Please be patient as the page loads
|
PIGO_HUMAN
|
||||||
| SwissProt Accessions | Q8TEQ8, Q6P154, Q6UX80, Q8TDS8, Q96CS9, Q9BVN9, Q9Y4B0 | Gene names | PIGO | |||
|
Domain Architecture |

|
|||||
| Description | GPI ethanolamine phosphate transferase 3 (EC 2.-.-.-) (Phosphatidylinositol-glycan biosynthesis class O protein) (PIG-O). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
PIGO_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8TEQ8, Q6P154, Q6UX80, Q8TDS8, Q96CS9, Q9BVN9, Q9Y4B0 | Gene names | PIGO | |||
|
Domain Architecture |

|
|||||
| Description | GPI ethanolamine phosphate transferase 3 (EC 2.-.-.-) (Phosphatidylinositol-glycan biosynthesis class O protein) (PIG-O). | |||||
|
PIGO_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.998121 (rank : 2) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9JJI6, Q9CRY2 | Gene names | Pigo | |||
|
Domain Architecture |

|
|||||
| Description | GPI ethanolamine phosphate transferase 3 (EC 2.-.-.-) (Phosphatidylinositol-glycan biosynthesis class O protein) (PIG-O). | |||||
|
PIGG_HUMAN
|
||||||
| θ value | 1.01294e-51 (rank : 3) | NC score | 0.786584 (rank : 3) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5H8A4, Q2TAK5, Q6UX31, Q7L5Y4, Q8N866, Q8NCC9, Q96SY9, Q9BVT7, Q9NXG5 | Gene names | PIGG, GPI7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GPI ethanolamine phosphate transferase 2 (EC 2.-.-.-) (Phosphatidylinositol-glycan biosynthesis class G protein) (PIG-G) (GPI7 homolog) (hGPI7). | |||||
|
PIGN_MOUSE
|
||||||
| θ value | 1.69304e-06 (rank : 4) | NC score | 0.316970 (rank : 4) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9R1S3, Q3V0S6, Q8VCC3, Q9R1S1, Q9R1S2 | Gene names | Pign | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GPI ethanolamine phosphate transferase 1 (EC 2.-.-.-) (Phosphatidylinositol-glycan biosynthesis class N protein) (PIG-N). | |||||
|
PIGN_HUMAN
|
||||||
| θ value | 8.40245e-06 (rank : 5) | NC score | 0.302082 (rank : 5) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O95427, Q7L8F8, Q8TC01, Q9NT05 | Gene names | PIGN, MCD4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GPI ethanolamine phosphate transferase 1 (EC 2.-.-.-) (Phosphatidylinositol-glycan biosynthesis class N protein) (PIG-N) (MCD4 homolog). | |||||
|
ENPP7_HUMAN
|
||||||
| θ value | 0.000158464 (rank : 6) | NC score | 0.226173 (rank : 6) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q6UWV6, Q6ZTS5, Q8IUS8 | Gene names | ENPP7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ectonucleotide pyrophosphatase/phosphodiesterase 7 precursor (EC 3.1.4.12) (E-NPP7) (NPP-7) (Alkaline sphingomyelin phosphodiesterase) (Intestinal alkaline sphingomyelinase) (Alk-SMase). | |||||
|
ENPP2_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 7) | NC score | 0.169607 (rank : 11) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q13822, Q13827, Q15117 | Gene names | ENPP2, ATX, PDNP2 | |||
|
Domain Architecture |

|
|||||
| Description | Ectonucleotide pyrophosphatase/phosphodiesterase 2 (E-NPP 2) (Phosphodiesterase I/nucleotide pyrophosphatase 2) (Phosphodiesterase I alpha) (PD-Ialpha) (Autotaxin) [Includes: Alkaline phosphodiesterase I (EC 3.1.4.1); Nucleotide pyrophosphatase (EC 3.6.1.9) (NPPase)]. | |||||
|
IDS_MOUSE
|
||||||
| θ value | 0.00869519 (rank : 8) | NC score | 0.110488 (rank : 16) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q08890, Q3TM30 | Gene names | Ids | |||
|
Domain Architecture |
|
|||||
| Description | Iduronate 2-sulfatase precursor (EC 3.1.6.13) (Alpha-L-iduronate sulfate sulfatase). | |||||
|
ENPP2_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 9) | NC score | 0.165290 (rank : 12) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9R1E6, Q99LG9 | Gene names | Enpp2, Npps2 | |||
|
Domain Architecture |

|
|||||
| Description | Ectonucleotide pyrophosphatase/phosphodiesterase 2 (E-NPP 2) (Phosphodiesterase I/nucleotide pyrophosphatase 2) (Phosphodiesterase I alpha) (PD-Ialpha) [Includes: Alkaline phosphodiesterase I (EC 3.1.4.1); Nucleotide pyrophosphatase (EC 3.6.1.9) (NPPase)]. | |||||
|
ENPP6_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 10) | NC score | 0.197288 (rank : 7) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BGN3, Q4VAH5 | Gene names | Enpp6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ectonucleotide pyrophosphatase/phosphodiesterase 6 precursor (EC 3.1.-.-) (E-NPP6) (NPP-6) [Contains: Ectonucleotide pyrophosphatase/phosphodiesterase 6 soluble form]. | |||||
|
ENPP5_MOUSE
|
||||||
| θ value | 0.125558 (rank : 11) | NC score | 0.180394 (rank : 8) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9EQG7, Q921P7 | Gene names | Enpp5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ectonucleotide pyrophosphatase/phosphodiesterase 5 precursor (EC 3.1.-.-) (E-NPP5) (NPP-5). | |||||
|
ENPP1_MOUSE
|
||||||
| θ value | 0.163984 (rank : 12) | NC score | 0.159645 (rank : 13) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P06802 | Gene names | Enpp1, Npps, Pc1, Pdnp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ectonucleotide pyrophosphatase/phosphodiesterase 1 (E-NPP 1) (Phosphodiesterase I/nucleotide pyrophosphatase 1) (Plasma-cell membrane glycoprotein PC-1) (Antigen Ly-41) [Includes: Alkaline phosphodiesterase I (EC 3.1.4.1); Nucleotide pyrophosphatase (EC 3.6.1.9) (NPPase)]. | |||||
|
ENPP5_HUMAN
|
||||||
| θ value | 0.279714 (rank : 13) | NC score | 0.179257 (rank : 9) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UJA9, Q6UX49 | Gene names | ENPP5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ectonucleotide pyrophosphatase/phosphodiesterase 5 precursor (EC 3.1.-.-) (E-NPP5) (NPP-5). | |||||
|
ENPP1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 14) | NC score | 0.138932 (rank : 14) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P22413, Q5T9R6, Q9NPZ3, Q9P1P6, Q9UP61, Q9Y6K3 | Gene names | ENPP1, NPPS, PC1, PDNP1 | |||
|
Domain Architecture |

|
|||||
| Description | Ectonucleotide pyrophosphatase/phosphodiesterase 1 (E-NPP 1) (Phosphodiesterase I/nucleotide pyrophosphatase 1) (Plasma-cell membrane glycoprotein PC-1) [Includes: Alkaline phosphodiesterase I (EC 3.1.4.1); Nucleotide pyrophosphatase (EC 3.6.1.9) (NPPase)]. | |||||
|
ENPP6_HUMAN
|
||||||
| θ value | 0.62314 (rank : 15) | NC score | 0.176520 (rank : 10) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q6UWR7, Q96M57 | Gene names | ENPP6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ectonucleotide pyrophosphatase/phosphodiesterase 6 precursor (EC 3.1.-.-) (E-NPP6) (NPP-6) [Contains: Ectonucleotide pyrophosphatase/phosphodiesterase 6 soluble form]. | |||||
|
ACOX3_HUMAN
|
||||||
| θ value | 1.38821 (rank : 16) | NC score | 0.024647 (rank : 19) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O15254 | Gene names | ACOX3, BRCOX, PRCOX | |||
|
Domain Architecture |

|
|||||
| Description | Acyl-coenzyme A oxidase 3, peroxisomal (EC 1.3.3.6) (Pristanoyl-CoA oxidase) (Branched-chain acyl-CoA oxidase) (BRCACox). | |||||
|
IDS_HUMAN
|
||||||
| θ value | 2.36792 (rank : 17) | NC score | 0.074136 (rank : 17) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P22304, Q14604 | Gene names | IDS | |||
|
Domain Architecture |
|
|||||
| Description | Iduronate 2-sulfatase precursor (EC 3.1.6.13) (Alpha-L-iduronate sulfate sulfatase) (Idursulfase) [Contains: Iduronate 2-sulfatase 42 kDa chain; Iduronate 2-sulfatase 14 kDa chain]. | |||||
|
STS_HUMAN
|
||||||
| θ value | 2.36792 (rank : 18) | NC score | 0.034444 (rank : 18) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P08842 | Gene names | STS | |||
|
Domain Architecture |
|
|||||
| Description | Steryl-sulfatase precursor (EC 3.1.6.2) (Steroid sulfatase) (Steryl- sulfate sulfohydrolase) (Arylsulfatase C) (ASC). | |||||
|
KREM2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 19) | NC score | 0.012075 (rank : 21) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8K1S7 | Gene names | Kremen2 | |||
|
Domain Architecture |
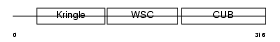
|
|||||
| Description | Kremen protein 2 precursor (Kringle-containing protein marking the eye and the nose) (Kringle domain-containing transmembrane protein 2) (Dickkopf receptor 2). | |||||
|
MEOX2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 20) | NC score | 0.003880 (rank : 24) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 340 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P50222, O75263, Q9UPL6 | Gene names | MEOX2, GAX, MOX2 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein MOX-2 (Mesenchyme homeobox 2) (Growth arrest-specific homeobox). | |||||
|
RHBT1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 21) | NC score | 0.006458 (rank : 23) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O94844 | Gene names | RHOBTB1, KIAA0740 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-related BTB domain-containing protein 1. | |||||
|
TRPV2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 22) | NC score | 0.011839 (rank : 22) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y5S1, Q9Y670 | Gene names | TRPV2, VRL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 2 (TrpV2) (osm-9-like TRP channel 2) (OTRPC2) (Vanilloid receptor-like protein 1) (VRL-1). | |||||
|
PPCE_MOUSE
|
||||||
| θ value | 8.99809 (rank : 23) | NC score | 0.012841 (rank : 20) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9QUR6, Q80YS1 | Gene names | Prep, Pep | |||
|
Domain Architecture |

|
|||||
| Description | Prolyl endopeptidase (EC 3.4.21.26) (Post-proline cleaving enzyme) (PE). | |||||
|
ZN438_HUMAN
|
||||||
| θ value | 8.99809 (rank : 24) | NC score | 0.001759 (rank : 25) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7Z4V0, Q5T426, Q658Q4, Q6ZN65 | Gene names | ZNF438 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 438. | |||||
|
ENPP3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 25) | NC score | 0.123205 (rank : 15) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O14638 | Gene names | ENPP3, PDNP3 | |||
|
Domain Architecture |

|
|||||
| Description | Ectonucleotide pyrophosphatase/phosphodiesterase 3 (E-NPP 3) (Phosphodiesterase I/nucleotide pyrophosphatase 3) (Phosphodiesterase I beta) (PD-Ibeta) (CD203c antigen) [Includes: Alkaline phosphodiesterase I (EC 3.1.4.1); Nucleotide pyrophosphatase (EC 3.6.1.9) (NPPase)]. | |||||
|
PIGO_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8TEQ8, Q6P154, Q6UX80, Q8TDS8, Q96CS9, Q9BVN9, Q9Y4B0 | Gene names | PIGO | |||
|
Domain Architecture |

|
|||||
| Description | GPI ethanolamine phosphate transferase 3 (EC 2.-.-.-) (Phosphatidylinositol-glycan biosynthesis class O protein) (PIG-O). | |||||
|
PIGO_MOUSE
|
||||||
| NC score | 0.998121 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9JJI6, Q9CRY2 | Gene names | Pigo | |||
|
Domain Architecture |

|
|||||
| Description | GPI ethanolamine phosphate transferase 3 (EC 2.-.-.-) (Phosphatidylinositol-glycan biosynthesis class O protein) (PIG-O). | |||||
|
PIGG_HUMAN
|
||||||
| NC score | 0.786584 (rank : 3) | θ value | 1.01294e-51 (rank : 3) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5H8A4, Q2TAK5, Q6UX31, Q7L5Y4, Q8N866, Q8NCC9, Q96SY9, Q9BVT7, Q9NXG5 | Gene names | PIGG, GPI7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GPI ethanolamine phosphate transferase 2 (EC 2.-.-.-) (Phosphatidylinositol-glycan biosynthesis class G protein) (PIG-G) (GPI7 homolog) (hGPI7). | |||||
|
PIGN_MOUSE
|
||||||
| NC score | 0.316970 (rank : 4) | θ value | 1.69304e-06 (rank : 4) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9R1S3, Q3V0S6, Q8VCC3, Q9R1S1, Q9R1S2 | Gene names | Pign | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GPI ethanolamine phosphate transferase 1 (EC 2.-.-.-) (Phosphatidylinositol-glycan biosynthesis class N protein) (PIG-N). | |||||
|
PIGN_HUMAN
|
||||||
| NC score | 0.302082 (rank : 5) | θ value | 8.40245e-06 (rank : 5) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O95427, Q7L8F8, Q8TC01, Q9NT05 | Gene names | PIGN, MCD4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GPI ethanolamine phosphate transferase 1 (EC 2.-.-.-) (Phosphatidylinositol-glycan biosynthesis class N protein) (PIG-N) (MCD4 homolog). | |||||
|
ENPP7_HUMAN
|
||||||
| NC score | 0.226173 (rank : 6) | θ value | 0.000158464 (rank : 6) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q6UWV6, Q6ZTS5, Q8IUS8 | Gene names | ENPP7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ectonucleotide pyrophosphatase/phosphodiesterase 7 precursor (EC 3.1.4.12) (E-NPP7) (NPP-7) (Alkaline sphingomyelin phosphodiesterase) (Intestinal alkaline sphingomyelinase) (Alk-SMase). | |||||
|
ENPP6_MOUSE
|
||||||
| NC score | 0.197288 (rank : 7) | θ value | 0.0563607 (rank : 10) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BGN3, Q4VAH5 | Gene names | Enpp6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ectonucleotide pyrophosphatase/phosphodiesterase 6 precursor (EC 3.1.-.-) (E-NPP6) (NPP-6) [Contains: Ectonucleotide pyrophosphatase/phosphodiesterase 6 soluble form]. | |||||
|
ENPP5_MOUSE
|
||||||
| NC score | 0.180394 (rank : 8) | θ value | 0.125558 (rank : 11) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9EQG7, Q921P7 | Gene names | Enpp5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ectonucleotide pyrophosphatase/phosphodiesterase 5 precursor (EC 3.1.-.-) (E-NPP5) (NPP-5). | |||||
|
ENPP5_HUMAN
|
||||||
| NC score | 0.179257 (rank : 9) | θ value | 0.279714 (rank : 13) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UJA9, Q6UX49 | Gene names | ENPP5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ectonucleotide pyrophosphatase/phosphodiesterase 5 precursor (EC 3.1.-.-) (E-NPP5) (NPP-5). | |||||
|
ENPP6_HUMAN
|
||||||
| NC score | 0.176520 (rank : 10) | θ value | 0.62314 (rank : 15) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q6UWR7, Q96M57 | Gene names | ENPP6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ectonucleotide pyrophosphatase/phosphodiesterase 6 precursor (EC 3.1.-.-) (E-NPP6) (NPP-6) [Contains: Ectonucleotide pyrophosphatase/phosphodiesterase 6 soluble form]. | |||||
|
ENPP2_HUMAN
|
||||||
| NC score | 0.169607 (rank : 11) | θ value | 0.00509761 (rank : 7) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q13822, Q13827, Q15117 | Gene names | ENPP2, ATX, PDNP2 | |||
|
Domain Architecture |

|
|||||
| Description | Ectonucleotide pyrophosphatase/phosphodiesterase 2 (E-NPP 2) (Phosphodiesterase I/nucleotide pyrophosphatase 2) (Phosphodiesterase I alpha) (PD-Ialpha) (Autotaxin) [Includes: Alkaline phosphodiesterase I (EC 3.1.4.1); Nucleotide pyrophosphatase (EC 3.6.1.9) (NPPase)]. | |||||
|
ENPP2_MOUSE
|
||||||
| NC score | 0.165290 (rank : 12) | θ value | 0.0113563 (rank : 9) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9R1E6, Q99LG9 | Gene names | Enpp2, Npps2 | |||
|
Domain Architecture |

|
|||||
| Description | Ectonucleotide pyrophosphatase/phosphodiesterase 2 (E-NPP 2) (Phosphodiesterase I/nucleotide pyrophosphatase 2) (Phosphodiesterase I alpha) (PD-Ialpha) [Includes: Alkaline phosphodiesterase I (EC 3.1.4.1); Nucleotide pyrophosphatase (EC 3.6.1.9) (NPPase)]. | |||||
|
ENPP1_MOUSE
|
||||||
| NC score | 0.159645 (rank : 13) | θ value | 0.163984 (rank : 12) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P06802 | Gene names | Enpp1, Npps, Pc1, Pdnp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ectonucleotide pyrophosphatase/phosphodiesterase 1 (E-NPP 1) (Phosphodiesterase I/nucleotide pyrophosphatase 1) (Plasma-cell membrane glycoprotein PC-1) (Antigen Ly-41) [Includes: Alkaline phosphodiesterase I (EC 3.1.4.1); Nucleotide pyrophosphatase (EC 3.6.1.9) (NPPase)]. | |||||
|
ENPP1_HUMAN
|
||||||
| NC score | 0.138932 (rank : 14) | θ value | 0.365318 (rank : 14) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P22413, Q5T9R6, Q9NPZ3, Q9P1P6, Q9UP61, Q9Y6K3 | Gene names | ENPP1, NPPS, PC1, PDNP1 | |||
|
Domain Architecture |

|
|||||
| Description | Ectonucleotide pyrophosphatase/phosphodiesterase 1 (E-NPP 1) (Phosphodiesterase I/nucleotide pyrophosphatase 1) (Plasma-cell membrane glycoprotein PC-1) [Includes: Alkaline phosphodiesterase I (EC 3.1.4.1); Nucleotide pyrophosphatase (EC 3.6.1.9) (NPPase)]. | |||||
|
ENPP3_HUMAN
|
||||||
| NC score | 0.123205 (rank : 15) | θ value | θ > 10 (rank : 25) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O14638 | Gene names | ENPP3, PDNP3 | |||
|
Domain Architecture |

|
|||||
| Description | Ectonucleotide pyrophosphatase/phosphodiesterase 3 (E-NPP 3) (Phosphodiesterase I/nucleotide pyrophosphatase 3) (Phosphodiesterase I beta) (PD-Ibeta) (CD203c antigen) [Includes: Alkaline phosphodiesterase I (EC 3.1.4.1); Nucleotide pyrophosphatase (EC 3.6.1.9) (NPPase)]. | |||||
|
IDS_MOUSE
|
||||||
| NC score | 0.110488 (rank : 16) | θ value | 0.00869519 (rank : 8) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q08890, Q3TM30 | Gene names | Ids | |||
|
Domain Architecture |

|
|||||
| Description | Iduronate 2-sulfatase precursor (EC 3.1.6.13) (Alpha-L-iduronate sulfate sulfatase). | |||||
|
IDS_HUMAN
|
||||||
| NC score | 0.074136 (rank : 17) | θ value | 2.36792 (rank : 17) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P22304, Q14604 | Gene names | IDS | |||
|
Domain Architecture |

|
|||||
| Description | Iduronate 2-sulfatase precursor (EC 3.1.6.13) (Alpha-L-iduronate sulfate sulfatase) (Idursulfase) [Contains: Iduronate 2-sulfatase 42 kDa chain; Iduronate 2-sulfatase 14 kDa chain]. | |||||
|
STS_HUMAN
|
||||||
| NC score | 0.034444 (rank : 18) | θ value | 2.36792 (rank : 18) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P08842 | Gene names | STS | |||
|
Domain Architecture |

|
|||||
| Description | Steryl-sulfatase precursor (EC 3.1.6.2) (Steroid sulfatase) (Steryl- sulfate sulfohydrolase) (Arylsulfatase C) (ASC). | |||||
|
ACOX3_HUMAN
|
||||||
| NC score | 0.024647 (rank : 19) | θ value | 1.38821 (rank : 16) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O15254 | Gene names | ACOX3, BRCOX, PRCOX | |||
|
Domain Architecture |

|
|||||
| Description | Acyl-coenzyme A oxidase 3, peroxisomal (EC 1.3.3.6) (Pristanoyl-CoA oxidase) (Branched-chain acyl-CoA oxidase) (BRCACox). | |||||
|
PPCE_MOUSE
|
||||||
| NC score | 0.012841 (rank : 20) | θ value | 8.99809 (rank : 23) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9QUR6, Q80YS1 | Gene names | Prep, Pep | |||
|
Domain Architecture |

|
|||||
| Description | Prolyl endopeptidase (EC 3.4.21.26) (Post-proline cleaving enzyme) (PE). | |||||
|
KREM2_MOUSE
|
||||||
| NC score | 0.012075 (rank : 21) | θ value | 5.27518 (rank : 19) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8K1S7 | Gene names | Kremen2 | |||
|
Domain Architecture |

|
|||||
| Description | Kremen protein 2 precursor (Kringle-containing protein marking the eye and the nose) (Kringle domain-containing transmembrane protein 2) (Dickkopf receptor 2). | |||||
|
TRPV2_HUMAN
|
||||||
| NC score | 0.011839 (rank : 22) | θ value | 6.88961 (rank : 22) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y5S1, Q9Y670 | Gene names | TRPV2, VRL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 2 (TrpV2) (osm-9-like TRP channel 2) (OTRPC2) (Vanilloid receptor-like protein 1) (VRL-1). | |||||
|
RHBT1_HUMAN
|
||||||
| NC score | 0.006458 (rank : 23) | θ value | 6.88961 (rank : 21) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O94844 | Gene names | RHOBTB1, KIAA0740 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-related BTB domain-containing protein 1. | |||||
|
MEOX2_HUMAN
|
||||||
| NC score | 0.003880 (rank : 24) | θ value | 6.88961 (rank : 20) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 340 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P50222, O75263, Q9UPL6 | Gene names | MEOX2, GAX, MOX2 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein MOX-2 (Mesenchyme homeobox 2) (Growth arrest-specific homeobox). | |||||
|
ZN438_HUMAN
|
||||||
| NC score | 0.001759 (rank : 25) | θ value | 8.99809 (rank : 24) | |||
| Query Neighborhood Hits | 24 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7Z4V0, Q5T426, Q658Q4, Q6ZN65 | Gene names | ZNF438 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 438. | |||||