Please be patient as the page loads
|
SENP6_HUMAN
|
||||||
| SwissProt Accessions | Q9GZR1, O94891, Q8TBY4, Q9UJV5 | Gene names | SENP6, KIAA0797, SSP1, SUSP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 6 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP6) (SUMO-1-specific protease 1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
SENP6_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9GZR1, O94891, Q8TBY4, Q9UJV5 | Gene names | SENP6, KIAA0797, SSP1, SUSP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 6 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP6) (SUMO-1-specific protease 1). | |||||
|
SENP7_HUMAN
|
||||||
| θ value | 1.08223e-53 (rank : 2) | NC score | 0.771824 (rank : 2) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9BQF6, Q96PS5, Q9C0F6, Q9HBT5 | Gene names | SENP7, KIAA1707, SSP2, SUSP2 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 7 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP7) (SUMO-1-specific protease 2). | |||||
|
SENP2_MOUSE
|
||||||
| θ value | 3.18553e-13 (rank : 3) | NC score | 0.556761 (rank : 3) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q91ZX6, Q544T8, Q9D4Z0 | Gene names | Senp2, Smt3ip2 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 2 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP2) (SUMO-1/Smt3-specific isopeptidase 2) (Smt3ip2) (Axam2). | |||||
|
SENP1_HUMAN
|
||||||
| θ value | 3.52202e-12 (rank : 4) | NC score | 0.553209 (rank : 4) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9P0U3, Q86XC8 | Gene names | SENP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 1 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP1). | |||||
|
SENP1_MOUSE
|
||||||
| θ value | 7.84624e-12 (rank : 5) | NC score | 0.549693 (rank : 5) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P59110 | Gene names | Senp1 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 1 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP1). | |||||
|
SENP2_HUMAN
|
||||||
| θ value | 1.33837e-11 (rank : 6) | NC score | 0.547410 (rank : 6) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9HC62, Q96SR2, Q9P2L5 | Gene names | SENP2, KIAA1331 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 2 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP2) (SMT3-specific isopeptidase 2) (Smt3ip2) (Axam2). | |||||
|
SENP3_HUMAN
|
||||||
| θ value | 2.61198e-07 (rank : 7) | NC score | 0.481334 (rank : 8) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9H4L4, Q96PS4, Q9Y3W9 | Gene names | SENP3, SSP3, SUSP3 | |||
|
Domain Architecture |
|
|||||
| Description | Sentrin-specific protease 3 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP3) (SUMO-1-specific protease 3). | |||||
|
SENP3_MOUSE
|
||||||
| θ value | 2.61198e-07 (rank : 8) | NC score | 0.479389 (rank : 9) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9EP97 | Gene names | Senp3, Smt3ip, Smt3ip1 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 3 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP3) (SUMO-1-specific protease 3) (Smt3-specific isopeptidase 1) (Smt3ip1). | |||||
|
SENP5_HUMAN
|
||||||
| θ value | 3.41135e-07 (rank : 9) | NC score | 0.489656 (rank : 7) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96HI0, Q96SA5 | Gene names | SENP5 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 5 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP5). | |||||
|
IF2P_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 10) | NC score | 0.047204 (rank : 20) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 889 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O60841, O95805, Q9UF81, Q9UMN7 | Gene names | EIF5B, IF2, KIAA0741 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 5B (eIF-5B) (Translation initiation factor IF-2). | |||||
|
MAP2_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 11) | NC score | 0.056885 (rank : 16) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 348 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P11137, Q99975, Q99976 | Gene names | MAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 2 (MAP 2) (MAP-2). | |||||
|
DDX23_HUMAN
|
||||||
| θ value | 0.279714 (rank : 12) | NC score | 0.018047 (rank : 44) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BUQ8, O43188 | Gene names | DDX23 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX23 (EC 3.6.1.-) (DEAD box protein 23) (100 kDa U5 snRNP-specific protein) (U5-100kD) (PRP28 homolog). | |||||
|
SENP8_HUMAN
|
||||||
| θ value | 0.279714 (rank : 13) | NC score | 0.152415 (rank : 10) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96LD8, Q96QA4 | Gene names | SENP8, DEN1, NEDP1, PRSC2 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 8 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP8) (Protease, cysteine 2) (NEDD8-specific protease 1) (Deneddylase-1). | |||||
|
NCOR1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 14) | NC score | 0.039641 (rank : 26) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 577 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O75376, Q9UPV5, Q9UQ18 | Gene names | NCOR1, KIAA1047 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR). | |||||
|
CCD22_MOUSE
|
||||||
| θ value | 0.47712 (rank : 15) | NC score | 0.058947 (rank : 15) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 268 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9JIG7, Q8BYH4 | Gene names | Ccdc22, DXImx40e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 22. | |||||
|
VPS72_HUMAN
|
||||||
| θ value | 0.47712 (rank : 16) | NC score | 0.077842 (rank : 13) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q15906, Q53GJ2, Q5U0R4 | Gene names | VPS72, TCFL1, YL1 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 72 homolog (Transcription factor-like 1) (Protein YL-1). | |||||
|
VPS72_MOUSE
|
||||||
| θ value | 0.47712 (rank : 17) | NC score | 0.078308 (rank : 12) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q62481, Q3U2N4, Q810A9, Q99K81 | Gene names | Vps72, Tcfl1, Yl1 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 72 homolog (Transcription factor-like 1) (Protein YL-1). | |||||
|
PHAR2_HUMAN
|
||||||
| θ value | 0.62314 (rank : 18) | NC score | 0.046083 (rank : 23) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O75167, Q68DM2 | Gene names | PHACTR2, C6orf56, KIAA0680 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatase and actin regulator 2. | |||||
|
AP3B2_MOUSE
|
||||||
| θ value | 0.813845 (rank : 19) | NC score | 0.034164 (rank : 29) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9JME5, Q6QR53, Q8R1E5 | Gene names | Ap3b2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-3 complex subunit beta-2 (Adapter-related protein complex 3 beta-2 subunit) (Beta3B-adaptin) (Adaptor protein complex AP-3 beta-2 subunit) (Clathrin assembly protein complex 3 beta-2 large chain). | |||||
|
MEPE_HUMAN
|
||||||
| θ value | 0.813845 (rank : 20) | NC score | 0.064754 (rank : 14) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NQ76 | Gene names | MEPE | |||
|
Domain Architecture |
|
|||||
| Description | Matrix extracellular phosphoglycoprotein precursor (Osteoblast/osteocyte factor 45) (OF45). | |||||
|
BPA1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 21) | NC score | 0.014253 (rank : 49) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q91ZU6, Q91ZU7 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 1/2/3/4 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
CCD22_HUMAN
|
||||||
| θ value | 1.38821 (rank : 22) | NC score | 0.055209 (rank : 17) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O60826 | Gene names | CCDC22, CXorf37 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 22. | |||||
|
CD014_HUMAN
|
||||||
| θ value | 1.38821 (rank : 23) | NC score | 0.041779 (rank : 24) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NC60, Q8N7L6, Q9BSQ9 | Gene names | C4orf14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C4orf14. | |||||
|
AR13B_HUMAN
|
||||||
| θ value | 1.81305 (rank : 24) | NC score | 0.020891 (rank : 42) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 528 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q3SXY8, Q504W8 | Gene names | ARL13B, ARL2L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADP-ribosylation factor-like protein 13B (ADP-ribosylation factor-like protein 2-like 1) (ARL2-like protein 1). | |||||
|
CCNB3_MOUSE
|
||||||
| θ value | 1.81305 (rank : 25) | NC score | 0.028124 (rank : 35) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q810T2, Q810T3, Q8VDC8 | Gene names | Ccnb3, Cycb3 | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-B3. | |||||
|
PHF14_HUMAN
|
||||||
| θ value | 1.81305 (rank : 26) | NC score | 0.031723 (rank : 31) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O94880 | Gene names | PHF14, KIAA0783 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 14. | |||||
|
AP2B_HUMAN
|
||||||
| θ value | 2.36792 (rank : 27) | NC score | 0.049069 (rank : 19) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q92481, Q5JYX6, Q9NQ63, Q9NU99, Q9UJI7, Q9Y214, Q9Y3K3 | Gene names | TFAP2B | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor AP-2 beta (AP2-beta) (Activating enhancer-binding protein 2 beta). | |||||
|
AP2B_MOUSE
|
||||||
| θ value | 2.36792 (rank : 28) | NC score | 0.049070 (rank : 18) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q61313 | Gene names | Tfap2b, Tcfap2b | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor AP-2 beta (AP2-beta) (Activating enhancer-binding protein 2 beta). | |||||
|
DPOLZ_HUMAN
|
||||||
| θ value | 2.36792 (rank : 29) | NC score | 0.038915 (rank : 27) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O60673, O43214 | Gene names | REV3L, POLZ, REV3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase zeta catalytic subunit (EC 2.7.7.7) (hREV3). | |||||
|
DPOLZ_MOUSE
|
||||||
| θ value | 2.36792 (rank : 30) | NC score | 0.039699 (rank : 25) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61493, Q9JMD6, Q9QWX6 | Gene names | Rev3l, Polz, Sez4 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase zeta catalytic subunit (EC 2.7.7.7) (Seizure-related protein 4). | |||||
|
NFM_HUMAN
|
||||||
| θ value | 2.36792 (rank : 31) | NC score | 0.013247 (rank : 51) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1328 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P07197 | Gene names | NEF3, NEFM, NFM | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M) (Neurofilament 3). | |||||
|
NFM_MOUSE
|
||||||
| θ value | 2.36792 (rank : 32) | NC score | 0.014411 (rank : 48) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1159 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P08553, Q61961 | Gene names | Nef3, Nefm, Nfm | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M). | |||||
|
AMRP_HUMAN
|
||||||
| θ value | 3.0926 (rank : 33) | NC score | 0.031666 (rank : 32) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P30533 | Gene names | LRPAP1, A2MRAP | |||
|
Domain Architecture |
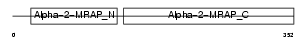
|
|||||
| Description | Alpha-2-macroglobulin receptor-associated protein precursor (Alpha-2- MRAP) (Low density lipoprotein receptor-related protein-associated protein 1) (RAP). | |||||
|
AP2A_HUMAN
|
||||||
| θ value | 3.0926 (rank : 34) | NC score | 0.046898 (rank : 22) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P05549, Q13777 | Gene names | TFAP2A, AP2TF, TFAP2 | |||
|
Domain Architecture |
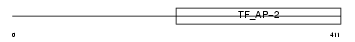
|
|||||
| Description | Transcription factor AP-2 alpha (AP2-alpha) (Activating enhancer- binding protein 2 alpha) (AP-2 transcription factor) (Activator protein 2) (AP-2). | |||||
|
AP2A_MOUSE
|
||||||
| θ value | 3.0926 (rank : 35) | NC score | 0.046947 (rank : 21) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P34056, Q60740, Q60741, Q60742, Q60743, Q62067, Q62068, Q62069, Q91VX0, Q9CRY4 | Gene names | Tfap2a, Ap2tf, Tcfap2a | |||
|
Domain Architecture |
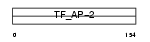
|
|||||
| Description | Transcription factor AP-2 alpha (AP2-alpha) (Activating enhancer- binding protein 2 alpha) (Activator protein 2) (AP-2). | |||||
|
FA50A_HUMAN
|
||||||
| θ value | 3.0926 (rank : 36) | NC score | 0.029918 (rank : 34) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q14320, Q5HY37 | Gene names | FAM50A, DXS9928E, HXC26, XAP5 | |||
|
Domain Architecture |
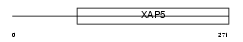
|
|||||
| Description | Protein FAM50A (Protein XAP-5) (Protein HXC-26). | |||||
|
MYH14_MOUSE
|
||||||
| θ value | 3.0926 (rank : 37) | NC score | 0.008618 (rank : 59) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1225 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6URW6, Q80V64, Q80ZE6 | Gene names | Myh14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-14 (Myosin heavy chain, nonmuscle IIc) (Nonmuscle myosin heavy chain IIc) (NMHC II-C). | |||||
|
SENP8_MOUSE
|
||||||
| θ value | 3.0926 (rank : 38) | NC score | 0.127520 (rank : 11) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9D2Z4, Q543P0 | Gene names | Senp8, Den1, Nedp1 | |||
|
Domain Architecture |
|
|||||
| Description | Sentrin-specific protease 8 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP8) (NEDD8-specific protease 1) (Deneddylase-1). | |||||
|
BMP5_HUMAN
|
||||||
| θ value | 4.03905 (rank : 39) | NC score | 0.009526 (rank : 58) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P22003, Q9H547, Q9NTM5 | Gene names | BMP5 | |||
|
Domain Architecture |

|
|||||
| Description | Bone morphogenetic protein 5 precursor (BMP-5). | |||||
|
FA50A_MOUSE
|
||||||
| θ value | 4.03905 (rank : 40) | NC score | 0.027231 (rank : 36) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 293 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9WV03, Q9D866 | Gene names | Fam50a, D0HXS9928E, Xap5 | |||
|
Domain Architecture |
|
|||||
| Description | Protein FAM50A (XAP-5 protein). | |||||
|
PHF14_MOUSE
|
||||||
| θ value | 4.03905 (rank : 41) | NC score | 0.025415 (rank : 38) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9D4H9, Q810Z6, Q8C284, Q8CGF8, Q9D1Q8 | Gene names | Phf14 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 14. | |||||
|
ADA30_HUMAN
|
||||||
| θ value | 5.27518 (rank : 42) | NC score | 0.008518 (rank : 60) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 391 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UKF2, Q9UKF1 | Gene names | ADAM30 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 30 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 30). | |||||
|
CBX2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 43) | NC score | 0.018697 (rank : 43) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P30658, O35731, Q8CIA0 | Gene names | Cbx2, M33 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 2 (Modifier 3 protein) (M33). | |||||
|
DC1I2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 44) | NC score | 0.013693 (rank : 50) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13409, Q96NG7, Q96S87, Q9BXZ5, Q9NT58 | Gene names | DYNC1I2, DNCI2, DNCIC2 | |||
|
Domain Architecture |

|
|||||
| Description | Cytoplasmic dynein 1 intermediate chain 2 (Dynein intermediate chain 2, cytosolic) (DH IC-2) (Cytoplasmic dynein intermediate chain 2). | |||||
|
NEDD4_MOUSE
|
||||||
| θ value | 5.27518 (rank : 45) | NC score | 0.007806 (rank : 62) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P46935, O08758, Q3UZI2, Q8BGB3 | Gene names | Nedd4, Kiaa0093, Nedd-4, Nedd4a | |||
|
Domain Architecture |

|
|||||
| Description | E3 ubiquitin-protein ligase NEDD4 (EC 6.3.2.-) (Neural precursor cell expressed developmentally down-regulated protein 4). | |||||
|
PIWL4_MOUSE
|
||||||
| θ value | 5.27518 (rank : 46) | NC score | 0.012987 (rank : 52) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CGT6, Q8CC75 | Gene names | Piwil4, Miwi2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Piwi-like protein 4 (mAgo5). | |||||
|
SIAL_HUMAN
|
||||||
| θ value | 5.27518 (rank : 47) | NC score | 0.029985 (rank : 33) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P21815 | Gene names | IBSP, BNSP | |||
|
Domain Architecture |
|
|||||
| Description | Bone sialoprotein-2 precursor (Bone sialoprotein II) (BSP II) (Cell- binding sialoprotein) (Integrin-binding sialoprotein). | |||||
|
TRDN_HUMAN
|
||||||
| θ value | 5.27518 (rank : 48) | NC score | 0.022018 (rank : 41) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q13061 | Gene names | TRDN | |||
|
Domain Architecture |

|
|||||
| Description | Triadin. | |||||
|
ZN384_HUMAN
|
||||||
| θ value | 5.27518 (rank : 49) | NC score | -0.000212 (rank : 69) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8TF68, O15407, Q8N938 | Gene names | ZNF384, CAGH1, CIZ, NMP4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 384 (Nuclear matrix transcription factor 4) (Nuclear matrix protein 4) (CAG repeat protein 1) (CAS-interacting zinc finger protein). | |||||
|
CSKI1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 50) | NC score | 0.004010 (rank : 67) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1031 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6P9K8, Q6ZPU2, Q8BWU2, Q8BX99, Q9CXH0 | Gene names | Caskin1, Kiaa1306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-1 (CASK-interacting protein 1). | |||||
|
DNL4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 51) | NC score | 0.016384 (rank : 47) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BTF7, Q3UG76 | Gene names | Lig4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA ligase 4 (EC 6.5.1.1) (DNA ligase IV) (Polydeoxyribonucleotide synthase [ATP] 4). | |||||
|
DSG2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 52) | NC score | 0.004167 (rank : 66) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q14126 | Gene names | DSG2 | |||
|
Domain Architecture |

|
|||||
| Description | Desmoglein-2 precursor (HDGC). | |||||
|
DSPP_MOUSE
|
||||||
| θ value | 6.88961 (rank : 53) | NC score | 0.031803 (rank : 30) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
RNPS1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 54) | NC score | 0.024523 (rank : 39) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15287, O75308, Q8WY42, Q9NYG3 | Gene names | RNPS1, LDC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein with serine-rich domain 1 (SR-related protein LDC2). | |||||
|
RNPS1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 55) | NC score | 0.024523 (rank : 40) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99M28, Q3TMJ1, Q922H8 | Gene names | Rnps1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein with serine-rich domain 1. | |||||
|
CCD82_MOUSE
|
||||||
| θ value | 8.99809 (rank : 56) | NC score | 0.016821 (rank : 46) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6PG04, Q3V462 | Gene names | Ccdc82 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 82. | |||||
|
CENPB_HUMAN
|
||||||
| θ value | 8.99809 (rank : 57) | NC score | 0.017665 (rank : 45) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P07199 | Gene names | CENPB | |||
|
Domain Architecture |

|
|||||
| Description | Major centromere autoantigen B (Centromere protein B) (CENP-B). | |||||
|
CYLC1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 58) | NC score | 0.035173 (rank : 28) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P35663, Q5JQQ9 | Gene names | CYLC1, CYL, CYL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-1 (Cylicin I) (Multiple-band polypeptide I). | |||||
|
DMP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 59) | NC score | 0.025443 (rank : 37) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
FOG1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 60) | NC score | 0.001942 (rank : 68) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1237 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O35615 | Gene names | Zfpm1, Fog, Fog1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZFPM1 (Zinc finger protein multitype 1) (Friend of GATA protein 1) (Friend of GATA-1) (FOG-1). | |||||
|
GOGA3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 61) | NC score | 0.008151 (rank : 61) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1437 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P55937, Q80VF5, Q8CCK4, Q9QYT2, Q9QYT3 | Gene names | Golga3, Mea2 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 3 (Golgin-160) (Male-enhanced antigen 2) (MEA-2). | |||||
|
KELL_HUMAN
|
||||||
| θ value | 8.99809 (rank : 62) | NC score | 0.009942 (rank : 57) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P23276, Q96RS8, Q99885 | Gene names | KEL | |||
|
Domain Architecture |

|
|||||
| Description | Kell blood group glycoprotein (EC 3.4.24.-) (CD238 antigen). | |||||
|
MDR1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 63) | NC score | 0.004255 (rank : 65) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P06795 | Gene names | Abcb1, Abcb1b, Mdr1, Mdr1b, Pgy1, Pgy1-1 | |||
|
Domain Architecture |

|
|||||
| Description | Multidrug resistance protein 1 (EC 3.6.3.44) (ATP-binding cassette sub-family B member 1) (P-glycoprotein 1) (CD243 antigen). | |||||
|
MTMR3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 64) | NC score | 0.007162 (rank : 63) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13615, Q9NYN5, Q9NYN6, Q9UDX6, Q9UEG3 | Gene names | MTMR3, KIAA0371, ZFYVE10 | |||
|
Domain Architecture |

|
|||||
| Description | Myotubularin-related protein 3 (EC 3.1.3.48) (FYVE domain-containing dual specificity protein phosphatase 1) (FYVE-DSP1) (Zinc finger FYVE domain-containing protein 10). | |||||
|
MYH14_HUMAN
|
||||||
| θ value | 8.99809 (rank : 65) | NC score | 0.006959 (rank : 64) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 995 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q7Z406, Q5CZ75, Q6XYE4, Q76B62, Q8WV23, Q96I22, Q9BT27, Q9BW35, Q9H882 | Gene names | MYH14, KIAA2034 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-14 (Myosin heavy chain, nonmuscle IIc) (Nonmuscle myosin heavy chain IIc) (NMHC II-C). | |||||
|
NRAP_MOUSE
|
||||||
| θ value | 8.99809 (rank : 66) | NC score | 0.010875 (rank : 55) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80XB4, O35884, Q3UTY7, Q3UU10, Q80V40 | Gene names | Nrap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nebulin-related anchoring protein (N-RAP). | |||||
|
PCLO_HUMAN
|
||||||
| θ value | 8.99809 (rank : 67) | NC score | 0.010080 (rank : 56) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
RECQ4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 68) | NC score | 0.011593 (rank : 54) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q75NR7, Q76MT1, Q99PV9 | Gene names | Recql4, Recq4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent DNA helicase Q4 (EC 3.6.1.-) (RecQ protein-like 4). | |||||
|
TOB1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 69) | NC score | 0.011705 (rank : 53) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q61471 | Gene names | Tob1, Tob, Trob | |||
|
Domain Architecture |

|
|||||
| Description | Tob1 protein (Transducer of erbB-2 1). | |||||
|
SENP6_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9GZR1, O94891, Q8TBY4, Q9UJV5 | Gene names | SENP6, KIAA0797, SSP1, SUSP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 6 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP6) (SUMO-1-specific protease 1). | |||||
|
SENP7_HUMAN
|
||||||
| NC score | 0.771824 (rank : 2) | θ value | 1.08223e-53 (rank : 2) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9BQF6, Q96PS5, Q9C0F6, Q9HBT5 | Gene names | SENP7, KIAA1707, SSP2, SUSP2 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 7 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP7) (SUMO-1-specific protease 2). | |||||
|
SENP2_MOUSE
|
||||||
| NC score | 0.556761 (rank : 3) | θ value | 3.18553e-13 (rank : 3) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q91ZX6, Q544T8, Q9D4Z0 | Gene names | Senp2, Smt3ip2 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 2 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP2) (SUMO-1/Smt3-specific isopeptidase 2) (Smt3ip2) (Axam2). | |||||
|
SENP1_HUMAN
|
||||||
| NC score | 0.553209 (rank : 4) | θ value | 3.52202e-12 (rank : 4) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9P0U3, Q86XC8 | Gene names | SENP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 1 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP1). | |||||
|
SENP1_MOUSE
|
||||||
| NC score | 0.549693 (rank : 5) | θ value | 7.84624e-12 (rank : 5) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P59110 | Gene names | Senp1 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 1 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP1). | |||||
|
SENP2_HUMAN
|
||||||
| NC score | 0.547410 (rank : 6) | θ value | 1.33837e-11 (rank : 6) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9HC62, Q96SR2, Q9P2L5 | Gene names | SENP2, KIAA1331 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 2 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP2) (SMT3-specific isopeptidase 2) (Smt3ip2) (Axam2). | |||||
|
SENP5_HUMAN
|
||||||
| NC score | 0.489656 (rank : 7) | θ value | 3.41135e-07 (rank : 9) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96HI0, Q96SA5 | Gene names | SENP5 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 5 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP5). | |||||
|
SENP3_HUMAN
|
||||||
| NC score | 0.481334 (rank : 8) | θ value | 2.61198e-07 (rank : 7) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9H4L4, Q96PS4, Q9Y3W9 | Gene names | SENP3, SSP3, SUSP3 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 3 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP3) (SUMO-1-specific protease 3). | |||||
|
SENP3_MOUSE
|
||||||
| NC score | 0.479389 (rank : 9) | θ value | 2.61198e-07 (rank : 8) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9EP97 | Gene names | Senp3, Smt3ip, Smt3ip1 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 3 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP3) (SUMO-1-specific protease 3) (Smt3-specific isopeptidase 1) (Smt3ip1). | |||||
|
SENP8_HUMAN
|
||||||
| NC score | 0.152415 (rank : 10) | θ value | 0.279714 (rank : 13) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96LD8, Q96QA4 | Gene names | SENP8, DEN1, NEDP1, PRSC2 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 8 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP8) (Protease, cysteine 2) (NEDD8-specific protease 1) (Deneddylase-1). | |||||
|
SENP8_MOUSE
|
||||||
| NC score | 0.127520 (rank : 11) | θ value | 3.0926 (rank : 38) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9D2Z4, Q543P0 | Gene names | Senp8, Den1, Nedp1 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 8 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP8) (NEDD8-specific protease 1) (Deneddylase-1). | |||||
|
VPS72_MOUSE
|
||||||
| NC score | 0.078308 (rank : 12) | θ value | 0.47712 (rank : 17) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q62481, Q3U2N4, Q810A9, Q99K81 | Gene names | Vps72, Tcfl1, Yl1 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 72 homolog (Transcription factor-like 1) (Protein YL-1). | |||||
|
VPS72_HUMAN
|
||||||
| NC score | 0.077842 (rank : 13) | θ value | 0.47712 (rank : 16) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q15906, Q53GJ2, Q5U0R4 | Gene names | VPS72, TCFL1, YL1 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 72 homolog (Transcription factor-like 1) (Protein YL-1). | |||||
|
MEPE_HUMAN
|
||||||
| NC score | 0.064754 (rank : 14) | θ value | 0.813845 (rank : 20) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NQ76 | Gene names | MEPE | |||
|
Domain Architecture |

|
|||||
| Description | Matrix extracellular phosphoglycoprotein precursor (Osteoblast/osteocyte factor 45) (OF45). | |||||
|
CCD22_MOUSE
|
||||||
| NC score | 0.058947 (rank : 15) | θ value | 0.47712 (rank : 15) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 268 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9JIG7, Q8BYH4 | Gene names | Ccdc22, DXImx40e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 22. | |||||
|
MAP2_HUMAN
|
||||||
| NC score | 0.056885 (rank : 16) | θ value | 0.0961366 (rank : 11) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 348 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P11137, Q99975, Q99976 | Gene names | MAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 2 (MAP 2) (MAP-2). | |||||
|
CCD22_HUMAN
|
||||||
| NC score | 0.055209 (rank : 17) | θ value | 1.38821 (rank : 22) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O60826 | Gene names | CCDC22, CXorf37 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 22. | |||||
|
AP2B_MOUSE
|
||||||
| NC score | 0.049070 (rank : 18) | θ value | 2.36792 (rank : 28) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q61313 | Gene names | Tfap2b, Tcfap2b | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor AP-2 beta (AP2-beta) (Activating enhancer-binding protein 2 beta). | |||||
|
AP2B_HUMAN
|
||||||
| NC score | 0.049069 (rank : 19) | θ value | 2.36792 (rank : 27) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q92481, Q5JYX6, Q9NQ63, Q9NU99, Q9UJI7, Q9Y214, Q9Y3K3 | Gene names | TFAP2B | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor AP-2 beta (AP2-beta) (Activating enhancer-binding protein 2 beta). | |||||
|
IF2P_HUMAN
|
||||||
| NC score | 0.047204 (rank : 20) | θ value | 0.0736092 (rank : 10) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 889 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O60841, O95805, Q9UF81, Q9UMN7 | Gene names | EIF5B, IF2, KIAA0741 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 5B (eIF-5B) (Translation initiation factor IF-2). | |||||
|
AP2A_MOUSE
|
||||||
| NC score | 0.046947 (rank : 21) | θ value | 3.0926 (rank : 35) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P34056, Q60740, Q60741, Q60742, Q60743, Q62067, Q62068, Q62069, Q91VX0, Q9CRY4 | Gene names | Tfap2a, Ap2tf, Tcfap2a | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor AP-2 alpha (AP2-alpha) (Activating enhancer- binding protein 2 alpha) (Activator protein 2) (AP-2). | |||||
|
AP2A_HUMAN
|
||||||
| NC score | 0.046898 (rank : 22) | θ value | 3.0926 (rank : 34) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P05549, Q13777 | Gene names | TFAP2A, AP2TF, TFAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor AP-2 alpha (AP2-alpha) (Activating enhancer- binding protein 2 alpha) (AP-2 transcription factor) (Activator protein 2) (AP-2). | |||||
|
PHAR2_HUMAN
|
||||||
| NC score | 0.046083 (rank : 23) | θ value | 0.62314 (rank : 18) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O75167, Q68DM2 | Gene names | PHACTR2, C6orf56, KIAA0680 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatase and actin regulator 2. | |||||
|
CD014_HUMAN
|
||||||
| NC score | 0.041779 (rank : 24) | θ value | 1.38821 (rank : 23) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NC60, Q8N7L6, Q9BSQ9 | Gene names | C4orf14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C4orf14. | |||||
|
DPOLZ_MOUSE
|
||||||
| NC score | 0.039699 (rank : 25) | θ value | 2.36792 (rank : 30) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61493, Q9JMD6, Q9QWX6 | Gene names | Rev3l, Polz, Sez4 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase zeta catalytic subunit (EC 2.7.7.7) (Seizure-related protein 4). | |||||
|
NCOR1_HUMAN
|
||||||
| NC score | 0.039641 (rank : 26) | θ value | 0.365318 (rank : 14) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 577 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O75376, Q9UPV5, Q9UQ18 | Gene names | NCOR1, KIAA1047 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR). | |||||
|
DPOLZ_HUMAN
|
||||||
| NC score | 0.038915 (rank : 27) | θ value | 2.36792 (rank : 29) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O60673, O43214 | Gene names | REV3L, POLZ, REV3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase zeta catalytic subunit (EC 2.7.7.7) (hREV3). | |||||
|
CYLC1_HUMAN
|
||||||
| NC score | 0.035173 (rank : 28) | θ value | 8.99809 (rank : 58) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P35663, Q5JQQ9 | Gene names | CYLC1, CYL, CYL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-1 (Cylicin I) (Multiple-band polypeptide I). | |||||
|
AP3B2_MOUSE
|
||||||
| NC score | 0.034164 (rank : 29) | θ value | 0.813845 (rank : 19) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9JME5, Q6QR53, Q8R1E5 | Gene names | Ap3b2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-3 complex subunit beta-2 (Adapter-related protein complex 3 beta-2 subunit) (Beta3B-adaptin) (Adaptor protein complex AP-3 beta-2 subunit) (Clathrin assembly protein complex 3 beta-2 large chain). | |||||
|
DSPP_MOUSE
|
||||||
| NC score | 0.031803 (rank : 30) | θ value | 6.88961 (rank : 53) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
PHF14_HUMAN
|
||||||
| NC score | 0.031723 (rank : 31) | θ value | 1.81305 (rank : 26) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O94880 | Gene names | PHF14, KIAA0783 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 14. | |||||
|
AMRP_HUMAN
|
||||||
| NC score | 0.031666 (rank : 32) | θ value | 3.0926 (rank : 33) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P30533 | Gene names | LRPAP1, A2MRAP | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-2-macroglobulin receptor-associated protein precursor (Alpha-2- MRAP) (Low density lipoprotein receptor-related protein-associated protein 1) (RAP). | |||||
|
SIAL_HUMAN
|
||||||
| NC score | 0.029985 (rank : 33) | θ value | 5.27518 (rank : 47) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P21815 | Gene names | IBSP, BNSP | |||
|
Domain Architecture |

|
|||||
| Description | Bone sialoprotein-2 precursor (Bone sialoprotein II) (BSP II) (Cell- binding sialoprotein) (Integrin-binding sialoprotein). | |||||
|
FA50A_HUMAN
|
||||||
| NC score | 0.029918 (rank : 34) | θ value | 3.0926 (rank : 36) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q14320, Q5HY37 | Gene names | FAM50A, DXS9928E, HXC26, XAP5 | |||
|
Domain Architecture |

|
|||||
| Description | Protein FAM50A (Protein XAP-5) (Protein HXC-26). | |||||
|
CCNB3_MOUSE
|
||||||
| NC score | 0.028124 (rank : 35) | θ value | 1.81305 (rank : 25) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q810T2, Q810T3, Q8VDC8 | Gene names | Ccnb3, Cycb3 | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-B3. | |||||
|
FA50A_MOUSE
|
||||||
| NC score | 0.027231 (rank : 36) | θ value | 4.03905 (rank : 40) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 293 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9WV03, Q9D866 | Gene names | Fam50a, D0HXS9928E, Xap5 | |||
|
Domain Architecture |

|
|||||
| Description | Protein FAM50A (XAP-5 protein). | |||||
|
DMP1_HUMAN
|
||||||
| NC score | 0.025443 (rank : 37) | θ value | 8.99809 (rank : 59) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
PHF14_MOUSE
|
||||||
| NC score | 0.025415 (rank : 38) | θ value | 4.03905 (rank : 41) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9D4H9, Q810Z6, Q8C284, Q8CGF8, Q9D1Q8 | Gene names | Phf14 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 14. | |||||
|
RNPS1_HUMAN
|
||||||
| NC score | 0.024523 (rank : 39) | θ value | 6.88961 (rank : 54) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15287, O75308, Q8WY42, Q9NYG3 | Gene names | RNPS1, LDC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein with serine-rich domain 1 (SR-related protein LDC2). | |||||
|
RNPS1_MOUSE
|
||||||
| NC score | 0.024523 (rank : 40) | θ value | 6.88961 (rank : 55) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99M28, Q3TMJ1, Q922H8 | Gene names | Rnps1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein with serine-rich domain 1. | |||||
|
TRDN_HUMAN
|
||||||
| NC score | 0.022018 (rank : 41) | θ value | 5.27518 (rank : 48) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q13061 | Gene names | TRDN | |||
|
Domain Architecture |

|
|||||
| Description | Triadin. | |||||
|
AR13B_HUMAN
|
||||||
| NC score | 0.020891 (rank : 42) | θ value | 1.81305 (rank : 24) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 528 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q3SXY8, Q504W8 | Gene names | ARL13B, ARL2L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADP-ribosylation factor-like protein 13B (ADP-ribosylation factor-like protein 2-like 1) (ARL2-like protein 1). | |||||
|
CBX2_MOUSE
|
||||||
| NC score | 0.018697 (rank : 43) | θ value | 5.27518 (rank : 43) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P30658, O35731, Q8CIA0 | Gene names | Cbx2, M33 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 2 (Modifier 3 protein) (M33). | |||||
|
DDX23_HUMAN
|
||||||
| NC score | 0.018047 (rank : 44) | θ value | 0.279714 (rank : 12) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BUQ8, O43188 | Gene names | DDX23 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX23 (EC 3.6.1.-) (DEAD box protein 23) (100 kDa U5 snRNP-specific protein) (U5-100kD) (PRP28 homolog). | |||||
|
CENPB_HUMAN
|
||||||
| NC score | 0.017665 (rank : 45) | θ value | 8.99809 (rank : 57) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P07199 | Gene names | CENPB | |||
|
Domain Architecture |

|
|||||
| Description | Major centromere autoantigen B (Centromere protein B) (CENP-B). | |||||
|
CCD82_MOUSE
|
||||||
| NC score | 0.016821 (rank : 46) | θ value | 8.99809 (rank : 56) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6PG04, Q3V462 | Gene names | Ccdc82 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 82. | |||||
|
DNL4_MOUSE
|
||||||
| NC score | 0.016384 (rank : 47) | θ value | 6.88961 (rank : 51) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BTF7, Q3UG76 | Gene names | Lig4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA ligase 4 (EC 6.5.1.1) (DNA ligase IV) (Polydeoxyribonucleotide synthase [ATP] 4). | |||||
|
NFM_MOUSE
|
||||||
| NC score | 0.014411 (rank : 48) | θ value | 2.36792 (rank : 32) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1159 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P08553, Q61961 | Gene names | Nef3, Nefm, Nfm | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M). | |||||
|
BPA1_MOUSE
|
||||||
| NC score | 0.014253 (rank : 49) | θ value | 1.38821 (rank : 21) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q91ZU6, Q91ZU7 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 1/2/3/4 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
DC1I2_HUMAN
|
||||||
| NC score | 0.013693 (rank : 50) | θ value | 5.27518 (rank : 44) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13409, Q96NG7, Q96S87, Q9BXZ5, Q9NT58 | Gene names | DYNC1I2, DNCI2, DNCIC2 | |||
|
Domain Architecture |

|
|||||
| Description | Cytoplasmic dynein 1 intermediate chain 2 (Dynein intermediate chain 2, cytosolic) (DH IC-2) (Cytoplasmic dynein intermediate chain 2). | |||||
|
NFM_HUMAN
|
||||||
| NC score | 0.013247 (rank : 51) | θ value | 2.36792 (rank : 31) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1328 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P07197 | Gene names | NEF3, NEFM, NFM | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M) (Neurofilament 3). | |||||
|
PIWL4_MOUSE
|
||||||
| NC score | 0.012987 (rank : 52) | θ value | 5.27518 (rank : 46) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CGT6, Q8CC75 | Gene names | Piwil4, Miwi2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Piwi-like protein 4 (mAgo5). | |||||
|
TOB1_MOUSE
|
||||||
| NC score | 0.011705 (rank : 53) | θ value | 8.99809 (rank : 69) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q61471 | Gene names | Tob1, Tob, Trob | |||
|
Domain Architecture |

|
|||||
| Description | Tob1 protein (Transducer of erbB-2 1). | |||||
|
RECQ4_MOUSE
|
||||||
| NC score | 0.011593 (rank : 54) | θ value | 8.99809 (rank : 68) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q75NR7, Q76MT1, Q99PV9 | Gene names | Recql4, Recq4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent DNA helicase Q4 (EC 3.6.1.-) (RecQ protein-like 4). | |||||
|
NRAP_MOUSE
|
||||||
| NC score | 0.010875 (rank : 55) | θ value | 8.99809 (rank : 66) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80XB4, O35884, Q3UTY7, Q3UU10, Q80V40 | Gene names | Nrap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nebulin-related anchoring protein (N-RAP). | |||||
|
PCLO_HUMAN
|
||||||
| NC score | 0.010080 (rank : 56) | θ value | 8.99809 (rank : 67) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
KELL_HUMAN
|
||||||
| NC score | 0.009942 (rank : 57) | θ value | 8.99809 (rank : 62) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P23276, Q96RS8, Q99885 | Gene names | KEL | |||
|
Domain Architecture |

|
|||||
| Description | Kell blood group glycoprotein (EC 3.4.24.-) (CD238 antigen). | |||||
|
BMP5_HUMAN
|
||||||
| NC score | 0.009526 (rank : 58) | θ value | 4.03905 (rank : 39) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P22003, Q9H547, Q9NTM5 | Gene names | BMP5 | |||
|
Domain Architecture |

|
|||||
| Description | Bone morphogenetic protein 5 precursor (BMP-5). | |||||
|
MYH14_MOUSE
|
||||||
| NC score | 0.008618 (rank : 59) | θ value | 3.0926 (rank : 37) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1225 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6URW6, Q80V64, Q80ZE6 | Gene names | Myh14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-14 (Myosin heavy chain, nonmuscle IIc) (Nonmuscle myosin heavy chain IIc) (NMHC II-C). | |||||
|
ADA30_HUMAN
|
||||||
| NC score | 0.008518 (rank : 60) | θ value | 5.27518 (rank : 42) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 391 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UKF2, Q9UKF1 | Gene names | ADAM30 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 30 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 30). | |||||
|
GOGA3_MOUSE
|
||||||
| NC score | 0.008151 (rank : 61) | θ value | 8.99809 (rank : 61) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1437 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P55937, Q80VF5, Q8CCK4, Q9QYT2, Q9QYT3 | Gene names | Golga3, Mea2 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 3 (Golgin-160) (Male-enhanced antigen 2) (MEA-2). | |||||
|
NEDD4_MOUSE
|
||||||
| NC score | 0.007806 (rank : 62) | θ value | 5.27518 (rank : 45) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P46935, O08758, Q3UZI2, Q8BGB3 | Gene names | Nedd4, Kiaa0093, Nedd-4, Nedd4a | |||
|
Domain Architecture |

|
|||||
| Description | E3 ubiquitin-protein ligase NEDD4 (EC 6.3.2.-) (Neural precursor cell expressed developmentally down-regulated protein 4). | |||||
|
MTMR3_HUMAN
|
||||||
| NC score | 0.007162 (rank : 63) | θ value | 8.99809 (rank : 64) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13615, Q9NYN5, Q9NYN6, Q9UDX6, Q9UEG3 | Gene names | MTMR3, KIAA0371, ZFYVE10 | |||
|
Domain Architecture |

|
|||||
| Description | Myotubularin-related protein 3 (EC 3.1.3.48) (FYVE domain-containing dual specificity protein phosphatase 1) (FYVE-DSP1) (Zinc finger FYVE domain-containing protein 10). | |||||
|
MYH14_HUMAN
|
||||||
| NC score | 0.006959 (rank : 64) | θ value | 8.99809 (rank : 65) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 995 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q7Z406, Q5CZ75, Q6XYE4, Q76B62, Q8WV23, Q96I22, Q9BT27, Q9BW35, Q9H882 | Gene names | MYH14, KIAA2034 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-14 (Myosin heavy chain, nonmuscle IIc) (Nonmuscle myosin heavy chain IIc) (NMHC II-C). | |||||
|
MDR1_MOUSE
|
||||||
| NC score | 0.004255 (rank : 65) | θ value | 8.99809 (rank : 63) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P06795 | Gene names | Abcb1, Abcb1b, Mdr1, Mdr1b, Pgy1, Pgy1-1 | |||
|
Domain Architecture |

|
|||||
| Description | Multidrug resistance protein 1 (EC 3.6.3.44) (ATP-binding cassette sub-family B member 1) (P-glycoprotein 1) (CD243 antigen). | |||||
|
DSG2_HUMAN
|
||||||
| NC score | 0.004167 (rank : 66) | θ value | 6.88961 (rank : 52) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q14126 | Gene names | DSG2 | |||
|
Domain Architecture |

|
|||||
| Description | Desmoglein-2 precursor (HDGC). | |||||
|
CSKI1_MOUSE
|
||||||
| NC score | 0.004010 (rank : 67) | θ value | 6.88961 (rank : 50) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1031 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6P9K8, Q6ZPU2, Q8BWU2, Q8BX99, Q9CXH0 | Gene names | Caskin1, Kiaa1306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-1 (CASK-interacting protein 1). | |||||
|
FOG1_MOUSE
|
||||||
| NC score | 0.001942 (rank : 68) | θ value | 8.99809 (rank : 60) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 1237 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O35615 | Gene names | Zfpm1, Fog, Fog1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZFPM1 (Zinc finger protein multitype 1) (Friend of GATA protein 1) (Friend of GATA-1) (FOG-1). | |||||
|
ZN384_HUMAN
|
||||||
| NC score | -0.000212 (rank : 69) | θ value | 5.27518 (rank : 49) | |||
| Query Neighborhood Hits | 69 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8TF68, O15407, Q8N938 | Gene names | ZNF384, CAGH1, CIZ, NMP4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 384 (Nuclear matrix transcription factor 4) (Nuclear matrix protein 4) (CAG repeat protein 1) (CAS-interacting zinc finger protein). | |||||