Please be patient as the page loads
|
CAN3_HUMAN
|
||||||
| SwissProt Accessions | P20807, Q9BTU4, Q9Y5S6, Q9Y5S7 | Gene names | CAPN3, CANP3, CANPL3, NCL1 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-3 (EC 3.4.22.-) (Calpain L3) (Calpain p94) (Calcium-activated neutral proteinase 3) (CANP 3) (Muscle-specific calcium-activated neutral protease 3) (nCL-1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
CAN11_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.989216 (rank : 4) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9UMQ6, Q8N4R5 | Gene names | CAPN11 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-11 (EC 3.4.22.-) (Calcium-activated neutral proteinase 11) (CANP 11). | |||||
|
CAN1_HUMAN
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.988442 (rank : 6) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P07384 | Gene names | CAPN1, CANPL1 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-1 catalytic subunit (EC 3.4.22.52) (Calpain-1 large subunit) (Calcium-activated neutral proteinase 1) (CANP 1) (Calpain mu-type) (muCANP) (Micromolar-calpain). | |||||
|
CAN1_MOUSE
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.988691 (rank : 5) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | O35350, O88666 | Gene names | Capn1, Canp1, Capa1 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-1 catalytic subunit (EC 3.4.22.52) (Calpain-1 large subunit) (Calcium-activated neutral proteinase 1) (CANP 1) (Calpain mu-type) (muCANP) (Micromolar-calpain). | |||||
|
CAN2_HUMAN
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.988242 (rank : 8) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P17655, Q16738, Q6PJT3, Q8WU26, Q9HBB1 | Gene names | CAPN2, CANPL2 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-2 catalytic subunit precursor (EC 3.4.22.53) (Calpain-2 large subunit) (Calcium-activated neutral proteinase 2) (CANP 2) (Calpain M- type) (M-calpain) (Millimolar-calpain) (Calpain large polypeptide L2). | |||||
|
CAN2_MOUSE
|
||||||
| θ value | 0 (rank : 5) | NC score | 0.988268 (rank : 7) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O08529, O35518, O54843 | Gene names | Capn2 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-2 catalytic subunit precursor (EC 3.4.22.53) (Calpain-2 large subunit) (Calcium-activated neutral proteinase 2) (CANP 2) (Calpain M- type) (M-calpain) (Millimolar-calpain) (80 kDa M-calpain subunit) (CALP80). | |||||
|
CAN3_HUMAN
|
||||||
| θ value | 0 (rank : 6) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 96 | |
| SwissProt Accessions | P20807, Q9BTU4, Q9Y5S6, Q9Y5S7 | Gene names | CAPN3, CANP3, CANPL3, NCL1 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-3 (EC 3.4.22.-) (Calpain L3) (Calpain p94) (Calcium-activated neutral proteinase 3) (CANP 3) (Muscle-specific calcium-activated neutral protease 3) (nCL-1). | |||||
|
CAN3_MOUSE
|
||||||
| θ value | 0 (rank : 7) | NC score | 0.996198 (rank : 2) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q64691, Q9WUC5 | Gene names | Capn3 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-3 (EC 3.4.22.-) (Calpain L3) (Calpain p94) (Calcium-activated neutral proteinase 3) (CANP 3) (Muscle-specific calcium-activated neutral protease 3). | |||||
|
CAN9_HUMAN
|
||||||
| θ value | 0 (rank : 8) | NC score | 0.990272 (rank : 3) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O14815, Q9NS74 | Gene names | CAPN9, NCL4 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-9 (EC 3.4.22.-) (Digestive tract-specific calpain) (nCL-4) (CG36 protein). | |||||
|
CAN9_MOUSE
|
||||||
| θ value | 0 (rank : 9) | NC score | 0.988110 (rank : 9) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9D805, O35919 | Gene names | Capn9, Ncl4 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-9 (EC 3.4.22.-) (Digestive tract-specific calpain) (nCL-4). | |||||
|
CAN12_HUMAN
|
||||||
| θ value | 2.20485e-131 (rank : 10) | NC score | 0.971465 (rank : 11) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q6ZSI9 | Gene names | CAPN12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calpain-12 (EC 3.4.22.-). | |||||
|
CAN12_MOUSE
|
||||||
| θ value | 8.37844e-131 (rank : 11) | NC score | 0.971614 (rank : 10) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9ER56, Q9ER53, Q9ER54, Q9ER55 | Gene names | Capn12 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-12 (EC 3.4.22.-). | |||||
|
CAN5_MOUSE
|
||||||
| θ value | 1.18001e-100 (rank : 12) | NC score | 0.880215 (rank : 12) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O08688, Q91YU0 | Gene names | Capn5 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-5 (EC 3.4.22.-) (nCL-3). | |||||
|
CAN5_HUMAN
|
||||||
| θ value | 9.98939e-100 (rank : 13) | NC score | 0.879663 (rank : 13) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O15484, O00263 | Gene names | CAPN5 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-5 (EC 3.4.22.-) (nCL-3) (htra-3). | |||||
|
CAN6_HUMAN
|
||||||
| θ value | 4.36332e-79 (rank : 14) | NC score | 0.867223 (rank : 14) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Y6Q1, Q9UEQ1, Q9UJA8 | Gene names | CAPN6, CALPM, CANPX | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-6 (Calpamodulin) (CalpM) (Calpain-like protease X-linked). | |||||
|
CAN6_MOUSE
|
||||||
| θ value | 3.12697e-77 (rank : 15) | NC score | 0.866187 (rank : 15) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O35646 | Gene names | Capn6, Capa6 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-6. | |||||
|
CPNS1_MOUSE
|
||||||
| θ value | 2.94036e-59 (rank : 16) | NC score | 0.836444 (rank : 18) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O88456, Q3V0X5, Q5BKQ2, Q8CEI2, Q8VEK4, Q9R1C5 | Gene names | Capns1, Capn4 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain small subunit 1 (CSS1) (Calcium-dependent protease small subunit 1) (Calcium-dependent protease small subunit) (CDPS) (Calpain regulatory subunit) (Calcium-activated neutral proteinase small subunit) (CANP small subunit). | |||||
|
CPNS1_HUMAN
|
||||||
| θ value | 1.45929e-58 (rank : 17) | NC score | 0.834652 (rank : 19) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P04632, Q8WTX3, Q96EW0 | Gene names | CAPNS1, CAPN4, CAPNS | |||
|
Domain Architecture |

|
|||||
| Description | Calpain small subunit 1 (CSS1) (Calcium-dependent protease small subunit 1) (Calcium-dependent protease small subunit) (CDPS) (Calpain regulatory subunit) (Calcium-activated neutral proteinase small subunit) (CANP small subunit). | |||||
|
CAN10_MOUSE
|
||||||
| θ value | 4.24587e-58 (rank : 18) | NC score | 0.852987 (rank : 16) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9ESK3, Q99J13, Q9WVF0 | Gene names | Capn10, Capn8 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-10 (EC 3.4.22.-) (Calcium-activated neutral proteinase 10) (CANP 10). | |||||
|
CAN10_HUMAN
|
||||||
| θ value | 1.2784e-54 (rank : 19) | NC score | 0.848882 (rank : 17) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9HC96, Q8NCD4, Q96IG4, Q96JI2, Q9HC89, Q9HC90, Q9HC91, Q9HC92, Q9HC93, Q9HC94, Q9HC95 | Gene names | CAPN10, KIAA1845 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-10 (EC 3.4.22.-) (Calcium-activated neutral proteinase 10) (CANP 10). | |||||
|
CPNS2_HUMAN
|
||||||
| θ value | 7.75577e-52 (rank : 20) | NC score | 0.830558 (rank : 20) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q96L46, Q9BPV4 | Gene names | CAPNS2 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain small subunit 2 (CSS2) (Calcium-dependent protease small subunit 2). | |||||
|
CPNS2_MOUSE
|
||||||
| θ value | 1.32293e-51 (rank : 21) | NC score | 0.826076 (rank : 21) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9D7J7 | Gene names | Capns2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calpain small subunit 2 (CSS2) (Calcium-dependent protease small subunit 2). | |||||
|
GRAN_MOUSE
|
||||||
| θ value | 1.42661e-21 (rank : 22) | NC score | 0.716613 (rank : 23) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8VC88, Q8BL53, Q8K3Z6 | Gene names | Gca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Grancalcin. | |||||
|
SORCN_HUMAN
|
||||||
| θ value | 4.15078e-21 (rank : 23) | NC score | 0.712365 (rank : 24) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P30626 | Gene names | SRI | |||
|
Domain Architecture |
|
|||||
| Description | Sorcin (22 kDa protein) (CP-22) (V19). | |||||
|
SORCN_MOUSE
|
||||||
| θ value | 4.15078e-21 (rank : 24) | NC score | 0.717150 (rank : 22) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q6P069, Q3UKC5, Q9CR38, Q9D7V8 | Gene names | Sri | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorcin. | |||||
|
GRAN_HUMAN
|
||||||
| θ value | 5.99374e-20 (rank : 25) | NC score | 0.708926 (rank : 25) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P28676 | Gene names | GCA, GCL | |||
|
Domain Architecture |
|
|||||
| Description | Grancalcin. | |||||
|
PDCD6_HUMAN
|
||||||
| θ value | 9.90251e-15 (rank : 26) | NC score | 0.597291 (rank : 30) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | O75340 | Gene names | PDCD6, ALG2 | |||
|
Domain Architecture |
|
|||||
| Description | Programmed cell death protein 6 (Probable calcium-binding protein ALG- 2). | |||||
|
PDCD6_MOUSE
|
||||||
| θ value | 9.90251e-15 (rank : 27) | NC score | 0.595394 (rank : 31) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P12815, Q545I0, Q61145 | Gene names | Pdcd6, Alg2 | |||
|
Domain Architecture |
|
|||||
| Description | Programmed cell death protein 6 (Probable calcium-binding protein ALG- 2) (PMP41) (ALG-257). | |||||
|
CAN7_HUMAN
|
||||||
| θ value | 2.88119e-14 (rank : 28) | NC score | 0.644967 (rank : 29) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Y6W3 | Gene names | CAPN7, PALBH | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-7 (EC 3.4.22.-) (PalB homolog) (PalBH). | |||||
|
CAN7_MOUSE
|
||||||
| θ value | 2.88119e-14 (rank : 29) | NC score | 0.647208 (rank : 28) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9R1S8, Q9Z0P9 | Gene names | Capn7, Palbh | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-7 (EC 3.4.22.-) (PalB homolog) (PalBH). | |||||
|
PEF1_HUMAN
|
||||||
| θ value | 1.09485e-13 (rank : 30) | NC score | 0.654497 (rank : 27) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9UBV8 | Gene names | PEF1, ABP32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peflin (PEF protein with a long N-terminal hydrophobic domain) (Penta- EF hand domain-containing protein 1). | |||||
|
PEF1_MOUSE
|
||||||
| θ value | 1.86753e-13 (rank : 31) | NC score | 0.667816 (rank : 26) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q8BFY6, Q8VCT5, Q9CYW8, Q9D934 | Gene names | Pef1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peflin (PEF protein with a long N-terminal hydrophobic domain) (Penta- EF hand domain-containing protein 1). | |||||
|
CALL5_HUMAN
|
||||||
| θ value | 1.87187e-05 (rank : 32) | NC score | 0.208335 (rank : 32) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9NZT1, Q8IXU8 | Gene names | CALML5, CLSP | |||
|
Domain Architecture |
|
|||||
| Description | Calmodulin-like protein 5 (Calmodulin-like skin protein). | |||||
|
CALM_HUMAN
|
||||||
| θ value | 0.00134147 (rank : 33) | NC score | 0.135356 (rank : 35) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P62158, P02593, P70667, P99014, Q13942, Q53S29, Q61379, Q61380, Q96HK3 | Gene names | CALM1, CALM, CAM, CAM1 | |||
|
Domain Architecture |
|
|||||
| Description | Calmodulin (CaM). | |||||
|
CALM_MOUSE
|
||||||
| θ value | 0.00134147 (rank : 34) | NC score | 0.135356 (rank : 36) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P62204, P02593, P70667, P99014, Q61379, Q61380 | Gene names | Calm1, Calm, Cam, Cam1 | |||
|
Domain Architecture |
|
|||||
| Description | Calmodulin (CaM). | |||||
|
CALM4_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 35) | NC score | 0.165648 (rank : 33) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 229 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q9JM83, Q9CR31, Q9D1E9 | Gene names | Calm4, Dd112 | |||
|
Domain Architecture |
|
|||||
| Description | Calmodulin-4 (Calcium-binding protein Dd112). | |||||
|
CALB1_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 36) | NC score | 0.105003 (rank : 49) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P05937 | Gene names | CALB1, CAB27 | |||
|
Domain Architecture |
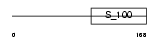
|
|||||
| Description | Calbindin (Vitamin D-dependent calcium-binding protein, avian-type) (Calbindin D28) (D-28K). | |||||
|
CALB1_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 37) | NC score | 0.109555 (rank : 47) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P12658, Q545N6 | Gene names | Calb1 | |||
|
Domain Architecture |
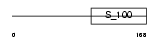
|
|||||
| Description | Calbindin (Vitamin D-dependent calcium-binding protein, avian-type) (Calbindin D28) (D-28K) (Spot 35 protein) (PCD-29). | |||||
|
CALL3_HUMAN
|
||||||
| θ value | 0.163984 (rank : 38) | NC score | 0.128063 (rank : 37) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P27482 | Gene names | CALML3 | |||
|
Domain Architecture |
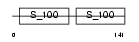
|
|||||
| Description | Calmodulin-like protein 3 (Calmodulin-related protein NB-1) (CaM-like protein) (CLP). | |||||
|
CETN2_MOUSE
|
||||||
| θ value | 0.21417 (rank : 39) | NC score | 0.118502 (rank : 42) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9R1K9, Q3UBB4, Q9CWM0 | Gene names | Cetn2, Calt | |||
|
Domain Architecture |
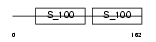
|
|||||
| Description | Centrin-2 (Caltractin isoform 1). | |||||
|
CALB2_HUMAN
|
||||||
| θ value | 0.279714 (rank : 40) | NC score | 0.110548 (rank : 46) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P22676, Q53HD2, Q96BK4 | Gene names | CALB2, CAB29 | |||
|
Domain Architecture |
|
|||||
| Description | Calretinin (CR) (29 kDa calbindin). | |||||
|
CABP1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 41) | NC score | 0.093825 (rank : 54) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9NZU7, O95663, Q9NZU8 | Gene names | CABP1 | |||
|
Domain Architecture |
|
|||||
| Description | Calcium-binding protein 1 (CaBP1) (Calbrain). | |||||
|
CABP1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 42) | NC score | 0.094403 (rank : 51) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9JLK7, Q9JLK6 | Gene names | Cabp1 | |||
|
Domain Architecture |
|
|||||
| Description | Calcium-binding protein 1 (CaBP1). | |||||
|
CETN2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 43) | NC score | 0.114027 (rank : 44) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P41208, Q53XW1 | Gene names | CETN2, CALT, CEN2 | |||
|
Domain Architecture |
|
|||||
| Description | Centrin-2 (Caltractin isoform 1). | |||||
|
CTNL1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 44) | NC score | 0.023040 (rank : 82) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UBT7, O76084, Q53FQ2, Q5JTQ7, Q5JTQ8, Q9Y401 | Gene names | CTNNAL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alpha-catulin (Catenin alpha-like protein 1) (Alpha-catenin-related protein) (ACRP). | |||||
|
SPTA2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 45) | NC score | 0.026065 (rank : 79) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 888 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q13813, Q13186, Q15324, Q16606, Q59EF1, Q5VXV5, Q5VXV6, Q7Z6M5, Q9P0V0 | Gene names | SPTAN1, SPTA2 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin alpha chain, brain (Spectrin, non-erythroid alpha chain) (Alpha-II spectrin) (Fodrin alpha chain). | |||||
|
TNNC2_MOUSE
|
||||||
| θ value | 0.47712 (rank : 46) | NC score | 0.111370 (rank : 45) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P20801 | Gene names | Tnnc2, Tncs | |||
|
Domain Architecture |
|
|||||
| Description | Troponin C, skeletal muscle (STNC). | |||||
|
TSH1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 47) | NC score | 0.010747 (rank : 87) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 675 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5DTH5, Q6PE60, Q8VD19, Q9JLD5 | Gene names | Tshz1, Kiaa4206, Sdccag33, Tsh1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 1 (Serologically defined colon cancer antigen 3 homolog). | |||||
|
TNNC2_HUMAN
|
||||||
| θ value | 0.62314 (rank : 48) | NC score | 0.107718 (rank : 48) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P02585 | Gene names | TNNC2 | |||
|
Domain Architecture |
|
|||||
| Description | Troponin C, skeletal muscle. | |||||
|
UBP48_MOUSE
|
||||||
| θ value | 0.62314 (rank : 49) | NC score | 0.008229 (rank : 91) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q3V0C5, Q3TM39, Q571J9, Q8VDJ1 | Gene names | Usp48, Kiaa4202 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 48 (EC 3.1.2.15) (Ubiquitin thioesterase 48) (Ubiquitin-specific-processing protease 48) (Deubiquitinating enzyme 48). | |||||
|
CALL6_HUMAN
|
||||||
| θ value | 0.813845 (rank : 50) | NC score | 0.082346 (rank : 65) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8TD86, Q6Q2C4 | Gene names | CALML6, CAGLP, CALGP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calmodulin-like protein 6 (Calglandulin-like protein). | |||||
|
EFHB_MOUSE
|
||||||
| θ value | 0.813845 (rank : 51) | NC score | 0.047819 (rank : 76) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8CDU5 | Gene names | Efhb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EF-hand domain-containing family member B. | |||||
|
EFHB_HUMAN
|
||||||
| θ value | 1.06291 (rank : 52) | NC score | 0.054952 (rank : 75) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8N7U6, Q6ZWK9, Q8IV58, Q96LQ6 | Gene names | EFHB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EF-hand domain-containing family member B. | |||||
|
LRC17_MOUSE
|
||||||
| θ value | 1.06291 (rank : 53) | NC score | 0.004186 (rank : 97) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 268 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9CXD9 | Gene names | Lrrc17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 17 precursor. | |||||
|
BRPF1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 54) | NC score | 0.006090 (rank : 92) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P55201, Q9UHI0 | Gene names | BRPF1, BR140 | |||
|
Domain Architecture |

|
|||||
| Description | Peregrin (Bromodomain and PHD finger-containing protein 1) (BR140 protein). | |||||
|
ELF2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 55) | NC score | 0.012592 (rank : 85) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9JHC9, Q6NST2, Q8BTX8, Q9JHC7, Q9JHC8, Q9JHD0 | Gene names | Elf2, Nerf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-2 (E74-like factor 2) (New ETS- related factor). | |||||
|
CETN1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 56) | NC score | 0.138021 (rank : 34) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q12798 | Gene names | CETN1, CEN1, CETN | |||
|
Domain Architecture |
|
|||||
| Description | Centrin-1 (Caltractin isoform 2). | |||||
|
FANCM_HUMAN
|
||||||
| θ value | 1.81305 (rank : 57) | NC score | 0.008968 (rank : 90) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8IYD8, Q3YFH9, Q8N9X6, Q9HCH6 | Gene names | FANCM, KIAA1596 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fanconi anemia group M protein (EC 3.6.1.-) (ATP-dependent RNA helicase FANCM) (Protein FACM) (Fanconi anemia-associated polypeptide of 250 kDa) (FAAP250) (Protein Hef ortholog). | |||||
|
SEGN_HUMAN
|
||||||
| θ value | 1.81305 (rank : 58) | NC score | 0.094857 (rank : 50) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O76038, Q5VV44, Q96QV7, Q9UJF6 | Gene names | SCGN, SECRET | |||
|
Domain Architecture |
|
|||||
| Description | Secretagogin. | |||||
|
TFAM_HUMAN
|
||||||
| θ value | 1.81305 (rank : 59) | NC score | 0.011265 (rank : 86) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q00059 | Gene names | TFAM, TCF6L2 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor A, mitochondrial precursor (mtTFA) (Mitochondrial transcription factor 1) (MtTF1) (Transcription factor 6-like 2). | |||||
|
CABP5_HUMAN
|
||||||
| θ value | 2.36792 (rank : 60) | NC score | 0.086061 (rank : 61) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9NP86 | Gene names | CABP5 | |||
|
Domain Architecture |
|
|||||
| Description | Calcium-binding protein 5 (CaBP5). | |||||
|
CETN1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 61) | NC score | 0.125688 (rank : 39) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P41209, Q3V119, Q9D9G9, Q9DAL6 | Gene names | Cetn1, Calt | |||
|
Domain Architecture |
|
|||||
| Description | Centrin-1 (Caltractin). | |||||
|
ELF2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 62) | NC score | 0.009515 (rank : 88) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q15723, Q15724, Q15725, Q6P1K5 | Gene names | ELF2, NERF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-2 (E74-like factor 2) (New ETS- related factor). | |||||
|
PLSI_HUMAN
|
||||||
| θ value | 2.36792 (rank : 63) | NC score | 0.086950 (rank : 59) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q14651 | Gene names | PLS1 | |||
|
Domain Architecture |

|
|||||
| Description | Plastin-1 (I-plastin) (Intestine-specific plastin). | |||||
|
SEGN_MOUSE
|
||||||
| θ value | 2.36792 (rank : 64) | NC score | 0.081150 (rank : 66) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q91WD9 | Gene names | Scgn | |||
|
Domain Architecture |

|
|||||
| Description | Secretagogin. | |||||
|
CABP4_MOUSE
|
||||||
| θ value | 3.0926 (rank : 65) | NC score | 0.087533 (rank : 58) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8VHC5 | Gene names | Cabp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium-binding protein 4 (CaBP4). | |||||
|
EFCB1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 66) | NC score | 0.028802 (rank : 78) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9HAE3 | Gene names | EFCAB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EF-hand calcium-binding domain-containing protein 1. | |||||
|
ACTN1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 67) | NC score | 0.025760 (rank : 80) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q7TPR4 | Gene names | Actn1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-actinin-1 (Alpha-actinin cytoskeletal isoform) (Non-muscle alpha-actinin-1) (F-actin cross-linking protein). | |||||
|
BSN_MOUSE
|
||||||
| θ value | 4.03905 (rank : 68) | NC score | 0.001990 (rank : 102) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
ACTN1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 69) | NC score | 0.024920 (rank : 81) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P12814, Q9BTN1 | Gene names | ACTN1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-actinin-1 (Alpha-actinin cytoskeletal isoform) (Non-muscle alpha-actinin-1) (F-actin cross-linking protein). | |||||
|
CABP2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 70) | NC score | 0.086256 (rank : 60) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9NPB3 | Gene names | CABP2 | |||
|
Domain Architecture |
|
|||||
| Description | Calcium-binding protein 2 (CaBP2). | |||||
|
IF2B_MOUSE
|
||||||
| θ value | 5.27518 (rank : 71) | NC score | 0.009183 (rank : 89) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99L45, Q9CSH6, Q9CT12 | Gene names | Eif2s2 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2 subunit 2 (Eukaryotic translation initiation factor 2 subunit beta) (eIF-2-beta). | |||||
|
IWS1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 72) | NC score | -0.000514 (rank : 108) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
RBBP6_MOUSE
|
||||||
| θ value | 5.27518 (rank : 73) | NC score | 0.001050 (rank : 105) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
TNNC1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 74) | NC score | 0.126071 (rank : 38) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | P19123 | Gene names | Tnnc1, Tncc | |||
|
Domain Architecture |
|
|||||
| Description | Troponin C, slow skeletal and cardiac muscles (TN-C). | |||||
|
KI13B_HUMAN
|
||||||
| θ value | 6.88961 (rank : 75) | NC score | 0.000084 (rank : 107) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 492 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NQT8, O75134, Q9BYJ6 | Gene names | KIF13B, GAKIN, KIAA0639 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF13B (Kinesin-like protein GAKIN). | |||||
|
LBXCO_MOUSE
|
||||||
| θ value | 6.88961 (rank : 76) | NC score | 0.005131 (rank : 93) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BX46, Q3UYA4, Q5W8I2, Q8C0T2 | Gene names | Lbxcor1, Corl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladybird homeobox corepressor 1 (Transcriptional corepressor Corl1). | |||||
|
LRRN1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 77) | NC score | -0.000927 (rank : 109) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 518 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6UXK5, Q3LID5, Q8IYV5, Q9H8V1, Q9P231 | Gene names | LRRN1, KIAA1497 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats neuronal protein 1 precursor (Neuronal leucine- rich repeat protein 1) (NLRR-1). | |||||
|
NOX5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 78) | NC score | 0.016718 (rank : 84) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96PH1, Q8TEQ1, Q8TER4, Q96PH2, Q96PJ8, Q96PJ9, Q9H6E0, Q9HAM8 | Gene names | NOX5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NADPH oxidase 5 (EC 1.6.3.-). | |||||
|
PO121_HUMAN
|
||||||
| θ value | 6.88961 (rank : 79) | NC score | 0.003951 (rank : 98) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y2N3, O75115, Q9Y4S7 | Gene names | POM121, KIAA0618, NUP121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear envelope pore membrane protein POM 121 (Pore membrane protein of 121 kDa) (P145). | |||||
|
RYR2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 80) | NC score | 0.019782 (rank : 83) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q92736, Q15411, Q546N8 | Gene names | RYR2 | |||
|
Domain Architecture |

|
|||||
| Description | Ryanodine receptor 2 (Cardiac muscle-type ryanodine receptor) (RyR2) (RYR-2) (Cardiac muscle ryanodine receptor-calcium release channel) (hRYR-2). | |||||
|
SFPQ_MOUSE
|
||||||
| θ value | 6.88961 (rank : 81) | NC score | 0.002962 (rank : 100) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8VIJ6, Q9ERW2 | Gene names | Sfpq | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, proline- and glutamine-rich (Polypyrimidine tract- binding protein-associated-splicing factor) (PTB-associated-splicing factor) (PSF) (DNA-binding p52/p100 complex, 100 kDa subunit). | |||||
|
TNNC1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 82) | NC score | 0.119822 (rank : 40) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P63316, O14800, P02590, P04463 | Gene names | TNNC1, TNNC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Troponin C, slow skeletal and cardiac muscles (TN-C). | |||||
|
ZN226_HUMAN
|
||||||
| θ value | 6.88961 (rank : 83) | NC score | -0.004666 (rank : 110) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 770 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NYT6, Q96TE6, Q9NS44 | Gene names | ZNF226 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger protein 226. | |||||
|
CABP5_MOUSE
|
||||||
| θ value | 8.99809 (rank : 84) | NC score | 0.070548 (rank : 69) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9JLK3 | Gene names | Cabp5 | |||
|
Domain Architecture |
|
|||||
| Description | Calcium-binding protein 5 (CaBP5). | |||||
|
CANB1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 85) | NC score | 0.085329 (rank : 63) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P63098, P06705, P15117, Q08044 | Gene names | PPP3R1, CNA2, CNB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcineurin subunit B isoform 1 (Protein phosphatase 2B regulatory subunit 1) (Protein phosphatase 3 regulatory subunit B alpha isoform 1). | |||||
|
CANB1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 86) | NC score | 0.085333 (rank : 62) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q63810 | Gene names | Ppp3r1, Cnb | |||
|
Domain Architecture |
|
|||||
| Description | Calcineurin subunit B isoform 1 (Protein phosphatase 2B regulatory subunit 1) (Protein phosphatase 3 regulatory subunit B alpha isoform 1). | |||||
|
CANB2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 87) | NC score | 0.071551 (rank : 68) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q96LZ3, Q7Z4V8, Q8WYJ4 | Gene names | PPP3R2, CBLP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcineurin subunit B isoform 2 (Protein phosphatase 2B regulatory subunit 2) (Protein phosphatase 3 regulatory subunit B beta isoform) (Calcineurin B-like protein) (CBLP) (CNBII). | |||||
|
DSG3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 88) | NC score | 0.001375 (rank : 103) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O35902, Q8CB02, Q8CE48 | Gene names | Dsg3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Desmoglein-3 precursor (130 kDa pemphigus vulgaris antigen homolog). | |||||
|
FOXD1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 89) | NC score | 0.000100 (rank : 106) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q61345 | Gene names | Foxd1, Fkhl8, Freac4, Hfhbf2 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein D1 (Forkhead-related protein FKHL8) (Forkhead- related transcription factor 4) (FREAC-4) (Brain factor 2) (HFH-BF-2). | |||||
|
LEO1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 90) | NC score | 0.001233 (rank : 104) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
MAP9_HUMAN
|
||||||
| θ value | 8.99809 (rank : 91) | NC score | 0.002134 (rank : 101) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 518 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q49MG5, Q4W5I7, Q68DU1, Q9H781, Q9H7B6 | Gene names | MAP9, ASAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 9 (Aster-associated protein). | |||||
|
MOR2A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 92) | NC score | 0.004619 (rank : 95) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q69ZX6, Q5QNQ7, Q6P547 | Gene names | Morc2a, Kiaa0852, Zcwcc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORC family CW-type zinc finger protein 2A (Zinc finger CW-type coiled-coil domain protein 1). | |||||
|
MORC2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 93) | NC score | 0.004577 (rank : 96) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y6X9, Q9UF28, Q9Y6V2 | Gene names | MORC2, KIAA0852, ZCWCC1 | |||
|
Domain Architecture |

|
|||||
| Description | MORC family CW-type zinc finger protein 2 (Zinc finger CW-type coiled- coil domain protein 1). | |||||
|
ONCO_MOUSE
|
||||||
| θ value | 8.99809 (rank : 94) | NC score | 0.046352 (rank : 77) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P51879, Q62004 | Gene names | Ocm | |||
|
Domain Architecture |
|
|||||
| Description | Oncomodulin (OM) (Parvalbumin beta). | |||||
|
RANB9_HUMAN
|
||||||
| θ value | 8.99809 (rank : 95) | NC score | 0.004755 (rank : 94) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 179 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96S59, O94764, Q6P3T7, Q7LBR2, Q7Z7F9 | Gene names | RANBP9, RANBPM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 9 (RanBP9) (RanBP7) (Ran-binding protein M) (RanBPM) (BPM90) (BPM-L). | |||||
|
SIN3B_HUMAN
|
||||||
| θ value | 8.99809 (rank : 96) | NC score | 0.003643 (rank : 99) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O75182, Q68GC2, Q6P4B8, Q8TB34, Q9BSC8 | Gene names | SIN3B, KIAA0700 | |||
|
Domain Architecture |
|
|||||
| Description | Paired amphipathic helix protein Sin3b (Transcriptional corepressor Sin3b) (Histone deacetylase complex subunit Sin3b). | |||||
|
CABP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.089074 (rank : 55) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9JLK4, Q9JLK5 | Gene names | Cabp2 | |||
|
Domain Architecture |
|
|||||
| Description | Calcium-binding protein 2 (CaBP2). | |||||
|
CABP3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 98) | NC score | 0.067293 (rank : 71) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9NZU6, Q9NYY0, Q9NZT9 | Gene names | CABP3 | |||
|
Domain Architecture |
|
|||||
| Description | Calcium-binding protein 3 (CaBP3). | |||||
|
CABP4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 99) | NC score | 0.117494 (rank : 43) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P57796, Q8N4Z2, Q8WWY5 | Gene names | CABP4 | |||
|
Domain Architecture |
|
|||||
| Description | Calcium-binding protein 4 (CaBP4). | |||||
|
CABP7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.088847 (rank : 56) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q86V35 | Gene names | CABP7 | |||
|
Domain Architecture |
|
|||||
| Description | Calcium-binding protein 7 (CaBP7). | |||||
|
CABP7_MOUSE
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.088847 (rank : 57) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q91ZM8 | Gene names | Cabp7 | |||
|
Domain Architecture |
|
|||||
| Description | Calcium-binding protein 7 (CaBP7). | |||||
|
CALB2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.082844 (rank : 64) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q08331, Q60964, Q9JM81 | Gene names | Calb2 | |||
|
Domain Architecture |

|
|||||
| Description | Calretinin (CR). | |||||
|
CALN1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.057394 (rank : 72) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9BXU9 | Gene names | CALN1, CABP8 | |||
|
Domain Architecture |
|
|||||
| Description | Calneuron-1 (Calcium-binding protein CaBP8). | |||||
|
CALN1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.057305 (rank : 73) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9JJG7 | Gene names | Caln1 | |||
|
Domain Architecture |
|
|||||
| Description | Calneuron-1. | |||||
|
CANB2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.073929 (rank : 67) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q63811 | Gene names | Ppp3r2 | |||
|
Domain Architecture |
|
|||||
| Description | Calcineurin subunit B isoform 2 (Protein phosphatase 2B regulatory subunit 2) (Protein phosphatase 3 regulatory subunit B beta isoform). | |||||
|
CAYP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.069588 (rank : 70) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q13938, Q3MJA7, Q8WUZ3 | Gene names | CAPS | |||
|
Domain Architecture |
|
|||||
| Description | Calcyphosin (Calcyphosine). | |||||
|
CETN3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.094115 (rank : 53) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | O15182, Q9BS23 | Gene names | CETN3, CEN3 | |||
|
Domain Architecture |
|
|||||
| Description | Centrin-3. | |||||
|
CETN3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 108) | NC score | 0.094395 (rank : 52) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | O35648 | Gene names | Cetn3, Cen3 | |||
|
Domain Architecture |
|
|||||
| Description | Centrin-3. | |||||
|
CF103_HUMAN
|
||||||
| θ value | θ > 10 (rank : 109) | NC score | 0.119476 (rank : 41) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8N7X0, Q5T402 | Gene names | C6orf103 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf103. | |||||
|
PRVA_MOUSE
|
||||||
| θ value | θ > 10 (rank : 110) | NC score | 0.056054 (rank : 74) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P32848 | Gene names | Pvalb, Pva | |||
|
Domain Architecture |
|
|||||
| Description | Parvalbumin alpha. | |||||
|
CAN3_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 6) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 96 | |
| SwissProt Accessions | P20807, Q9BTU4, Q9Y5S6, Q9Y5S7 | Gene names | CAPN3, CANP3, CANPL3, NCL1 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-3 (EC 3.4.22.-) (Calpain L3) (Calpain p94) (Calcium-activated neutral proteinase 3) (CANP 3) (Muscle-specific calcium-activated neutral protease 3) (nCL-1). | |||||
|
CAN3_MOUSE
|
||||||
| NC score | 0.996198 (rank : 2) | θ value | 0 (rank : 7) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q64691, Q9WUC5 | Gene names | Capn3 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-3 (EC 3.4.22.-) (Calpain L3) (Calpain p94) (Calcium-activated neutral proteinase 3) (CANP 3) (Muscle-specific calcium-activated neutral protease 3). | |||||
|
CAN9_HUMAN
|
||||||
| NC score | 0.990272 (rank : 3) | θ value | 0 (rank : 8) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O14815, Q9NS74 | Gene names | CAPN9, NCL4 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-9 (EC 3.4.22.-) (Digestive tract-specific calpain) (nCL-4) (CG36 protein). | |||||
|
CAN11_HUMAN
|
||||||
| NC score | 0.989216 (rank : 4) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9UMQ6, Q8N4R5 | Gene names | CAPN11 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-11 (EC 3.4.22.-) (Calcium-activated neutral proteinase 11) (CANP 11). | |||||
|
CAN1_MOUSE
|
||||||
| NC score | 0.988691 (rank : 5) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | O35350, O88666 | Gene names | Capn1, Canp1, Capa1 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-1 catalytic subunit (EC 3.4.22.52) (Calpain-1 large subunit) (Calcium-activated neutral proteinase 1) (CANP 1) (Calpain mu-type) (muCANP) (Micromolar-calpain). | |||||
|
CAN1_HUMAN
|
||||||
| NC score | 0.988442 (rank : 6) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P07384 | Gene names | CAPN1, CANPL1 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-1 catalytic subunit (EC 3.4.22.52) (Calpain-1 large subunit) (Calcium-activated neutral proteinase 1) (CANP 1) (Calpain mu-type) (muCANP) (Micromolar-calpain). | |||||
|
CAN2_MOUSE
|
||||||
| NC score | 0.988268 (rank : 7) | θ value | 0 (rank : 5) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O08529, O35518, O54843 | Gene names | Capn2 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-2 catalytic subunit precursor (EC 3.4.22.53) (Calpain-2 large subunit) (Calcium-activated neutral proteinase 2) (CANP 2) (Calpain M- type) (M-calpain) (Millimolar-calpain) (80 kDa M-calpain subunit) (CALP80). | |||||
|
CAN2_HUMAN
|
||||||
| NC score | 0.988242 (rank : 8) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P17655, Q16738, Q6PJT3, Q8WU26, Q9HBB1 | Gene names | CAPN2, CANPL2 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-2 catalytic subunit precursor (EC 3.4.22.53) (Calpain-2 large subunit) (Calcium-activated neutral proteinase 2) (CANP 2) (Calpain M- type) (M-calpain) (Millimolar-calpain) (Calpain large polypeptide L2). | |||||
|
CAN9_MOUSE
|
||||||
| NC score | 0.988110 (rank : 9) | θ value | 0 (rank : 9) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9D805, O35919 | Gene names | Capn9, Ncl4 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-9 (EC 3.4.22.-) (Digestive tract-specific calpain) (nCL-4). | |||||
|
CAN12_MOUSE
|
||||||
| NC score | 0.971614 (rank : 10) | θ value | 8.37844e-131 (rank : 11) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9ER56, Q9ER53, Q9ER54, Q9ER55 | Gene names | Capn12 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-12 (EC 3.4.22.-). | |||||
|
CAN12_HUMAN
|
||||||
| NC score | 0.971465 (rank : 11) | θ value | 2.20485e-131 (rank : 10) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q6ZSI9 | Gene names | CAPN12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calpain-12 (EC 3.4.22.-). | |||||
|
CAN5_MOUSE
|
||||||
| NC score | 0.880215 (rank : 12) | θ value | 1.18001e-100 (rank : 12) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O08688, Q91YU0 | Gene names | Capn5 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-5 (EC 3.4.22.-) (nCL-3). | |||||
|
CAN5_HUMAN
|
||||||
| NC score | 0.879663 (rank : 13) | θ value | 9.98939e-100 (rank : 13) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O15484, O00263 | Gene names | CAPN5 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-5 (EC 3.4.22.-) (nCL-3) (htra-3). | |||||
|
CAN6_HUMAN
|
||||||
| NC score | 0.867223 (rank : 14) | θ value | 4.36332e-79 (rank : 14) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Y6Q1, Q9UEQ1, Q9UJA8 | Gene names | CAPN6, CALPM, CANPX | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-6 (Calpamodulin) (CalpM) (Calpain-like protease X-linked). | |||||
|
CAN6_MOUSE
|
||||||
| NC score | 0.866187 (rank : 15) | θ value | 3.12697e-77 (rank : 15) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O35646 | Gene names | Capn6, Capa6 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-6. | |||||
|
CAN10_MOUSE
|
||||||
| NC score | 0.852987 (rank : 16) | θ value | 4.24587e-58 (rank : 18) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9ESK3, Q99J13, Q9WVF0 | Gene names | Capn10, Capn8 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-10 (EC 3.4.22.-) (Calcium-activated neutral proteinase 10) (CANP 10). | |||||
|
CAN10_HUMAN
|
||||||
| NC score | 0.848882 (rank : 17) | θ value | 1.2784e-54 (rank : 19) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9HC96, Q8NCD4, Q96IG4, Q96JI2, Q9HC89, Q9HC90, Q9HC91, Q9HC92, Q9HC93, Q9HC94, Q9HC95 | Gene names | CAPN10, KIAA1845 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-10 (EC 3.4.22.-) (Calcium-activated neutral proteinase 10) (CANP 10). | |||||
|
CPNS1_MOUSE
|
||||||
| NC score | 0.836444 (rank : 18) | θ value | 2.94036e-59 (rank : 16) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O88456, Q3V0X5, Q5BKQ2, Q8CEI2, Q8VEK4, Q9R1C5 | Gene names | Capns1, Capn4 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain small subunit 1 (CSS1) (Calcium-dependent protease small subunit 1) (Calcium-dependent protease small subunit) (CDPS) (Calpain regulatory subunit) (Calcium-activated neutral proteinase small subunit) (CANP small subunit). | |||||
|
CPNS1_HUMAN
|
||||||
| NC score | 0.834652 (rank : 19) | θ value | 1.45929e-58 (rank : 17) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P04632, Q8WTX3, Q96EW0 | Gene names | CAPNS1, CAPN4, CAPNS | |||
|
Domain Architecture |

|
|||||
| Description | Calpain small subunit 1 (CSS1) (Calcium-dependent protease small subunit 1) (Calcium-dependent protease small subunit) (CDPS) (Calpain regulatory subunit) (Calcium-activated neutral proteinase small subunit) (CANP small subunit). | |||||
|
CPNS2_HUMAN
|
||||||
| NC score | 0.830558 (rank : 20) | θ value | 7.75577e-52 (rank : 20) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q96L46, Q9BPV4 | Gene names | CAPNS2 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain small subunit 2 (CSS2) (Calcium-dependent protease small subunit 2). | |||||
|
CPNS2_MOUSE
|
||||||
| NC score | 0.826076 (rank : 21) | θ value | 1.32293e-51 (rank : 21) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9D7J7 | Gene names | Capns2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calpain small subunit 2 (CSS2) (Calcium-dependent protease small subunit 2). | |||||
|
SORCN_MOUSE
|
||||||
| NC score | 0.717150 (rank : 22) | θ value | 4.15078e-21 (rank : 24) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q6P069, Q3UKC5, Q9CR38, Q9D7V8 | Gene names | Sri | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorcin. | |||||
|
GRAN_MOUSE
|
||||||
| NC score | 0.716613 (rank : 23) | θ value | 1.42661e-21 (rank : 22) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8VC88, Q8BL53, Q8K3Z6 | Gene names | Gca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Grancalcin. | |||||
|
SORCN_HUMAN
|
||||||
| NC score | 0.712365 (rank : 24) | θ value | 4.15078e-21 (rank : 23) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P30626 | Gene names | SRI | |||
|
Domain Architecture |

|
|||||
| Description | Sorcin (22 kDa protein) (CP-22) (V19). | |||||
|
GRAN_HUMAN
|
||||||
| NC score | 0.708926 (rank : 25) | θ value | 5.99374e-20 (rank : 25) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P28676 | Gene names | GCA, GCL | |||
|
Domain Architecture |

|
|||||
| Description | Grancalcin. | |||||
|
PEF1_MOUSE
|
||||||
| NC score | 0.667816 (rank : 26) | θ value | 1.86753e-13 (rank : 31) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q8BFY6, Q8VCT5, Q9CYW8, Q9D934 | Gene names | Pef1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peflin (PEF protein with a long N-terminal hydrophobic domain) (Penta- EF hand domain-containing protein 1). | |||||
|
PEF1_HUMAN
|
||||||
| NC score | 0.654497 (rank : 27) | θ value | 1.09485e-13 (rank : 30) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9UBV8 | Gene names | PEF1, ABP32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peflin (PEF protein with a long N-terminal hydrophobic domain) (Penta- EF hand domain-containing protein 1). | |||||
|
CAN7_MOUSE
|
||||||
| NC score | 0.647208 (rank : 28) | θ value | 2.88119e-14 (rank : 29) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9R1S8, Q9Z0P9 | Gene names | Capn7, Palbh | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-7 (EC 3.4.22.-) (PalB homolog) (PalBH). | |||||
|
CAN7_HUMAN
|
||||||
| NC score | 0.644967 (rank : 29) | θ value | 2.88119e-14 (rank : 28) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9Y6W3 | Gene names | CAPN7, PALBH | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-7 (EC 3.4.22.-) (PalB homolog) (PalBH). | |||||
|
PDCD6_HUMAN
|
||||||
| NC score | 0.597291 (rank : 30) | θ value | 9.90251e-15 (rank : 26) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | O75340 | Gene names | PDCD6, ALG2 | |||
|
Domain Architecture |

|
|||||
| Description | Programmed cell death protein 6 (Probable calcium-binding protein ALG- 2). | |||||
|
PDCD6_MOUSE
|
||||||
| NC score | 0.595394 (rank : 31) | θ value | 9.90251e-15 (rank : 27) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P12815, Q545I0, Q61145 | Gene names | Pdcd6, Alg2 | |||
|
Domain Architecture |

|
|||||
| Description | Programmed cell death protein 6 (Probable calcium-binding protein ALG- 2) (PMP41) (ALG-257). | |||||
|
CALL5_HUMAN
|
||||||
| NC score | 0.208335 (rank : 32) | θ value | 1.87187e-05 (rank : 32) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9NZT1, Q8IXU8 | Gene names | CALML5, CLSP | |||
|
Domain Architecture |

|
|||||
| Description | Calmodulin-like protein 5 (Calmodulin-like skin protein). | |||||
|
CALM4_MOUSE
|
||||||
| NC score | 0.165648 (rank : 33) | θ value | 0.0113563 (rank : 35) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 229 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q9JM83, Q9CR31, Q9D1E9 | Gene names | Calm4, Dd112 | |||
|
Domain Architecture |

|
|||||
| Description | Calmodulin-4 (Calcium-binding protein Dd112). | |||||
|
CETN1_HUMAN
|
||||||
| NC score | 0.138021 (rank : 34) | θ value | 1.81305 (rank : 56) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q12798 | Gene names | CETN1, CEN1, CETN | |||
|
Domain Architecture |

|
|||||
| Description | Centrin-1 (Caltractin isoform 2). | |||||
|
CALM_HUMAN
|
||||||
| NC score | 0.135356 (rank : 35) | θ value | 0.00134147 (rank : 33) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P62158, P02593, P70667, P99014, Q13942, Q53S29, Q61379, Q61380, Q96HK3 | Gene names | CALM1, CALM, CAM, CAM1 | |||
|
Domain Architecture |

|
|||||
| Description | Calmodulin (CaM). | |||||
|
CALM_MOUSE
|
||||||
| NC score | 0.135356 (rank : 36) | θ value | 0.00134147 (rank : 34) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P62204, P02593, P70667, P99014, Q61379, Q61380 | Gene names | Calm1, Calm, Cam, Cam1 | |||
|
Domain Architecture |

|
|||||
| Description | Calmodulin (CaM). | |||||
|
CALL3_HUMAN
|
||||||
| NC score | 0.128063 (rank : 37) | θ value | 0.163984 (rank : 38) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P27482 | Gene names | CALML3 | |||
|
Domain Architecture |

|
|||||
| Description | Calmodulin-like protein 3 (Calmodulin-related protein NB-1) (CaM-like protein) (CLP). | |||||
|
TNNC1_MOUSE
|
||||||
| NC score | 0.126071 (rank : 38) | θ value | 5.27518 (rank : 74) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | P19123 | Gene names | Tnnc1, Tncc | |||
|
Domain Architecture |

|
|||||
| Description | Troponin C, slow skeletal and cardiac muscles (TN-C). | |||||
|
CETN1_MOUSE
|
||||||
| NC score | 0.125688 (rank : 39) | θ value | 2.36792 (rank : 61) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P41209, Q3V119, Q9D9G9, Q9DAL6 | Gene names | Cetn1, Calt | |||
|
Domain Architecture |

|
|||||
| Description | Centrin-1 (Caltractin). | |||||
|
TNNC1_HUMAN
|
||||||
| NC score | 0.119822 (rank : 40) | θ value | 6.88961 (rank : 82) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P63316, O14800, P02590, P04463 | Gene names | TNNC1, TNNC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Troponin C, slow skeletal and cardiac muscles (TN-C). | |||||
|
CF103_HUMAN
|
||||||
| NC score | 0.119476 (rank : 41) | θ value | θ > 10 (rank : 109) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8N7X0, Q5T402 | Gene names | C6orf103 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf103. | |||||
|
CETN2_MOUSE
|
||||||
| NC score | 0.118502 (rank : 42) | θ value | 0.21417 (rank : 39) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9R1K9, Q3UBB4, Q9CWM0 | Gene names | Cetn2, Calt | |||
|
Domain Architecture |

|
|||||
| Description | Centrin-2 (Caltractin isoform 1). | |||||
|
CABP4_HUMAN
|
||||||
| NC score | 0.117494 (rank : 43) | θ value | θ > 10 (rank : 99) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P57796, Q8N4Z2, Q8WWY5 | Gene names | CABP4 | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-binding protein 4 (CaBP4). | |||||
|
CETN2_HUMAN
|
||||||
| NC score | 0.114027 (rank : 44) | θ value | 0.365318 (rank : 43) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P41208, Q53XW1 | Gene names | CETN2, CALT, CEN2 | |||
|
Domain Architecture |

|
|||||
| Description | Centrin-2 (Caltractin isoform 1). | |||||
|
TNNC2_MOUSE
|
||||||
| NC score | 0.111370 (rank : 45) | θ value | 0.47712 (rank : 46) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P20801 | Gene names | Tnnc2, Tncs | |||
|
Domain Architecture |

|
|||||
| Description | Troponin C, skeletal muscle (STNC). | |||||
|
CALB2_HUMAN
|
||||||
| NC score | 0.110548 (rank : 46) | θ value | 0.279714 (rank : 40) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P22676, Q53HD2, Q96BK4 | Gene names | CALB2, CAB29 | |||
|
Domain Architecture |

|
|||||
| Description | Calretinin (CR) (29 kDa calbindin). | |||||
|
CALB1_MOUSE
|
||||||
| NC score | 0.109555 (rank : 47) | θ value | 0.0961366 (rank : 37) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P12658, Q545N6 | Gene names | Calb1 | |||
|
Domain Architecture |

|
|||||
| Description | Calbindin (Vitamin D-dependent calcium-binding protein, avian-type) (Calbindin D28) (D-28K) (Spot 35 protein) (PCD-29). | |||||
|
TNNC2_HUMAN
|
||||||
| NC score | 0.107718 (rank : 48) | θ value | 0.62314 (rank : 48) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P02585 | Gene names | TNNC2 | |||
|
Domain Architecture |

|
|||||
| Description | Troponin C, skeletal muscle. | |||||
|
CALB1_HUMAN
|
||||||
| NC score | 0.105003 (rank : 49) | θ value | 0.0961366 (rank : 36) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P05937 | Gene names | CALB1, CAB27 | |||
|
Domain Architecture |

|
|||||
| Description | Calbindin (Vitamin D-dependent calcium-binding protein, avian-type) (Calbindin D28) (D-28K). | |||||
|
SEGN_HUMAN
|
||||||
| NC score | 0.094857 (rank : 50) | θ value | 1.81305 (rank : 58) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O76038, Q5VV44, Q96QV7, Q9UJF6 | Gene names | SCGN, SECRET | |||
|
Domain Architecture |

|
|||||
| Description | Secretagogin. | |||||
|
CABP1_MOUSE
|
||||||
| NC score | 0.094403 (rank : 51) | θ value | 0.365318 (rank : 42) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9JLK7, Q9JLK6 | Gene names | Cabp1 | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-binding protein 1 (CaBP1). | |||||
|
CETN3_MOUSE
|
||||||
| NC score | 0.094395 (rank : 52) | θ value | θ > 10 (rank : 108) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | O35648 | Gene names | Cetn3, Cen3 | |||
|
Domain Architecture |

|
|||||
| Description | Centrin-3. | |||||
|
CETN3_HUMAN
|
||||||
| NC score | 0.094115 (rank : 53) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | O15182, Q9BS23 | Gene names | CETN3, CEN3 | |||
|
Domain Architecture |

|
|||||
| Description | Centrin-3. | |||||
|
CABP1_HUMAN
|
||||||
| NC score | 0.093825 (rank : 54) | θ value | 0.365318 (rank : 41) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9NZU7, O95663, Q9NZU8 | Gene names | CABP1 | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-binding protein 1 (CaBP1) (Calbrain). | |||||
|
CABP2_MOUSE
|
||||||
| NC score | 0.089074 (rank : 55) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9JLK4, Q9JLK5 | Gene names | Cabp2 | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-binding protein 2 (CaBP2). | |||||
|
CABP7_HUMAN
|
||||||
| NC score | 0.088847 (rank : 56) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q86V35 | Gene names | CABP7 | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-binding protein 7 (CaBP7). | |||||
|
CABP7_MOUSE
|
||||||
| NC score | 0.088847 (rank : 57) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q91ZM8 | Gene names | Cabp7 | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-binding protein 7 (CaBP7). | |||||
|
CABP4_MOUSE
|
||||||
| NC score | 0.087533 (rank : 58) | θ value | 3.0926 (rank : 65) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8VHC5 | Gene names | Cabp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium-binding protein 4 (CaBP4). | |||||
|
PLSI_HUMAN
|
||||||
| NC score | 0.086950 (rank : 59) | θ value | 2.36792 (rank : 63) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q14651 | Gene names | PLS1 | |||
|
Domain Architecture |

|
|||||
| Description | Plastin-1 (I-plastin) (Intestine-specific plastin). | |||||
|
CABP2_HUMAN
|
||||||
| NC score | 0.086256 (rank : 60) | θ value | 5.27518 (rank : 70) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9NPB3 | Gene names | CABP2 | |||
|
Domain Architecture |
|
|||||
| Description | Calcium-binding protein 2 (CaBP2). | |||||
|
CABP5_HUMAN
|
||||||
| NC score | 0.086061 (rank : 61) | θ value | 2.36792 (rank : 60) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9NP86 | Gene names | CABP5 | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-binding protein 5 (CaBP5). | |||||
|
CANB1_MOUSE
|
||||||
| NC score | 0.085333 (rank : 62) | θ value | 8.99809 (rank : 86) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q63810 | Gene names | Ppp3r1, Cnb | |||
|
Domain Architecture |

|
|||||
| Description | Calcineurin subunit B isoform 1 (Protein phosphatase 2B regulatory subunit 1) (Protein phosphatase 3 regulatory subunit B alpha isoform 1). | |||||
|
CANB1_HUMAN
|
||||||
| NC score | 0.085329 (rank : 63) | θ value | 8.99809 (rank : 85) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P63098, P06705, P15117, Q08044 | Gene names | PPP3R1, CNA2, CNB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcineurin subunit B isoform 1 (Protein phosphatase 2B regulatory subunit 1) (Protein phosphatase 3 regulatory subunit B alpha isoform 1). | |||||
|
CALB2_MOUSE
|
||||||
| NC score | 0.082844 (rank : 64) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q08331, Q60964, Q9JM81 | Gene names | Calb2 | |||
|
Domain Architecture |

|
|||||
| Description | Calretinin (CR). | |||||
|
CALL6_HUMAN
|
||||||
| NC score | 0.082346 (rank : 65) | θ value | 0.813845 (rank : 50) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8TD86, Q6Q2C4 | Gene names | CALML6, CAGLP, CALGP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calmodulin-like protein 6 (Calglandulin-like protein). | |||||
|
SEGN_MOUSE
|
||||||
| NC score | 0.081150 (rank : 66) | θ value | 2.36792 (rank : 64) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q91WD9 | Gene names | Scgn | |||
|
Domain Architecture |

|
|||||
| Description | Secretagogin. | |||||
|
CANB2_MOUSE
|
||||||
| NC score | 0.073929 (rank : 67) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q63811 | Gene names | Ppp3r2 | |||
|
Domain Architecture |

|
|||||
| Description | Calcineurin subunit B isoform 2 (Protein phosphatase 2B regulatory subunit 2) (Protein phosphatase 3 regulatory subunit B beta isoform). | |||||
|
CANB2_HUMAN
|
||||||
| NC score | 0.071551 (rank : 68) | θ value | 8.99809 (rank : 87) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q96LZ3, Q7Z4V8, Q8WYJ4 | Gene names | PPP3R2, CBLP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcineurin subunit B isoform 2 (Protein phosphatase 2B regulatory subunit 2) (Protein phosphatase 3 regulatory subunit B beta isoform) (Calcineurin B-like protein) (CBLP) (CNBII). | |||||
|
CABP5_MOUSE
|
||||||
| NC score | 0.070548 (rank : 69) | θ value | 8.99809 (rank : 84) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9JLK3 | Gene names | Cabp5 | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-binding protein 5 (CaBP5). | |||||
|
CAYP1_HUMAN
|
||||||
| NC score | 0.069588 (rank : 70) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q13938, Q3MJA7, Q8WUZ3 | Gene names | CAPS | |||
|
Domain Architecture |

|
|||||
| Description | Calcyphosin (Calcyphosine). | |||||
|
CABP3_HUMAN
|
||||||
| NC score | 0.067293 (rank : 71) | θ value | θ > 10 (rank : 98) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9NZU6, Q9NYY0, Q9NZT9 | Gene names | CABP3 | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-binding protein 3 (CaBP3). | |||||
|
CALN1_HUMAN
|
||||||
| NC score | 0.057394 (rank : 72) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9BXU9 | Gene names | CALN1, CABP8 | |||
|
Domain Architecture |

|
|||||
| Description | Calneuron-1 (Calcium-binding protein CaBP8). | |||||
|
CALN1_MOUSE
|
||||||
| NC score | 0.057305 (rank : 73) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9JJG7 | Gene names | Caln1 | |||
|
Domain Architecture |

|
|||||
| Description | Calneuron-1. | |||||
|
PRVA_MOUSE
|
||||||
| NC score | 0.056054 (rank : 74) | θ value | θ > 10 (rank : 110) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P32848 | Gene names | Pvalb, Pva | |||
|
Domain Architecture |

|
|||||
| Description | Parvalbumin alpha. | |||||
|
EFHB_HUMAN
|
||||||
| NC score | 0.054952 (rank : 75) | θ value | 1.06291 (rank : 52) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8N7U6, Q6ZWK9, Q8IV58, Q96LQ6 | Gene names | EFHB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EF-hand domain-containing family member B. | |||||
|
EFHB_MOUSE
|
||||||
| NC score | 0.047819 (rank : 76) | θ value | 0.813845 (rank : 51) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8CDU5 | Gene names | Efhb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EF-hand domain-containing family member B. | |||||
|
ONCO_MOUSE
|
||||||
| NC score | 0.046352 (rank : 77) | θ value | 8.99809 (rank : 94) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P51879, Q62004 | Gene names | Ocm | |||
|
Domain Architecture |

|
|||||
| Description | Oncomodulin (OM) (Parvalbumin beta). | |||||
|
EFCB1_HUMAN
|
||||||
| NC score | 0.028802 (rank : 78) | θ value | 3.0926 (rank : 66) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9HAE3 | Gene names | EFCAB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EF-hand calcium-binding domain-containing protein 1. | |||||
|
SPTA2_HUMAN
|
||||||
| NC score | 0.026065 (rank : 79) | θ value | 0.47712 (rank : 45) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 888 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q13813, Q13186, Q15324, Q16606, Q59EF1, Q5VXV5, Q5VXV6, Q7Z6M5, Q9P0V0 | Gene names | SPTAN1, SPTA2 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin alpha chain, brain (Spectrin, non-erythroid alpha chain) (Alpha-II spectrin) (Fodrin alpha chain). | |||||
|
ACTN1_MOUSE
|
||||||
| NC score | 0.025760 (rank : 80) | θ value | 4.03905 (rank : 67) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q7TPR4 | Gene names | Actn1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-actinin-1 (Alpha-actinin cytoskeletal isoform) (Non-muscle alpha-actinin-1) (F-actin cross-linking protein). | |||||
|
ACTN1_HUMAN
|
||||||
| NC score | 0.024920 (rank : 81) | θ value | 5.27518 (rank : 69) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P12814, Q9BTN1 | Gene names | ACTN1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-actinin-1 (Alpha-actinin cytoskeletal isoform) (Non-muscle alpha-actinin-1) (F-actin cross-linking protein). | |||||
|
CTNL1_HUMAN
|
||||||
| NC score | 0.023040 (rank : 82) | θ value | 0.47712 (rank : 44) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UBT7, O76084, Q53FQ2, Q5JTQ7, Q5JTQ8, Q9Y401 | Gene names | CTNNAL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alpha-catulin (Catenin alpha-like protein 1) (Alpha-catenin-related protein) (ACRP). | |||||
|
RYR2_HUMAN
|
||||||
| NC score | 0.019782 (rank : 83) | θ value | 6.88961 (rank : 80) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q92736, Q15411, Q546N8 | Gene names | RYR2 | |||
|
Domain Architecture |

|
|||||
| Description | Ryanodine receptor 2 (Cardiac muscle-type ryanodine receptor) (RyR2) (RYR-2) (Cardiac muscle ryanodine receptor-calcium release channel) (hRYR-2). | |||||
|
NOX5_HUMAN
|
||||||
| NC score | 0.016718 (rank : 84) | θ value | 6.88961 (rank : 78) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96PH1, Q8TEQ1, Q8TER4, Q96PH2, Q96PJ8, Q96PJ9, Q9H6E0, Q9HAM8 | Gene names | NOX5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NADPH oxidase 5 (EC 1.6.3.-). | |||||
|
ELF2_MOUSE
|
||||||
| NC score | 0.012592 (rank : 85) | θ value | 1.38821 (rank : 55) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9JHC9, Q6NST2, Q8BTX8, Q9JHC7, Q9JHC8, Q9JHD0 | Gene names | Elf2, Nerf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-2 (E74-like factor 2) (New ETS- related factor). | |||||
|
TFAM_HUMAN
|
||||||
| NC score | 0.011265 (rank : 86) | θ value | 1.81305 (rank : 59) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q00059 | Gene names | TFAM, TCF6L2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor A, mitochondrial precursor (mtTFA) (Mitochondrial transcription factor 1) (MtTF1) (Transcription factor 6-like 2). | |||||
|
TSH1_MOUSE
|
||||||
| NC score | 0.010747 (rank : 87) | θ value | 0.47712 (rank : 47) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 675 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5DTH5, Q6PE60, Q8VD19, Q9JLD5 | Gene names | Tshz1, Kiaa4206, Sdccag33, Tsh1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 1 (Serologically defined colon cancer antigen 3 homolog). | |||||
|
ELF2_HUMAN
|
||||||
| NC score | 0.009515 (rank : 88) | θ value | 2.36792 (rank : 62) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q15723, Q15724, Q15725, Q6P1K5 | Gene names | ELF2, NERF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-2 (E74-like factor 2) (New ETS- related factor). | |||||
|
IF2B_MOUSE
|
||||||
| NC score | 0.009183 (rank : 89) | θ value | 5.27518 (rank : 71) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99L45, Q9CSH6, Q9CT12 | Gene names | Eif2s2 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2 subunit 2 (Eukaryotic translation initiation factor 2 subunit beta) (eIF-2-beta). | |||||
|
FANCM_HUMAN
|
||||||
| NC score | 0.008968 (rank : 90) | θ value | 1.81305 (rank : 57) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8IYD8, Q3YFH9, Q8N9X6, Q9HCH6 | Gene names | FANCM, KIAA1596 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fanconi anemia group M protein (EC 3.6.1.-) (ATP-dependent RNA helicase FANCM) (Protein FACM) (Fanconi anemia-associated polypeptide of 250 kDa) (FAAP250) (Protein Hef ortholog). | |||||
|
UBP48_MOUSE
|
||||||
| NC score | 0.008229 (rank : 91) | θ value | 0.62314 (rank : 49) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q3V0C5, Q3TM39, Q571J9, Q8VDJ1 | Gene names | Usp48, Kiaa4202 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 48 (EC 3.1.2.15) (Ubiquitin thioesterase 48) (Ubiquitin-specific-processing protease 48) (Deubiquitinating enzyme 48). | |||||
|
BRPF1_HUMAN
|
||||||
| NC score | 0.006090 (rank : 92) | θ value | 1.38821 (rank : 54) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P55201, Q9UHI0 | Gene names | BRPF1, BR140 | |||
|
Domain Architecture |

|
|||||
| Description | Peregrin (Bromodomain and PHD finger-containing protein 1) (BR140 protein). | |||||
|
LBXCO_MOUSE
|
||||||
| NC score | 0.005131 (rank : 93) | θ value | 6.88961 (rank : 76) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BX46, Q3UYA4, Q5W8I2, Q8C0T2 | Gene names | Lbxcor1, Corl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladybird homeobox corepressor 1 (Transcriptional corepressor Corl1). | |||||
|
RANB9_HUMAN
|
||||||
| NC score | 0.004755 (rank : 94) | θ value | 8.99809 (rank : 95) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 179 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96S59, O94764, Q6P3T7, Q7LBR2, Q7Z7F9 | Gene names | RANBP9, RANBPM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 9 (RanBP9) (RanBP7) (Ran-binding protein M) (RanBPM) (BPM90) (BPM-L). | |||||
|
MOR2A_MOUSE
|
||||||
| NC score | 0.004619 (rank : 95) | θ value | 8.99809 (rank : 92) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q69ZX6, Q5QNQ7, Q6P547 | Gene names | Morc2a, Kiaa0852, Zcwcc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORC family CW-type zinc finger protein 2A (Zinc finger CW-type coiled-coil domain protein 1). | |||||
|
MORC2_HUMAN
|
||||||
| NC score | 0.004577 (rank : 96) | θ value | 8.99809 (rank : 93) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y6X9, Q9UF28, Q9Y6V2 | Gene names | MORC2, KIAA0852, ZCWCC1 | |||
|
Domain Architecture |

|
|||||
| Description | MORC family CW-type zinc finger protein 2 (Zinc finger CW-type coiled- coil domain protein 1). | |||||
|
LRC17_MOUSE
|
||||||
| NC score | 0.004186 (rank : 97) | θ value | 1.06291 (rank : 53) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 268 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9CXD9 | Gene names | Lrrc17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 17 precursor. | |||||
|
PO121_HUMAN
|
||||||
| NC score | 0.003951 (rank : 98) | θ value | 6.88961 (rank : 79) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y2N3, O75115, Q9Y4S7 | Gene names | POM121, KIAA0618, NUP121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear envelope pore membrane protein POM 121 (Pore membrane protein of 121 kDa) (P145). | |||||
|
SIN3B_HUMAN
|
||||||
| NC score | 0.003643 (rank : 99) | θ value | 8.99809 (rank : 96) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O75182, Q68GC2, Q6P4B8, Q8TB34, Q9BSC8 | Gene names | SIN3B, KIAA0700 | |||
|
Domain Architecture |

|
|||||
| Description | Paired amphipathic helix protein Sin3b (Transcriptional corepressor Sin3b) (Histone deacetylase complex subunit Sin3b). | |||||
|
SFPQ_MOUSE
|
||||||
| NC score | 0.002962 (rank : 100) | θ value | 6.88961 (rank : 81) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8VIJ6, Q9ERW2 | Gene names | Sfpq | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, proline- and glutamine-rich (Polypyrimidine tract- binding protein-associated-splicing factor) (PTB-associated-splicing factor) (PSF) (DNA-binding p52/p100 complex, 100 kDa subunit). | |||||
|
MAP9_HUMAN
|
||||||
| NC score | 0.002134 (rank : 101) | θ value | 8.99809 (rank : 91) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 518 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q49MG5, Q4W5I7, Q68DU1, Q9H781, Q9H7B6 | Gene names | MAP9, ASAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 9 (Aster-associated protein). | |||||
|
BSN_MOUSE
|
||||||
| NC score | 0.001990 (rank : 102) | θ value | 4.03905 (rank : 68) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
DSG3_MOUSE
|
||||||
| NC score | 0.001375 (rank : 103) | θ value | 8.99809 (rank : 88) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O35902, Q8CB02, Q8CE48 | Gene names | Dsg3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Desmoglein-3 precursor (130 kDa pemphigus vulgaris antigen homolog). | |||||
|
LEO1_HUMAN
|
||||||
| NC score | 0.001233 (rank : 104) | θ value | 8.99809 (rank : 90) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
RBBP6_MOUSE
|
||||||
| NC score | 0.001050 (rank : 105) | θ value | 5.27518 (rank : 73) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
FOXD1_MOUSE
|
||||||
| NC score | 0.000100 (rank : 106) | θ value | 8.99809 (rank : 89) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q61345 | Gene names | Foxd1, Fkhl8, Freac4, Hfhbf2 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein D1 (Forkhead-related protein FKHL8) (Forkhead- related transcription factor 4) (FREAC-4) (Brain factor 2) (HFH-BF-2). | |||||
|
KI13B_HUMAN
|
||||||
| NC score | 0.000084 (rank : 107) | θ value | 6.88961 (rank : 75) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 492 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NQT8, O75134, Q9BYJ6 | Gene names | KIF13B, GAKIN, KIAA0639 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF13B (Kinesin-like protein GAKIN). | |||||
|
IWS1_MOUSE
|
||||||
| NC score | -0.000514 (rank : 108) | θ value | 5.27518 (rank : 72) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
LRRN1_HUMAN
|
||||||
| NC score | -0.000927 (rank : 109) | θ value | 6.88961 (rank : 77) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 518 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6UXK5, Q3LID5, Q8IYV5, Q9H8V1, Q9P231 | Gene names | LRRN1, KIAA1497 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats neuronal protein 1 precursor (Neuronal leucine- rich repeat protein 1) (NLRR-1). | |||||
|
ZN226_HUMAN
|
||||||
| NC score | -0.004666 (rank : 110) | θ value | 6.88961 (rank : 83) | |||
| Query Neighborhood Hits | 96 | Target Neighborhood Hits | 770 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NYT6, Q96TE6, Q9NS44 | Gene names | ZNF226 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 226. | |||||