017725 (rank : 77) | θ value | 8.99809 (rank : 111) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P07199 | Gene names | CENPB | |||
|
Domain Architecture |

|
|||||
| Description | Major centromere autoantigen B (Centromere protein B) (CENP-B). | |||||
|
CI004_HUMAN
|
||||||
| NC score | 0.016407 (rank : 78) | θ value | 6.88961 (rank : 101) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 5 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9P0K9, Q5T4G4 | Gene names | C9orf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C9orf4 (Brain protein CG-6). | |||||
|
OSTP_HUMAN
|
||||||
| NC score | 0.016125 (rank : 79) | θ value | 5.27518 (rank : 97) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P10451, Q15681, Q15682, Q15683, Q8NBK2, Q96IZ1 | Gene names | SPP1, OPN | |||
|
Domain Architecture |

|
|||||
| Description | Osteopontin precursor (Bone sialoprotein-1) (Secreted phosphoprotein 1) (SPP-1) (Urinary stone protein) (Nephropontin) (Uropontin). | |||||
|
RPGR1_MOUSE
|
||||||
| NC score | 0.016108 (rank : 80) | θ value | 6.88961 (rank : 107) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 713 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9EPQ2, Q8CAC2, Q8CDJ9, Q91WE0, Q9CUK6, Q9D5Q1 | Gene names | Rpgrip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator-interacting protein 1 (RPGR-interacting protein 1). | |||||
|
NRDC_MOUSE
|
||||||
| NC score | 0.015703 (rank : 81) | θ value | 1.81305 (rank : 63) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BHG1 | Gene names | Nrd1 | |||
|
Domain Architecture |

|
|||||
| Description | Nardilysin precursor (EC 3.4.24.61) (N-arginine dibasic convertase) (NRD convertase) (NRD-C). | |||||
|
PP2CG_HUMAN
|
||||||
| NC score | 0.015396 (rank : 82) | θ value | 5.27518 (rank : 98) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O15355 | Gene names | PPM1G, PPM1C | |||
|
Domain Architecture |

|
|||||
| Description | Protein phosphatase 2C isoform gamma (EC 3.1.3.16) (PP2C-gamma) (Protein phosphatase magnesium-dependent 1 gamma) (Protein phosphatase 1C). | |||||
|
PP2CG_MOUSE
|
||||||
| NC score | 0.015377 (rank : 83) | θ value | 3.0926 (rank : 78) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q61074 | Gene names | Ppm1g, Fin13, Ppm1c | |||
|
Domain Architecture |

|
|||||
| Description | Protein phosphatase 2C isoform gamma (EC 3.1.3.16) (PP2C-gamma) (Protein phosphatase magnesium-dependent 1 gamma) (Protein phosphatase 1C) (Fibroblast growth factor-inducible protein 13) (FIN13). | |||||
|
CK060_HUMAN
|
||||||
| NC score | 0.014003 (rank : 84) | θ value | 5.27518 (rank : 95) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NQC8 | Gene names | C11orf60, C11orf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UPF0360 protein C11orf60. | |||||
|
PHF14_HUMAN
|
||||||
| NC score | 0.013999 (rank : 85) | θ value | 1.81305 (rank : 64) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O94880 | Gene names | PHF14, KIAA0783 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 14. | |||||
|
NP1L1_MOUSE
|
||||||
| NC score | 0.013687 (rank : 86) | θ value | 6.88961 (rank : 106) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P28656, Q3UL14 | Gene names | Nap1l1, Nrp | |||
|
Domain Architecture |

|
|||||
| Description | Nucleosome assembly protein 1-like 1 (NAP-1-related protein) (Brain protein DN38). | |||||
|
NP1L1_HUMAN
|
||||||
| NC score | 0.013425 (rank : 87) | θ value | 6.88961 (rank : 105) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P55209 | Gene names | NAP1L1, NRP | |||
|
Domain Architecture |

|
|||||
| Description | Nucleosome assembly protein 1-like 1 (NAP-1-related protein) (hNRP). | |||||
|
PHF14_MOUSE
|
||||||
| NC score | 0.013211 (rank : 88) | θ value | 4.03905 (rank : 88) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9D4H9, Q810Z6, Q8C284, Q8CGF8, Q9D1Q8 | Gene names | Phf14 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 14. | |||||
|
CYLC1_HUMAN
|
||||||
| NC score | 0.013100 (rank : 89) | θ value | 4.03905 (rank : 86) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P35663, Q5JQQ9 | Gene names | CYLC1, CYL, CYL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-1 (Cylicin I) (Multiple-band polypeptide I). | |||||
|
IF2P_HUMAN
|
||||||
| NC score | 0.013048 (rank : 90) | θ value | 1.38821 (rank : 56) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 889 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O60841, O95805, Q9UF81, Q9UMN7 | Gene names | EIF5B, IF2, KIAA0741 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 5B (eIF-5B) (Translation initiation factor IF-2). | |||||
|
KI21B_MOUSE
|
||||||
| NC score | 0.011876 (rank : 91) | θ value | 0.813845 (rank : 49) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 997 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9QXL1, P97424 | Gene names | Kif21b, Kif6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 21B (Kinesin-like protein KIF6). | |||||
|
CFDP1_MOUSE
|
||||||
| NC score | 0.011309 (rank : 92) | θ value | 8.99809 (rank : 112) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O88271, O70565, Q9JMA5 | Gene names | Cfdp1, Bcnt, Cfdp, Cp27 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Craniofacial development protein 1 (Bucentaur) (27 kDa craniofacial protein) (Protein Cp27). | |||||
|
ABTAP_MOUSE
|
||||||
| NC score | 0.011305 (rank : 93) | θ value | 5.27518 (rank : 93) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q3V1V3 | Gene names | Abtap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ABT1-associated protein. | |||||
|
BAZ2B_HUMAN
|
||||||
| NC score | 0.010901 (rank : 94) | θ value | 2.36792 (rank : 69) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 803 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9UIF8, Q96EA1, Q96SQ8, Q9P252, Q9Y4N8 | Gene names | BAZ2B, KIAA1476 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2B (hWALp4). | |||||
|
HUWE1_MOUSE
|
||||||
| NC score | 0.010860 (rank : 95) | θ value | 3.0926 (rank : 77) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7TMY8, Q4G2Z1, Q5BMM7, Q6NS61, Q8BNJ7, Q8CFH2, Q8VD14, Q921M5, Q9R0P2 | Gene names | Huwe1, Kiaa0312, Ureb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (E3Histone). | |||||
|
NUCL_MOUSE
|
||||||
| NC score | 0.010838 (rank : 96) | θ value | 8.99809 (rank : 119) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P09405, Q548M9, Q61991, Q99K50 | Gene names | Ncl, Nuc | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolin (Protein C23). | |||||
|
HUWE1_HUMAN
|
||||||
| NC score | 0.010774 (rank : 97) | θ value | 3.0926 (rank : 76) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7Z6Z7, O15029, Q4G2Z2, Q5H961, Q6P4D0, Q8NG67, Q9BUI0, Q9HCJ4, Q9NSL6, Q9P0A9 | Gene names | HUWE1, KIAA0312, KIAA1578, UREB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (Mcl-1 ubiquitin ligase E3) (Mule) (ARF-binding protein 1) (ARF-BP1). | |||||
|
KI21A_MOUSE
|
||||||
| NC score | 0.010767 (rank : 98) | θ value | 1.81305 (rank : 62) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1317 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9QXL2, Q6P5H1, Q6ZPJ8, Q8BWZ9, Q8BXF1 | Gene names | Kif21a, Kiaa1708 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 21A. | |||||
|
KIF3B_HUMAN
|
||||||
| NC score | 0.010660 (rank : 99) | θ value | 2.36792 (rank : 71) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 540 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O15066 | Gene names | KIF3B, KIAA0359 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF3B (Microtubule plus end-directed kinesin motor 3B) (HH0048). | |||||
|
KI21B_HUMAN
|
||||||
| NC score | 0.010529 (rank : 100) | θ value | 4.03905 (rank : 87) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1017 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O75037, Q5T4J3 | Gene names | KIF21B, KIAA0449 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 21B. | |||||
|
FAM9A_HUMAN
|
||||||
| NC score | 0.010306 (rank : 101) | θ value | 8.99809 (rank : 114) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8IZU1 | Gene names | FAM9A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM9A. | |||||
|
MIA2_MOUSE
|
||||||
| NC score | 0.010265 (rank : 102) | θ value | 8.99809 (rank : 117) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q91ZV0 | Gene names | Mia2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma inhibitory activity protein 2 precursor. | |||||
|
KIF3B_MOUSE
|
||||||
| NC score | 0.009730 (rank : 103) | θ value | 5.27518 (rank : 96) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 533 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q61771 | Gene names | Kif3b | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF3B (Microtubule plus end-directed kinesin motor 3B). | |||||
|
AP3B1_HUMAN
|
||||||
| NC score | 0.008946 (rank : 104) | θ value | 3.0926 (rank : 75) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O00203, O00580, Q7Z393, Q9HD66 | Gene names | AP3B1, ADTB3A | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-1 (Adapter-related protein complex 3 beta-1 subunit) (Beta3A-adaptin) (Adaptor protein complex AP-3 beta-1 subunit) (Clathrin assembly protein complex 3 beta-1 large chain). | |||||
|
CN037_HUMAN
|
||||||
| NC score | 0.008843 (rank : 105) | θ value | 8.99809 (rank : 113) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q86TY3, Q6P5Q1, Q86TY1 | Gene names | C14orf37 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf37 precursor. | |||||
|
HIAL1_MOUSE
|
||||||
| NC score | 0.008640 (rank : 106) | θ value | 8.99809 (rank : 116) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CIA9, Q8VCW0 | Gene names | Hiatl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hippocampus abundant transcript-like protein 1. | |||||
|
HIAL1_HUMAN
|
||||||
| NC score | 0.008571 (rank : 107) | θ value | 8.99809 (rank : 115) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5SR56, Q3KQT4, Q53GU5, Q8WU95, Q96SM4 | Gene names | HIATL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hippocampus abundant transcript-like protein 1. | |||||
|
WDR43_HUMAN
|
||||||
| NC score | 0.008247 (rank : 108) | θ value | 6.88961 (rank : 108) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q15061, Q15395, Q92577 | Gene names | WDR43, KIAA0007 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 43. | |||||
|
TPR_HUMAN
|
||||||
| NC score | 0.007880 (rank : 109) | θ value | 0.813845 (rank : 51) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
ATAD2_HUMAN
|
||||||
| NC score | 0.007397 (rank : 110) | θ value | 4.03905 (rank : 84) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 322 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6PL18, Q658P2, Q68CQ0, Q6PJV6, Q8N890, Q9UHS5 | Gene names | ATAD2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATPase family AAA domain-containing protein 2. | |||||
|
CCD39_MOUSE
|
||||||
| NC score | 0.007369 (rank : 111) | θ value | 5.27518 (rank : 94) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1026 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9D5Y1, Q8CDG6 | Gene names | Ccdc39, D3Ertd789e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 39. | |||||
|
CALR_MOUSE
|
||||||
| NC score | 0.007138 (rank : 112) | θ value | 6.88961 (rank : 100) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P14211 | Gene names | Calr | |||
|
Domain Architecture |

|
|||||
| Description | Calreticulin precursor (CRP55) (Calregulin) (HACBP) (ERp60). | |||||
|
NFM_MOUSE
|
||||||
| NC score | 0.007094 (rank : 113) | θ value | 0.813845 (rank : 50) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1159 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P08553, Q61961 | Gene names | Nef3, Nefm, Nfm | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M). | |||||
|
CLMN_HUMAN
|
||||||
| NC score | 0.006997 (rank : 114) | θ value | 4.03905 (rank : 85) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q96JQ2, Q9H713, Q9HA23, Q9HA57, Q9UFP4, Q9ULN2 | Gene names | CLMN, KIAA1188 | |||
|
Domain Architecture |

|
|||||
| Description | Calmin. | |||||
|
RAD50_HUMAN
|
||||||
| NC score | 0.006755 (rank : 115) | θ value | 4.03905 (rank : 89) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1317 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q92878, O43254, Q6GMT7, Q6P5X3, Q9UP86 | Gene names | RAD50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (hRAD50). | |||||
|
RBMX2_HUMAN
|
||||||
| NC score | 0.006173 (rank : 116) | θ value | 4.03905 (rank : 90) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y388, Q5JY82, Q9Y3I8 | Gene names | RBMX2 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding motif protein, X-linked 2. | |||||
|
TRI44_HUMAN
|
||||||
| NC score | 0.005672 (rank : 117) | θ value | 8.99809 (rank : 124) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 393 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q96DX7, Q96QY2, Q9UGK0 | Gene names | TRIM44, DIPB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 44 (Protein DIPB). | |||||
|
PELP1_MOUSE
|
||||||
| NC score | 0.005229 (rank : 118) | θ value | 8.99809 (rank : 120) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9DBD5, Q5F2E2, Q6PEM0, Q91YM9 | Gene names | Pelp1, Mnar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor). | |||||
|
NFM_HUMAN
|
||||||
| NC score | 0.005194 (rank : 119) | θ value | 1.38821 (rank : 57) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1328 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P07197 | Gene names | NEF3, NEFM, NFM | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M) (Neurofilament 3). | |||||
|
PTN13_MOUSE
|
||||||
| NC score | 0.005121 (rank : 120) | θ value | 1.81305 (rank : 65) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 426 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q64512, Q61494, Q62135, Q64499 | Gene names | Ptpn13, Ptp14 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 13 (EC 3.1.3.48) (Protein tyrosine phosphatase PTP-BL) (Protein-tyrosine phosphatase RIP) (protein tyrosine phosphatase DPZPTP) (PTP36). | |||||
|
TRI26_MOUSE
|
||||||
| NC score | 0.003784 (rank : 121) | θ value | 3.0926 (rank : 81) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 339 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q99PN3, Q8C9E9, Q8JZT7, Q99PN2 | Gene names | Trim26, Znf173 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 26 (Zinc finger protein 173). | |||||
|
PEPL_HUMAN
|
||||||
| NC score | 0.003745 (rank : 122) | θ value | 8.99809 (rank : 121) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1141 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O60437, O60314, O60454 | Gene names | PPL, KIAA0568 | |||
|
Domain Architecture |

|
|||||
| Description | Periplakin (195 kDa cornified envelope precursor protein) (190 kDa paraneoplastic pemphigus antigen). | |||||
|
CAC1D_HUMAN
|
||||||
| NC score | 0.001875 (rank : 123) | θ value | 8.99809 (rank : 110) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q01668, Q13916, Q13931 | Gene names | CACNA1D, CACH3, CACN4, CACNL1A2, CCHL1A2 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1D (Voltage- gated calcium channel subunit alpha Cav1.3) (Calcium channel, L type, alpha-1 polypeptide, isoform 2). | |||||
|
PSYR_MOUSE
|
||||||
| NC score | -0.001377 (rank : 124) | θ value | 8.99809 (rank : 122) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 780 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61038, Q921N3 | Gene names | Gpr65, Gpcr25, Tdag8 | |||
|
Domain Architecture |

|
|||||
| Description | Psychosine receptor (G-protein coupled receptor 65) (T cell death- associated protein 8). | |||||