Please be patient as the page loads
|
PRR6_HUMAN
|
||||||
| SwissProt Accessions | Q7Z7K6, Q3L8N5, Q8NFH6 | Gene names | PRR6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 6 (Nuclear protein p30). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
PRR6_HUMAN
|
||||||
| θ value | 1.0648e-109 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q7Z7K6, Q3L8N5, Q8NFH6 | Gene names | PRR6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 6 (Nuclear protein p30). | |||||
|
PRR6_MOUSE
|
||||||
| θ value | 1.94955e-95 (rank : 2) | NC score | 0.973580 (rank : 2) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9CXS4, Q5SX21, Q6PIA5 | Gene names | Prr6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 6. | |||||
|
NFH_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 3) | NC score | 0.033693 (rank : 15) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
MKL1_MOUSE
|
||||||
| θ value | 0.125558 (rank : 4) | NC score | 0.055645 (rank : 3) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 667 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8K4J6 | Gene names | Mkl1, Bsac | |||
|
Domain Architecture |

|
|||||
| Description | MKL/myocardin-like protein 1 (Myocardin-related transcription factor A) (MRTF-A) (Megakaryoblastic leukemia 1 protein homolog) (Basic SAP coiled-coil transcription activator). | |||||
|
MYO15_HUMAN
|
||||||
| θ value | 0.125558 (rank : 5) | NC score | 0.028371 (rank : 19) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9UKN7 | Gene names | MYO15A, MYO15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
SPEG_MOUSE
|
||||||
| θ value | 0.279714 (rank : 6) | NC score | 0.019128 (rank : 29) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
RGS3_HUMAN
|
||||||
| θ value | 1.81305 (rank : 7) | NC score | 0.029831 (rank : 17) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
JADE2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 8) | NC score | 0.024160 (rank : 23) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6ZQF7, Q3UHD5, Q6IE83 | Gene names | Phf15, Jade2, Kiaa0239 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-2 (PHD finger protein 15). | |||||
|
PI5PA_MOUSE
|
||||||
| θ value | 2.36792 (rank : 9) | NC score | 0.038510 (rank : 12) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P59644 | Gene names | Pib5pa | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4,5-bisphosphate 5-phosphatase A (EC 3.1.3.56). | |||||
|
GAK11_HUMAN
|
||||||
| θ value | 3.0926 (rank : 10) | NC score | 0.049572 (rank : 6) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P63145, Q9UKI1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_22q11.21 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K101 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 11) | NC score | 0.049452 (rank : 7) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q7LDI9, Q9UKH5, Q9Y6I1, Q9YNA6, Q9YNB0 | Gene names | ERVK6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_7p22.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K(HML-2.HOM) Gag protein) (HERV-K108 Gag protein) (HERV-K(C7) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK4_HUMAN
|
||||||
| θ value | 3.0926 (rank : 12) | NC score | 0.049750 (rank : 5) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 415 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P63126, Q9UKH4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_6q14.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K109 Gag protein) (HERV-K(C6) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK6_HUMAN
|
||||||
| θ value | 3.0926 (rank : 13) | NC score | 0.049336 (rank : 8) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P62685 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_8p23.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K115 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
MAST4_HUMAN
|
||||||
| θ value | 3.0926 (rank : 14) | NC score | 0.009596 (rank : 36) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 1804 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O15021, Q6ZN07, Q96LY3 | Gene names | MAST4, KIAA0303 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 4 (EC 2.7.11.1). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 3.0926 (rank : 15) | NC score | 0.036660 (rank : 13) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
USP9X_HUMAN
|
||||||
| θ value | 3.0926 (rank : 16) | NC score | 0.027050 (rank : 21) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q93008, O75550, Q8WWT3, Q8WX12 | Gene names | USP9X, DFFRX, USP9 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase FAF-X (EC 3.1.2.15) (Ubiquitin thioesterase FAF-X) (Ubiquitin-specific-processing protease FAF-X) (Deubiquitinating enzyme FAF-X) (Fat facets protein-related, X- linked) (Ubiquitin-specific protease 9, X chromosome). | |||||
|
GAK12_HUMAN
|
||||||
| θ value | 4.03905 (rank : 17) | NC score | 0.049125 (rank : 9) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P63130, Q9UKI0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q22 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K102 Gag protein) (HERV-K(III) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
PO6F1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 18) | NC score | 0.015416 (rank : 34) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q07916, Q91XK3 | Gene names | Pou6f1, Emb | |||
|
Domain Architecture |
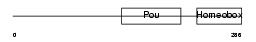
|
|||||
| Description | POU domain, class 6, transcription factor 1 (Octamer-binding transcription factor EMB) (Transcription regulatory protein MCP-1). | |||||
|
XKR7_HUMAN
|
||||||
| θ value | 4.03905 (rank : 19) | NC score | 0.020341 (rank : 27) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5GH72, Q9NUG5 | Gene names | XKR7, C20orf159, XRG7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | XK-related protein 7. | |||||
|
ZAR1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 20) | NC score | 0.034389 (rank : 14) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 254 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80SU3 | Gene names | Zar1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zygote arrest 1 (Oocyte-specific maternal effect factor). | |||||
|
ZN213_HUMAN
|
||||||
| θ value | 4.03905 (rank : 21) | NC score | 0.001904 (rank : 40) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 755 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O14771, Q96IS1 | Gene names | ZNF213 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 213 (Putative transcription factor CR53). | |||||
|
FOG1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 22) | NC score | 0.013416 (rank : 35) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 1237 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O35615 | Gene names | Zfpm1, Fog, Fog1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZFPM1 (Zinc finger protein multitype 1) (Friend of GATA protein 1) (Friend of GATA-1) (FOG-1). | |||||
|
GAK5_HUMAN
|
||||||
| θ value | 5.27518 (rank : 23) | NC score | 0.048076 (rank : 10) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 429 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P62684 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19p13.11 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K113 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
NUFP1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 24) | NC score | 0.031052 (rank : 16) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UHK0, Q8WVM5, Q96SG1 | Gene names | NUFIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear fragile X mental retardation-interacting protein 1 (Nuclear FMRP-interacting protein 1). | |||||
|
PER3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 25) | NC score | 0.018998 (rank : 30) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P56645, Q5H8X4, Q969K6, Q96S77, Q96S78, Q9C0J3, Q9NSP9, Q9UGU8 | Gene names | PER3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (hPER3). | |||||
|
RTP3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 26) | NC score | 0.041009 (rank : 11) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5QGU6, Q8VDC2 | Gene names | Rtp3, Tmem7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Receptor-transporting protein 3 (Transmembrane protein 7). | |||||
|
USP9X_MOUSE
|
||||||
| θ value | 5.27518 (rank : 27) | NC score | 0.025399 (rank : 22) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P70398, Q62497 | Gene names | Usp9x, Fafl, Fam | |||
|
Domain Architecture |

|
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase FAF-X (EC 3.1.2.15) (Ubiquitin thioesterase FAF-X) (Ubiquitin-specific-processing protease FAF-X) (Deubiquitinating enzyme FAF-X) (Fat facets protein-related, X- linked) (Ubiquitin-specific protease 9, X chromosome) (Ubiquitin carboxyl-terminal hydrolase FAM) (Fat facets homolog). | |||||
|
ELL_HUMAN
|
||||||
| θ value | 6.88961 (rank : 28) | NC score | 0.018126 (rank : 31) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P55199 | Gene names | ELL, C19orf17 | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase II elongation factor ELL (Eleven-nineteen lysine-rich leukemia protein). | |||||
|
HSF1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 29) | NC score | 0.022214 (rank : 26) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q00613 | Gene names | HSF1, HSTF1 | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock factor protein 1 (HSF 1) (Heat shock transcription factor 1) (HSTF 1). | |||||
|
MILK2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 30) | NC score | 0.027078 (rank : 20) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8IY33, Q7RTP4, Q7Z655, Q8TEQ4, Q9H5F9 | Gene names | ||||
|
Domain Architecture |
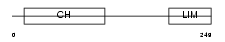
|
|||||
| Description | MICAL-like protein 2. | |||||
|
PAK4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 31) | NC score | 0.004463 (rank : 39) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 1098 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BTW9, Q6ZPX0, Q80Z97, Q9CS71 | Gene names | Pak4, Kiaa1142 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase PAK 4 (EC 2.7.11.1) (p21-activated kinase 4) (PAK-4). | |||||
|
PCIF1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 32) | NC score | 0.022405 (rank : 25) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H4Z3, Q9NT85 | Gene names | PCIF1, C20orf67 | |||
|
Domain Architecture |
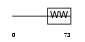
|
|||||
| Description | Phosphorylated CTD-interacting factor 1. | |||||
|
TMPS9_MOUSE
|
||||||
| θ value | 6.88961 (rank : 33) | NC score | 0.004651 (rank : 38) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P69525 | Gene names | Tmprss9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protease, serine 9 (EC 3.4.21.-) (Polyserase-1) (Polyserine protease 1) (Polyserase-I) [Contains: Serase-1; Serase-2; Serase-3]. | |||||
|
WBP1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 34) | NC score | 0.022977 (rank : 24) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P97764, Q8WUP5, Q99J20 | Gene names | Wbp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 1 (WBP-1). | |||||
|
AATK_HUMAN
|
||||||
| θ value | 8.99809 (rank : 35) | NC score | 0.008018 (rank : 37) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 1243 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6ZMQ8, O75136, Q6ZN31, Q86X28 | Gene names | AATK, KIAA0641 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Apoptosis-associated tyrosine-protein kinase (EC 2.7.10.2) (AATYK) (Brain apoptosis-associated tyrosine kinase) (CDK5-binding protein) (p35-binding protein) (p35BP). | |||||
|
KCD18_HUMAN
|
||||||
| θ value | 8.99809 (rank : 36) | NC score | 0.017705 (rank : 32) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6PI47, Q53T21, Q6NW26, Q6PCD8, Q8N9B7, Q96N73 | Gene names | KCTD18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD18. | |||||
|
RGS3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 37) | NC score | 0.029730 (rank : 18) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
SAPS1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 38) | NC score | 0.020089 (rank : 28) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UPN7, Q504V2, Q6NVJ6, Q9BU97 | Gene names | SAPS1, KIAA1115 | |||
|
Domain Architecture |

|
|||||
| Description | SAPS domain family member 1. | |||||
|
SFR14_MOUSE
|
||||||
| θ value | 8.99809 (rank : 39) | NC score | 0.017215 (rank : 33) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CH09, Q6PG19, Q80UY8, Q8BY32, Q8CFM0 | Gene names | Sfrs14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative splicing factor, arginine/serine-rich 14 (Arginine/serine- rich-splicing factor 14). | |||||
|
CN155_HUMAN
|
||||||
| θ value | θ > 10 (rank : 40) | NC score | 0.055003 (rank : 4) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q5H9T9, Q5H9U7, Q86YI2, Q9H0J3 | Gene names | C14orf155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf155. | |||||
|
PRR6_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 1.0648e-109 (rank : 1) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q7Z7K6, Q3L8N5, Q8NFH6 | Gene names | PRR6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 6 (Nuclear protein p30). | |||||
|
PRR6_MOUSE
|
||||||
| NC score | 0.973580 (rank : 2) | θ value | 1.94955e-95 (rank : 2) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9CXS4, Q5SX21, Q6PIA5 | Gene names | Prr6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 6. | |||||
|
MKL1_MOUSE
|
||||||
| NC score | 0.055645 (rank : 3) | θ value | 0.125558 (rank : 4) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 667 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8K4J6 | Gene names | Mkl1, Bsac | |||
|
Domain Architecture |

|
|||||
| Description | MKL/myocardin-like protein 1 (Myocardin-related transcription factor A) (MRTF-A) (Megakaryoblastic leukemia 1 protein homolog) (Basic SAP coiled-coil transcription activator). | |||||
|
CN155_HUMAN
|
||||||
| NC score | 0.055003 (rank : 4) | θ value | θ > 10 (rank : 40) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q5H9T9, Q5H9U7, Q86YI2, Q9H0J3 | Gene names | C14orf155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf155. | |||||
|
GAK4_HUMAN
|
||||||
| NC score | 0.049750 (rank : 5) | θ value | 3.0926 (rank : 12) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 415 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P63126, Q9UKH4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_6q14.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K109 Gag protein) (HERV-K(C6) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK11_HUMAN
|
||||||
| NC score | 0.049572 (rank : 6) | θ value | 3.0926 (rank : 10) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P63145, Q9UKI1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_22q11.21 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K101 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK2_HUMAN
|
||||||
| NC score | 0.049452 (rank : 7) | θ value | 3.0926 (rank : 11) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q7LDI9, Q9UKH5, Q9Y6I1, Q9YNA6, Q9YNB0 | Gene names | ERVK6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_7p22.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K(HML-2.HOM) Gag protein) (HERV-K108 Gag protein) (HERV-K(C7) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK6_HUMAN
|
||||||
| NC score | 0.049336 (rank : 8) | θ value | 3.0926 (rank : 13) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P62685 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_8p23.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K115 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK12_HUMAN
|
||||||
| NC score | 0.049125 (rank : 9) | θ value | 4.03905 (rank : 17) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P63130, Q9UKI0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q22 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K102 Gag protein) (HERV-K(III) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK5_HUMAN
|
||||||
| NC score | 0.048076 (rank : 10) | θ value | 5.27518 (rank : 23) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 429 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P62684 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19p13.11 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K113 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
RTP3_MOUSE
|
||||||
| NC score | 0.041009 (rank : 11) | θ value | 5.27518 (rank : 26) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5QGU6, Q8VDC2 | Gene names | Rtp3, Tmem7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Receptor-transporting protein 3 (Transmembrane protein 7). | |||||
|
PI5PA_MOUSE
|
||||||
| NC score | 0.038510 (rank : 12) | θ value | 2.36792 (rank : 9) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P59644 | Gene names | Pib5pa | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4,5-bisphosphate 5-phosphatase A (EC 3.1.3.56). | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.036660 (rank : 13) | θ value | 3.0926 (rank : 15) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
ZAR1_MOUSE
|
||||||
| NC score | 0.034389 (rank : 14) | θ value | 4.03905 (rank : 20) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 254 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80SU3 | Gene names | Zar1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zygote arrest 1 (Oocyte-specific maternal effect factor). | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.033693 (rank : 15) | θ value | 0.00390308 (rank : 3) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
NUFP1_HUMAN
|
||||||
| NC score | 0.031052 (rank : 16) | θ value | 5.27518 (rank : 24) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UHK0, Q8WVM5, Q96SG1 | Gene names | NUFIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear fragile X mental retardation-interacting protein 1 (Nuclear FMRP-interacting protein 1). | |||||
|
RGS3_HUMAN
|
||||||
| NC score | 0.029831 (rank : 17) | θ value | 1.81305 (rank : 7) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
RGS3_MOUSE
|
||||||
| NC score | 0.029730 (rank : 18) | θ value | 8.99809 (rank : 37) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
MYO15_HUMAN
|
||||||
| NC score | 0.028371 (rank : 19) | θ value | 0.125558 (rank : 5) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9UKN7 | Gene names | MYO15A, MYO15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
MILK2_HUMAN
|
||||||
| NC score | 0.027078 (rank : 20) | θ value | 6.88961 (rank : 30) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8IY33, Q7RTP4, Q7Z655, Q8TEQ4, Q9H5F9 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | MICAL-like protein 2. | |||||
|
USP9X_HUMAN
|
||||||
| NC score | 0.027050 (rank : 21) | θ value | 3.0926 (rank : 16) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q93008, O75550, Q8WWT3, Q8WX12 | Gene names | USP9X, DFFRX, USP9 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase FAF-X (EC 3.1.2.15) (Ubiquitin thioesterase FAF-X) (Ubiquitin-specific-processing protease FAF-X) (Deubiquitinating enzyme FAF-X) (Fat facets protein-related, X- linked) (Ubiquitin-specific protease 9, X chromosome). | |||||
|
USP9X_MOUSE
|
||||||
| NC score | 0.025399 (rank : 22) | θ value | 5.27518 (rank : 27) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P70398, Q62497 | Gene names | Usp9x, Fafl, Fam | |||
|
Domain Architecture |

|
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase FAF-X (EC 3.1.2.15) (Ubiquitin thioesterase FAF-X) (Ubiquitin-specific-processing protease FAF-X) (Deubiquitinating enzyme FAF-X) (Fat facets protein-related, X- linked) (Ubiquitin-specific protease 9, X chromosome) (Ubiquitin carboxyl-terminal hydrolase FAM) (Fat facets homolog). | |||||
|
JADE2_MOUSE
|
||||||
| NC score | 0.024160 (rank : 23) | θ value | 2.36792 (rank : 8) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6ZQF7, Q3UHD5, Q6IE83 | Gene names | Phf15, Jade2, Kiaa0239 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-2 (PHD finger protein 15). | |||||
|
WBP1_MOUSE
|
||||||
| NC score | 0.022977 (rank : 24) | θ value | 6.88961 (rank : 34) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P97764, Q8WUP5, Q99J20 | Gene names | Wbp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 1 (WBP-1). | |||||
|
PCIF1_HUMAN
|
||||||
| NC score | 0.022405 (rank : 25) | θ value | 6.88961 (rank : 32) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H4Z3, Q9NT85 | Gene names | PCIF1, C20orf67 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphorylated CTD-interacting factor 1. | |||||
|
HSF1_HUMAN
|
||||||
| NC score | 0.022214 (rank : 26) | θ value | 6.88961 (rank : 29) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q00613 | Gene names | HSF1, HSTF1 | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock factor protein 1 (HSF 1) (Heat shock transcription factor 1) (HSTF 1). | |||||
|
XKR7_HUMAN
|
||||||
| NC score | 0.020341 (rank : 27) | θ value | 4.03905 (rank : 19) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5GH72, Q9NUG5 | Gene names | XKR7, C20orf159, XRG7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | XK-related protein 7. | |||||
|
SAPS1_HUMAN
|
||||||
| NC score | 0.020089 (rank : 28) | θ value | 8.99809 (rank : 38) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UPN7, Q504V2, Q6NVJ6, Q9BU97 | Gene names | SAPS1, KIAA1115 | |||
|
Domain Architecture |

|
|||||
| Description | SAPS domain family member 1. | |||||
|
SPEG_MOUSE
|
||||||
| NC score | 0.019128 (rank : 29) | θ value | 0.279714 (rank : 6) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
PER3_HUMAN
|
||||||
| NC score | 0.018998 (rank : 30) | θ value | 5.27518 (rank : 25) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P56645, Q5H8X4, Q969K6, Q96S77, Q96S78, Q9C0J3, Q9NSP9, Q9UGU8 | Gene names | PER3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (hPER3). | |||||
|
ELL_HUMAN
|
||||||
| NC score | 0.018126 (rank : 31) | θ value | 6.88961 (rank : 28) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P55199 | Gene names | ELL, C19orf17 | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase II elongation factor ELL (Eleven-nineteen lysine-rich leukemia protein). | |||||
|
KCD18_HUMAN
|
||||||
| NC score | 0.017705 (rank : 32) | θ value | 8.99809 (rank : 36) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6PI47, Q53T21, Q6NW26, Q6PCD8, Q8N9B7, Q96N73 | Gene names | KCTD18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD18. | |||||
|
SFR14_MOUSE
|
||||||
| NC score | 0.017215 (rank : 33) | θ value | 8.99809 (rank : 39) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CH09, Q6PG19, Q80UY8, Q8BY32, Q8CFM0 | Gene names | Sfrs14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative splicing factor, arginine/serine-rich 14 (Arginine/serine- rich-splicing factor 14). | |||||
|
PO6F1_MOUSE
|
||||||
| NC score | 0.015416 (rank : 34) | θ value | 4.03905 (rank : 18) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q07916, Q91XK3 | Gene names | Pou6f1, Emb | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 6, transcription factor 1 (Octamer-binding transcription factor EMB) (Transcription regulatory protein MCP-1). | |||||
|
FOG1_MOUSE
|
||||||
| NC score | 0.013416 (rank : 35) | θ value | 5.27518 (rank : 22) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 1237 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O35615 | Gene names | Zfpm1, Fog, Fog1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZFPM1 (Zinc finger protein multitype 1) (Friend of GATA protein 1) (Friend of GATA-1) (FOG-1). | |||||
|
MAST4_HUMAN
|
||||||
| NC score | 0.009596 (rank : 36) | θ value | 3.0926 (rank : 14) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 1804 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O15021, Q6ZN07, Q96LY3 | Gene names | MAST4, KIAA0303 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 4 (EC 2.7.11.1). | |||||
|
AATK_HUMAN
|
||||||
| NC score | 0.008018 (rank : 37) | θ value | 8.99809 (rank : 35) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 1243 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6ZMQ8, O75136, Q6ZN31, Q86X28 | Gene names | AATK, KIAA0641 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Apoptosis-associated tyrosine-protein kinase (EC 2.7.10.2) (AATYK) (Brain apoptosis-associated tyrosine kinase) (CDK5-binding protein) (p35-binding protein) (p35BP). | |||||
|
TMPS9_MOUSE
|
||||||
| NC score | 0.004651 (rank : 38) | θ value | 6.88961 (rank : 33) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P69525 | Gene names | Tmprss9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protease, serine 9 (EC 3.4.21.-) (Polyserase-1) (Polyserine protease 1) (Polyserase-I) [Contains: Serase-1; Serase-2; Serase-3]. | |||||
|
PAK4_MOUSE
|
||||||
| NC score | 0.004463 (rank : 39) | θ value | 6.88961 (rank : 31) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 1098 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BTW9, Q6ZPX0, Q80Z97, Q9CS71 | Gene names | Pak4, Kiaa1142 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase PAK 4 (EC 2.7.11.1) (p21-activated kinase 4) (PAK-4). | |||||
|
ZN213_HUMAN
|
||||||
| NC score | 0.001904 (rank : 40) | θ value | 4.03905 (rank : 21) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 755 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O14771, Q96IS1 | Gene names | ZNF213 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 213 (Putative transcription factor CR53). | |||||