Please be patient as the page loads
|
PPR3A_HUMAN
|
||||||
| SwissProt Accessions | Q16821, O43476, Q75LN8, Q7KYM8, Q86UI6 | Gene names | PPP1R3A, PP1G | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein phosphatase 1 regulatory subunit 3A (Protein phosphatase 1 glycogen-associated regulatory subunit) (Protein phosphatase type-1 glycogen targeting subunit). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
PPR3A_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q16821, O43476, Q75LN8, Q7KYM8, Q86UI6 | Gene names | PPP1R3A, PP1G | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein phosphatase 1 regulatory subunit 3A (Protein phosphatase 1 glycogen-associated regulatory subunit) (Protein phosphatase type-1 glycogen targeting subunit). | |||||
|
PPR3A_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.963828 (rank : 2) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99MR9, Q8BUJ4, Q8BUL0 | Gene names | Ppp1r3a, Pp1g | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein phosphatase 1 regulatory subunit 3A (Protein phosphatase 1 glycogen-associated regulatory subunit) (Protein phosphatase type-1 glycogen targeting subunit) (RGL). | |||||
|
PPR3D_HUMAN
|
||||||
| θ value | 4.02038e-16 (rank : 3) | NC score | 0.621171 (rank : 3) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O95685, Q6DK02 | Gene names | PPP1R3D, PPP1R6 | |||
|
Domain Architecture |

|
|||||
| Description | Protein phosphatase 1 regulatory subunit 3D (Protein phosphatase 1, regulatory subunit 6) (Protein phosphatase 1-binding subunit R6). | |||||
|
NFM_MOUSE
|
||||||
| θ value | 0.00298849 (rank : 4) | NC score | 0.029421 (rank : 17) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1159 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P08553, Q61961 | Gene names | Nef3, Nefm, Nfm | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M). | |||||
|
SYNE2_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 5) | NC score | 0.041077 (rank : 10) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1323 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8WXH0, Q8N1S3, Q8NF49, Q8TER7, Q8WWW3, Q8WWW4, Q8WWW5, Q8WXH1, Q9NU50, Q9UFQ4, Q9Y2L4, Q9Y4R1 | Gene names | SYNE2, KIAA1011, NUA | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-2 (Nuclear envelope spectrin repeat protein 2) (Syne-2) (Synaptic nuclear envelope protein 2) (Nucleus and actin connecting element protein) (Protein NUANCE). | |||||
|
LAMA4_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 6) | NC score | 0.029123 (rank : 18) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 708 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q16363, Q14735, Q15335, Q4LE44, Q5SZG8, Q9UE18, Q9UJN9 | Gene names | LAMA4 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-4 chain precursor. | |||||
|
RBBP6_MOUSE
|
||||||
| θ value | 0.125558 (rank : 7) | NC score | 0.048605 (rank : 7) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
ATRX_MOUSE
|
||||||
| θ value | 0.163984 (rank : 8) | NC score | 0.042827 (rank : 9) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q61687 | Gene names | Atrx, Hp1bp2, Xnp | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked nuclear protein) (Heterochromatin protein 2) (HP1 alpha-interacting protein) (HP1-BP38 protein). | |||||
|
ANK2_HUMAN
|
||||||
| θ value | 0.21417 (rank : 9) | NC score | 0.019369 (rank : 29) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
LSP1_MOUSE
|
||||||
| θ value | 0.21417 (rank : 10) | NC score | 0.066246 (rank : 4) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P19973, P97339, Q04950, Q62022, Q62023, Q62024, Q8CD28, Q99L65 | Gene names | Lsp1, Pp52, S37, Wp34 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphocyte-specific protein 1 (Protein pp52) (52 kDa phosphoprotein) (Lymphocyte-specific antigen WP34) (S37 protein). | |||||
|
CEND1_MOUSE
|
||||||
| θ value | 0.279714 (rank : 11) | NC score | 0.025849 (rank : 23) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BZ05, Q80TX2, Q8C3T2, Q8VEL6 | Gene names | Centd1, Kiaa0580 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centaurin-delta 1 (Cnt-d1). | |||||
|
MBD4_HUMAN
|
||||||
| θ value | 0.47712 (rank : 12) | NC score | 0.050557 (rank : 6) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O95243, Q7Z4T3, Q96F09 | Gene names | MBD4, MED1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 4 (EC 3.2.2.-) (Methyl-CpG-binding protein MBD4) (Methyl-CpG-binding endonuclease 1) (Mismatch-specific DNA N-glycosylase). | |||||
|
TTF2_MOUSE
|
||||||
| θ value | 0.47712 (rank : 13) | NC score | 0.033355 (rank : 14) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q5NC05, Q5M924 | Gene names | Ttf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription termination factor 2 (EC 3.6.1.-) (RNA polymerase II termination factor) (Transcription release factor 2). | |||||
|
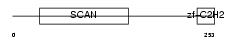
ZN435_HUMAN
|
||||||
| θ value | 0.47712 (rank : 14) | NC score | 0.004594 (rank : 51) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 774 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H4T2, Q9H6K2 | Gene names | ZNF435, ZSCAN16 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger protein 435 (Zinc finger and SCAN domain-containing protein 16). | |||||
|
CSPG2_HUMAN
|
||||||
| θ value | 0.62314 (rank : 15) | NC score | 0.016777 (rank : 36) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P13611, P20754, Q13010, Q13189, Q15123, Q9UCL9, Q9UNW5 | Gene names | CSPG2 | |||
|
Domain Architecture |

|
|||||
| Description | Versican core protein precursor (Large fibroblast proteoglycan) (Chondroitin sulfate proteoglycan core protein 2) (PG-M) (Glial hyaluronate-binding protein) (GHAP). | |||||
|
LRCH2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 16) | NC score | 0.019009 (rank : 30) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q3UMG5, Q7TNP9, Q80TC9 | Gene names | Lrch2, Kiaa1495 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 2. | |||||
|
SMC4_MOUSE
|
||||||
| θ value | 0.62314 (rank : 17) | NC score | 0.028107 (rank : 20) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 830 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8CG47, Q8BTS7, Q8BTY9, Q99K21 | Gene names | Smc4l1, Capc, Smc4 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosomes 4-like 1 protein (Chromosome- associated polypeptide C) (XCAP-C homolog). | |||||
|
KIF4A_MOUSE
|
||||||
| θ value | 1.06291 (rank : 18) | NC score | 0.015574 (rank : 39) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1033 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P33174 | Gene names | Kif4a, Kif4, Kns4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromosome-associated kinesin KIF4A (Chromokinesin). | |||||
|
AKA12_HUMAN
|
||||||
| θ value | 1.38821 (rank : 19) | NC score | 0.046003 (rank : 8) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q02952, O00310, O00498, Q99970 | Gene names | AKAP12, AKAP250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 12 (A-kinase anchor protein 250 kDa) (AKAP 250) (Myasthenia gravis autoantigen gravin). | |||||
|
IMPG1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 20) | NC score | 0.032976 (rank : 15) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q17R60, O43686, O95094, Q9BWZ1 | Gene names | IMPG1, IPM150, SPACR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interphotoreceptor matrix proteoglycan 1 precursor (Interphotoreceptor matrix proteoglycan of 150 kDa) (IPM-150) (Sialoprotein associated with cones and rods). | |||||
|
KCNH7_MOUSE
|
||||||
| θ value | 1.81305 (rank : 21) | NC score | 0.013852 (rank : 41) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9ER47 | Gene names | Kcnh7, Erg3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 7 (Voltage-gated potassium channel subunit Kv11.3) (Ether-a-go-go-related gene potassium channel 3) (Ether-a-go-go-related protein 3) (Eag-related protein 3). | |||||
|
NAR4_HUMAN
|
||||||
| θ value | 1.81305 (rank : 22) | NC score | 0.032406 (rank : 16) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q93070, Q9BZ50, Q9BZ51, Q9HB06 | Gene names | ART4, DO, DOK1 | |||
|
Domain Architecture |

|
|||||
| Description | Ecto-ADP-ribosyltransferase 4 precursor (EC 2.4.2.31) (NAD(P)(+)-- arginine ADP-ribosyltransferase 4) (Mono(ADP-ribosyl)transferase 4) (Dombrock blood group carrier molecule) (CD297 antigen). | |||||
|
RFIP1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 23) | NC score | 0.028854 (rank : 19) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6WKZ4, Q6AZK4, Q6WKZ2, Q6WKZ6, Q86YV4, Q8TDL1, Q9H642 | Gene names | RAB11FIP1, RCP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab11 family-interacting protein 1 (Rab11-FIP1) (Rab-coupling protein). | |||||
|
ANR26_HUMAN
|
||||||
| θ value | 2.36792 (rank : 24) | NC score | 0.018220 (rank : 32) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1614 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9UPS8, Q2TAZ3, Q6ZR14, Q9H1Q1, Q9NSK9, Q9NTD5, Q9NW69 | Gene names | ANKRD26, KIAA1074 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 26. | |||||
|
ATN1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 25) | NC score | 0.024826 (rank : 24) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
RB6I2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 26) | NC score | 0.017871 (rank : 34) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1584 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8IUD2, Q6NVK2, Q8IUD3, Q8IUD4, Q8IUD5, Q8NAS1, Q9NXN5, Q9UIK7, Q9UPS1 | Gene names | ERC1, ELKS, KIAA1081, RAB6IP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ELKS/RAB6-interacting/CAST family member 1 (RAB6-interacting protein 2) (ERC protein 1). | |||||
|
DSPP_HUMAN
|
||||||
| θ value | 3.0926 (rank : 27) | NC score | 0.054099 (rank : 5) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
GGNB1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 28) | NC score | 0.035619 (rank : 13) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6K1E7, Q8C5T9 | Gene names | Ggnbp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin-binding protein 1. | |||||
|
MYH14_HUMAN
|
||||||
| θ value | 3.0926 (rank : 29) | NC score | 0.010691 (rank : 46) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 995 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q7Z406, Q5CZ75, Q6XYE4, Q76B62, Q8WV23, Q96I22, Q9BT27, Q9BW35, Q9H882 | Gene names | MYH14, KIAA2034 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-14 (Myosin heavy chain, nonmuscle IIc) (Nonmuscle myosin heavy chain IIc) (NMHC II-C). | |||||
|
NELFD_HUMAN
|
||||||
| θ value | 3.0926 (rank : 30) | NC score | 0.040409 (rank : 11) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8IXH7, Q9BYL2, Q9H405, Q9H888, Q9H8T3, Q9NVX5, Q9P029, Q9UGN1, Q9UGN2, Q9UGN3 | Gene names | TH1L, NELFD, TH1 | |||
|
Domain Architecture |
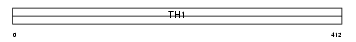
|
|||||
| Description | Negative elongation factor C/D (NELF-C/D) (TH1-like protein). | |||||
|
NELFD_MOUSE
|
||||||
| θ value | 3.0926 (rank : 31) | NC score | 0.040391 (rank : 12) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q922L6, Q9QXC8, Q9QXC9, Q9QXD0 | Gene names | Th1l, Nelfd, Th1 | |||
|
Domain Architecture |

|
|||||
| Description | Negative elongation factor D (NELF-D) (TH1-like protein). | |||||
|
RFC1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 32) | NC score | 0.027511 (rank : 21) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P35601 | Gene names | Rfc1, Ibf-1, Recc1 | |||
|
Domain Architecture |

|
|||||
| Description | Replication factor C subunit 1 (Replication factor C large subunit) (RF-C 140 kDa subunit) (Activator 1 140 kDa subunit) (Activator 1 large subunit) (A1 140 kDa subunit) (A1-P145) (Differentiation- specific element-binding protein) (ISRE-binding protein). | |||||
|
VCAM1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 33) | NC score | 0.010963 (rank : 45) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P19320, Q6NUP8 | Gene names | VCAM1, L1CAM | |||
|
Domain Architecture |

|
|||||
| Description | Vascular cell adhesion protein 1 precursor (V-CAM 1) (CD106 antigen) (INCAM-100). | |||||
|
NRK_HUMAN
|
||||||
| θ value | 4.03905 (rank : 34) | NC score | 0.002513 (rank : 53) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7Z2Y5, Q32ND6, Q5H9K2, Q6ZMP2 | Gene names | NRK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nik-related protein kinase (EC 2.7.11.1). | |||||
|
IWS1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 35) | NC score | 0.024336 (rank : 25) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
LZTS1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 36) | NC score | 0.023277 (rank : 26) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 532 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y250, Q9Y5V7, Q9Y5V8, Q9Y5V9, Q9Y5W0, Q9Y5W1, Q9Y5W2 | Gene names | LZTS1, FEZ1 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine zipper putative tumor suppressor 1 (F37/esophageal cancer- related gene-coding leucine-zipper motif) (Fez1). | |||||
|
MINT_MOUSE
|
||||||
| θ value | 5.27518 (rank : 37) | NC score | 0.017929 (rank : 33) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
MRP1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 38) | NC score | 0.006222 (rank : 50) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P33527, O14819, O43333, P78419, Q59GI9, Q9UQ97, Q9UQ99, Q9UQA0 | Gene names | ABCC1, MRP, MRP1 | |||
|
Domain Architecture |

|
|||||
| Description | Multidrug resistance-associated protein 1 (ATP-binding cassette sub- family C member 1) (Leukotriene C(4) transporter) (LTC4 transporter). | |||||
|
MYCPP_HUMAN
|
||||||
| θ value | 5.27518 (rank : 39) | NC score | 0.016451 (rank : 37) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7Z401, Q14655, Q86T77, Q8IVX2, Q8NB93 | Gene names | MYCPBP, IRLB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | C-myc promoter-binding protein. | |||||
|
STOX1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 40) | NC score | 0.027477 (rank : 22) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6ZVD7, Q4F8Q6, Q5I946, Q5I947, Q5I948, Q5VX38, Q5VX39, Q6ZRY3, Q96LR3, Q96LS0 | Gene names | STOX1, C10orf24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Storkhead-box protein 1 (Winged helix domain-containing protein). | |||||
|
VDP_HUMAN
|
||||||
| θ value | 5.27518 (rank : 41) | NC score | 0.021091 (rank : 27) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O60763 | Gene names | VDP | |||
|
Domain Architecture |

|
|||||
| Description | General vesicular transport factor p115 (Transcytosis-associated protein) (TAP) (Vesicle docking protein). | |||||
|
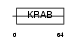
ZN224_HUMAN
|
||||||
| θ value | 5.27518 (rank : 42) | NC score | 0.000533 (rank : 54) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 808 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NZL3, Q86V10, Q8IZC8, Q9UID9, Q9Y2P6 | Gene names | ZNF224, BMZF2, ZNF255 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger protein 224 (Zinc finger protein 255) (Bone marrow zinc finger 2) (BMZF-2). | |||||
|
GP112_HUMAN
|
||||||
| θ value | 6.88961 (rank : 43) | NC score | 0.008233 (rank : 49) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IZF6, Q86SM6 | Gene names | GPR112 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 112. | |||||
|
LZTS1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 44) | NC score | 0.021033 (rank : 28) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 540 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P60853 | Gene names | Lzts1, Fez1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine zipper putative tumor suppressor 1 (F37/Esophageal cancer- related gene-coding leucine-zipper motif) (Fez1). | |||||
|
MCPH1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 45) | NC score | 0.018402 (rank : 31) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7TT79, Q8BNI9, Q8C9I7 | Gene names | Mcph1 | |||
|
Domain Architecture |

|
|||||
| Description | Microcephalin. | |||||
|
MYT1L_HUMAN
|
||||||
| θ value | 6.88961 (rank : 46) | NC score | 0.013800 (rank : 42) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UL68, Q6IQ17, Q9UPP6 | Gene names | MYT1L, KIAA1106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1-like protein (MyT1L protein) (MyT1-L). | |||||
|
PHC3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 47) | NC score | 0.013036 (rank : 43) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NDX5, Q5HYF0, Q6NSG2, Q8NFT7, Q8NFZ1, Q8TBM2, Q9H971, Q9H9I4 | Gene names | PHC3, EDR3, PH3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyhomeotic-like protein 3 (hPH3) (Homolog of polyhomeotic 3) (Early development regulatory protein 3). | |||||
|
TIAM1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 48) | NC score | 0.009742 (rank : 47) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60610 | Gene names | Tiam1, Tiam-1 | |||
|
Domain Architecture |

|
|||||
| Description | T-lymphoma invasion and metastasis-inducing protein 1 (TIAM-1 protein). | |||||
|
AFF4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 49) | NC score | 0.017113 (rank : 35) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9ESC8, Q8C6K4, Q8C6W3, Q8CCH3 | Gene names | Aff4, Alf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4. | |||||
|
DDX11_HUMAN
|
||||||
| θ value | 8.99809 (rank : 50) | NC score | 0.011324 (rank : 44) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96FC9, Q13333, Q86VQ4, Q86W62, Q92498, Q92770, Q92998, Q92999 | Gene names | DDX11, CHL1, CHLR1, KRG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX11 (EC 3.6.1.-) (DEAD/H box protein 11) (CHL1 homolog) (Keratinocyte growth factor-regulated gene 2 protein) (KRG-2). | |||||
|
PPRB_HUMAN
|
||||||
| θ value | 8.99809 (rank : 51) | NC score | 0.016333 (rank : 38) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q15648, O43810, O75447, Q9HD39 | Gene names | PPARBP, ARC205, DRIP205, TRAP220, TRIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor-binding protein (PBP) (PPAR-binding protein) (Thyroid hormone receptor-associated protein complex 220 kDa component) (Trap220) (Thyroid receptor-interacting protein 2) (TRIP-2) (p53 regulatory protein RB18A) (Vitamin D receptor-interacting protein complex component DRIP205) (Activator- recruited cofactor 205 kDa component) (ARC205). | |||||
|
RP1L1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 52) | NC score | 0.015044 (rank : 40) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1072 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8IWN7, Q86SQ1, Q8IWN8, Q8IWN9, Q8IWP0, Q8IWP1, Q8IWP2 | Gene names | RP1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinitis pigmentosa 1-like 1 protein. | |||||
|
SLIK3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 53) | NC score | 0.003946 (rank : 52) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q810B9 | Gene names | Slitrk3 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT and NTRK-like protein 3 precursor. | |||||
|
TEX14_MOUSE
|
||||||
| θ value | 8.99809 (rank : 54) | NC score | 0.008740 (rank : 48) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1333 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q7M6U3, Q3UT36, Q5NC10, Q5NC11, Q8CGK1, Q8CGK2, Q99MV8 | Gene names | Tex14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed protein 14 (Testis-expressed sequence 14). | |||||
|
PPR3A_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q16821, O43476, Q75LN8, Q7KYM8, Q86UI6 | Gene names | PPP1R3A, PP1G | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein phosphatase 1 regulatory subunit 3A (Protein phosphatase 1 glycogen-associated regulatory subunit) (Protein phosphatase type-1 glycogen targeting subunit). | |||||
|
PPR3A_MOUSE
|
||||||
| NC score | 0.963828 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99MR9, Q8BUJ4, Q8BUL0 | Gene names | Ppp1r3a, Pp1g | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein phosphatase 1 regulatory subunit 3A (Protein phosphatase 1 glycogen-associated regulatory subunit) (Protein phosphatase type-1 glycogen targeting subunit) (RGL). | |||||
|
PPR3D_HUMAN
|
||||||
| NC score | 0.621171 (rank : 3) | θ value | 4.02038e-16 (rank : 3) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O95685, Q6DK02 | Gene names | PPP1R3D, PPP1R6 | |||
|
Domain Architecture |

|
|||||
| Description | Protein phosphatase 1 regulatory subunit 3D (Protein phosphatase 1, regulatory subunit 6) (Protein phosphatase 1-binding subunit R6). | |||||
|
LSP1_MOUSE
|
||||||
| NC score | 0.066246 (rank : 4) | θ value | 0.21417 (rank : 10) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P19973, P97339, Q04950, Q62022, Q62023, Q62024, Q8CD28, Q99L65 | Gene names | Lsp1, Pp52, S37, Wp34 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphocyte-specific protein 1 (Protein pp52) (52 kDa phosphoprotein) (Lymphocyte-specific antigen WP34) (S37 protein). | |||||
|
DSPP_HUMAN
|
||||||
| NC score | 0.054099 (rank : 5) | θ value | 3.0926 (rank : 27) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
MBD4_HUMAN
|
||||||
| NC score | 0.050557 (rank : 6) | θ value | 0.47712 (rank : 12) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O95243, Q7Z4T3, Q96F09 | Gene names | MBD4, MED1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 4 (EC 3.2.2.-) (Methyl-CpG-binding protein MBD4) (Methyl-CpG-binding endonuclease 1) (Mismatch-specific DNA N-glycosylase). | |||||
|
RBBP6_MOUSE
|
||||||
| NC score | 0.048605 (rank : 7) | θ value | 0.125558 (rank : 7) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
AKA12_HUMAN
|
||||||
| NC score | 0.046003 (rank : 8) | θ value | 1.38821 (rank : 19) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q02952, O00310, O00498, Q99970 | Gene names | AKAP12, AKAP250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 12 (A-kinase anchor protein 250 kDa) (AKAP 250) (Myasthenia gravis autoantigen gravin). | |||||
|
ATRX_MOUSE
|
||||||
| NC score | 0.042827 (rank : 9) | θ value | 0.163984 (rank : 8) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q61687 | Gene names | Atrx, Hp1bp2, Xnp | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked nuclear protein) (Heterochromatin protein 2) (HP1 alpha-interacting protein) (HP1-BP38 protein). | |||||
|
SYNE2_HUMAN
|
||||||
| NC score | 0.041077 (rank : 10) | θ value | 0.00509761 (rank : 5) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1323 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8WXH0, Q8N1S3, Q8NF49, Q8TER7, Q8WWW3, Q8WWW4, Q8WWW5, Q8WXH1, Q9NU50, Q9UFQ4, Q9Y2L4, Q9Y4R1 | Gene names | SYNE2, KIAA1011, NUA | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-2 (Nuclear envelope spectrin repeat protein 2) (Syne-2) (Synaptic nuclear envelope protein 2) (Nucleus and actin connecting element protein) (Protein NUANCE). | |||||
|
NELFD_HUMAN
|
||||||
| NC score | 0.040409 (rank : 11) | θ value | 3.0926 (rank : 30) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8IXH7, Q9BYL2, Q9H405, Q9H888, Q9H8T3, Q9NVX5, Q9P029, Q9UGN1, Q9UGN2, Q9UGN3 | Gene names | TH1L, NELFD, TH1 | |||
|
Domain Architecture |

|
|||||
| Description | Negative elongation factor C/D (NELF-C/D) (TH1-like protein). | |||||
|
NELFD_MOUSE
|
||||||
| NC score | 0.040391 (rank : 12) | θ value | 3.0926 (rank : 31) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q922L6, Q9QXC8, Q9QXC9, Q9QXD0 | Gene names | Th1l, Nelfd, Th1 | |||
|
Domain Architecture |

|
|||||
| Description | Negative elongation factor D (NELF-D) (TH1-like protein). | |||||
|
GGNB1_MOUSE
|
||||||
| NC score | 0.035619 (rank : 13) | θ value | 3.0926 (rank : 28) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6K1E7, Q8C5T9 | Gene names | Ggnbp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin-binding protein 1. | |||||
|
TTF2_MOUSE
|
||||||
| NC score | 0.033355 (rank : 14) | θ value | 0.47712 (rank : 13) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q5NC05, Q5M924 | Gene names | Ttf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription termination factor 2 (EC 3.6.1.-) (RNA polymerase II termination factor) (Transcription release factor 2). | |||||
|
IMPG1_HUMAN
|
||||||
| NC score | 0.032976 (rank : 15) | θ value | 1.81305 (rank : 20) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q17R60, O43686, O95094, Q9BWZ1 | Gene names | IMPG1, IPM150, SPACR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interphotoreceptor matrix proteoglycan 1 precursor (Interphotoreceptor matrix proteoglycan of 150 kDa) (IPM-150) (Sialoprotein associated with cones and rods). | |||||
|
NAR4_HUMAN
|
||||||
| NC score | 0.032406 (rank : 16) | θ value | 1.81305 (rank : 22) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q93070, Q9BZ50, Q9BZ51, Q9HB06 | Gene names | ART4, DO, DOK1 | |||
|
Domain Architecture |

|
|||||
| Description | Ecto-ADP-ribosyltransferase 4 precursor (EC 2.4.2.31) (NAD(P)(+)-- arginine ADP-ribosyltransferase 4) (Mono(ADP-ribosyl)transferase 4) (Dombrock blood group carrier molecule) (CD297 antigen). | |||||
|
NFM_MOUSE
|
||||||
| NC score | 0.029421 (rank : 17) | θ value | 0.00298849 (rank : 4) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1159 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P08553, Q61961 | Gene names | Nef3, Nefm, Nfm | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M). | |||||
|
LAMA4_HUMAN
|
||||||
| NC score | 0.029123 (rank : 18) | θ value | 0.0563607 (rank : 6) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 708 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q16363, Q14735, Q15335, Q4LE44, Q5SZG8, Q9UE18, Q9UJN9 | Gene names | LAMA4 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-4 chain precursor. | |||||
|
RFIP1_HUMAN
|
||||||
| NC score | 0.028854 (rank : 19) | θ value | 1.81305 (rank : 23) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6WKZ4, Q6AZK4, Q6WKZ2, Q6WKZ6, Q86YV4, Q8TDL1, Q9H642 | Gene names | RAB11FIP1, RCP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab11 family-interacting protein 1 (Rab11-FIP1) (Rab-coupling protein). | |||||
|
SMC4_MOUSE
|
||||||
| NC score | 0.028107 (rank : 20) | θ value | 0.62314 (rank : 17) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 830 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8CG47, Q8BTS7, Q8BTY9, Q99K21 | Gene names | Smc4l1, Capc, Smc4 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosomes 4-like 1 protein (Chromosome- associated polypeptide C) (XCAP-C homolog). | |||||
|
RFC1_MOUSE
|
||||||
| NC score | 0.027511 (rank : 21) | θ value | 3.0926 (rank : 32) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P35601 | Gene names | Rfc1, Ibf-1, Recc1 | |||
|
Domain Architecture |

|
|||||
| Description | Replication factor C subunit 1 (Replication factor C large subunit) (RF-C 140 kDa subunit) (Activator 1 140 kDa subunit) (Activator 1 large subunit) (A1 140 kDa subunit) (A1-P145) (Differentiation- specific element-binding protein) (ISRE-binding protein). | |||||
|
STOX1_HUMAN
|
||||||
| NC score | 0.027477 (rank : 22) | θ value | 5.27518 (rank : 40) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6ZVD7, Q4F8Q6, Q5I946, Q5I947, Q5I948, Q5VX38, Q5VX39, Q6ZRY3, Q96LR3, Q96LS0 | Gene names | STOX1, C10orf24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Storkhead-box protein 1 (Winged helix domain-containing protein). | |||||
|
CEND1_MOUSE
|
||||||
| NC score | 0.025849 (rank : 23) | θ value | 0.279714 (rank : 11) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BZ05, Q80TX2, Q8C3T2, Q8VEL6 | Gene names | Centd1, Kiaa0580 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centaurin-delta 1 (Cnt-d1). | |||||
|
ATN1_HUMAN
|
||||||
| NC score | 0.024826 (rank : 24) | θ value | 2.36792 (rank : 25) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.024336 (rank : 25) | θ value | 5.27518 (rank : 35) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
LZTS1_HUMAN
|
||||||
| NC score | 0.023277 (rank : 26) | θ value | 5.27518 (rank : 36) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 532 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y250, Q9Y5V7, Q9Y5V8, Q9Y5V9, Q9Y5W0, Q9Y5W1, Q9Y5W2 | Gene names | LZTS1, FEZ1 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine zipper putative tumor suppressor 1 (F37/esophageal cancer- related gene-coding leucine-zipper motif) (Fez1). | |||||
|
VDP_HUMAN
|
||||||
| NC score | 0.021091 (rank : 27) | θ value | 5.27518 (rank : 41) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O60763 | Gene names | VDP | |||
|
Domain Architecture |

|
|||||
| Description | General vesicular transport factor p115 (Transcytosis-associated protein) (TAP) (Vesicle docking protein). | |||||
|
LZTS1_MOUSE
|
||||||
| NC score | 0.021033 (rank : 28) | θ value | 6.88961 (rank : 44) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 540 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P60853 | Gene names | Lzts1, Fez1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine zipper putative tumor suppressor 1 (F37/Esophageal cancer- related gene-coding leucine-zipper motif) (Fez1). | |||||
|
ANK2_HUMAN
|
||||||
| NC score | 0.019369 (rank : 29) | θ value | 0.21417 (rank : 9) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
LRCH2_MOUSE
|
||||||
| NC score | 0.019009 (rank : 30) | θ value | 0.62314 (rank : 16) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q3UMG5, Q7TNP9, Q80TC9 | Gene names | Lrch2, Kiaa1495 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 2. | |||||
|
MCPH1_MOUSE
|
||||||
| NC score | 0.018402 (rank : 31) | θ value | 6.88961 (rank : 45) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7TT79, Q8BNI9, Q8C9I7 | Gene names | Mcph1 | |||
|
Domain Architecture |

|
|||||
| Description | Microcephalin. | |||||
|
ANR26_HUMAN
|
||||||
| NC score | 0.018220 (rank : 32) | θ value | 2.36792 (rank : 24) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1614 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9UPS8, Q2TAZ3, Q6ZR14, Q9H1Q1, Q9NSK9, Q9NTD5, Q9NW69 | Gene names | ANKRD26, KIAA1074 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 26. | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.017929 (rank : 33) | θ value | 5.27518 (rank : 37) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
RB6I2_HUMAN
|
||||||
| NC score | 0.017871 (rank : 34) | θ value | 2.36792 (rank : 26) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1584 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8IUD2, Q6NVK2, Q8IUD3, Q8IUD4, Q8IUD5, Q8NAS1, Q9NXN5, Q9UIK7, Q9UPS1 | Gene names | ERC1, ELKS, KIAA1081, RAB6IP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ELKS/RAB6-interacting/CAST family member 1 (RAB6-interacting protein 2) (ERC protein 1). | |||||
|
AFF4_MOUSE
|
||||||
| NC score | 0.017113 (rank : 35) | θ value | 8.99809 (rank : 49) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9ESC8, Q8C6K4, Q8C6W3, Q8CCH3 | Gene names | Aff4, Alf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4. | |||||
|
CSPG2_HUMAN
|
||||||
| NC score | 0.016777 (rank : 36) | θ value | 0.62314 (rank : 15) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P13611, P20754, Q13010, Q13189, Q15123, Q9UCL9, Q9UNW5 | Gene names | CSPG2 | |||
|
Domain Architecture |

|
|||||
| Description | Versican core protein precursor (Large fibroblast proteoglycan) (Chondroitin sulfate proteoglycan core protein 2) (PG-M) (Glial hyaluronate-binding protein) (GHAP). | |||||
|
MYCPP_HUMAN
|
||||||
| NC score | 0.016451 (rank : 37) | θ value | 5.27518 (rank : 39) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7Z401, Q14655, Q86T77, Q8IVX2, Q8NB93 | Gene names | MYCPBP, IRLB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | C-myc promoter-binding protein. | |||||
|
PPRB_HUMAN
|
||||||
| NC score | 0.016333 (rank : 38) | θ value | 8.99809 (rank : 51) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q15648, O43810, O75447, Q9HD39 | Gene names | PPARBP, ARC205, DRIP205, TRAP220, TRIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor-binding protein (PBP) (PPAR-binding protein) (Thyroid hormone receptor-associated protein complex 220 kDa component) (Trap220) (Thyroid receptor-interacting protein 2) (TRIP-2) (p53 regulatory protein RB18A) (Vitamin D receptor-interacting protein complex component DRIP205) (Activator- recruited cofactor 205 kDa component) (ARC205). | |||||
|
KIF4A_MOUSE
|
||||||
| NC score | 0.015574 (rank : 39) | θ value | 1.06291 (rank : 18) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1033 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P33174 | Gene names | Kif4a, Kif4, Kns4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromosome-associated kinesin KIF4A (Chromokinesin). | |||||
|
RP1L1_HUMAN
|
||||||
| NC score | 0.015044 (rank : 40) | θ value | 8.99809 (rank : 52) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1072 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8IWN7, Q86SQ1, Q8IWN8, Q8IWN9, Q8IWP0, Q8IWP1, Q8IWP2 | Gene names | RP1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinitis pigmentosa 1-like 1 protein. | |||||
|
KCNH7_MOUSE
|
||||||
| NC score | 0.013852 (rank : 41) | θ value | 1.81305 (rank : 21) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9ER47 | Gene names | Kcnh7, Erg3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 7 (Voltage-gated potassium channel subunit Kv11.3) (Ether-a-go-go-related gene potassium channel 3) (Ether-a-go-go-related protein 3) (Eag-related protein 3). | |||||
|
MYT1L_HUMAN
|
||||||
| NC score | 0.013800 (rank : 42) | θ value | 6.88961 (rank : 46) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UL68, Q6IQ17, Q9UPP6 | Gene names | MYT1L, KIAA1106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1-like protein (MyT1L protein) (MyT1-L). | |||||
|
PHC3_HUMAN
|
||||||
| NC score | 0.013036 (rank : 43) | θ value | 6.88961 (rank : 47) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NDX5, Q5HYF0, Q6NSG2, Q8NFT7, Q8NFZ1, Q8TBM2, Q9H971, Q9H9I4 | Gene names | PHC3, EDR3, PH3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyhomeotic-like protein 3 (hPH3) (Homolog of polyhomeotic 3) (Early development regulatory protein 3). | |||||
|
DDX11_HUMAN
|
||||||
| NC score | 0.011324 (rank : 44) | θ value | 8.99809 (rank : 50) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96FC9, Q13333, Q86VQ4, Q86W62, Q92498, Q92770, Q92998, Q92999 | Gene names | DDX11, CHL1, CHLR1, KRG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX11 (EC 3.6.1.-) (DEAD/H box protein 11) (CHL1 homolog) (Keratinocyte growth factor-regulated gene 2 protein) (KRG-2). | |||||
|
VCAM1_HUMAN
|
||||||
| NC score | 0.010963 (rank : 45) | θ value | 3.0926 (rank : 33) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P19320, Q6NUP8 | Gene names | VCAM1, L1CAM | |||
|
Domain Architecture |

|
|||||
| Description | Vascular cell adhesion protein 1 precursor (V-CAM 1) (CD106 antigen) (INCAM-100). | |||||
|
MYH14_HUMAN
|
||||||
| NC score | 0.010691 (rank : 46) | θ value | 3.0926 (rank : 29) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 995 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q7Z406, Q5CZ75, Q6XYE4, Q76B62, Q8WV23, Q96I22, Q9BT27, Q9BW35, Q9H882 | Gene names | MYH14, KIAA2034 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-14 (Myosin heavy chain, nonmuscle IIc) (Nonmuscle myosin heavy chain IIc) (NMHC II-C). | |||||
|
TIAM1_MOUSE
|
||||||
| NC score | 0.009742 (rank : 47) | θ value | 6.88961 (rank : 48) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60610 | Gene names | Tiam1, Tiam-1 | |||
|
Domain Architecture |

|
|||||
| Description | T-lymphoma invasion and metastasis-inducing protein 1 (TIAM-1 protein). | |||||
|
TEX14_MOUSE
|
||||||
| NC score | 0.008740 (rank : 48) | θ value | 8.99809 (rank : 54) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1333 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q7M6U3, Q3UT36, Q5NC10, Q5NC11, Q8CGK1, Q8CGK2, Q99MV8 | Gene names | Tex14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed protein 14 (Testis-expressed sequence 14). | |||||
|
GP112_HUMAN
|
||||||
| NC score | 0.008233 (rank : 49) | θ value | 6.88961 (rank : 43) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IZF6, Q86SM6 | Gene names | GPR112 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 112. | |||||
|
MRP1_HUMAN
|
||||||
| NC score | 0.006222 (rank : 50) | θ value | 5.27518 (rank : 38) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P33527, O14819, O43333, P78419, Q59GI9, Q9UQ97, Q9UQ99, Q9UQA0 | Gene names | ABCC1, MRP, MRP1 | |||
|
Domain Architecture |

|
|||||
| Description | Multidrug resistance-associated protein 1 (ATP-binding cassette sub- family C member 1) (Leukotriene C(4) transporter) (LTC4 transporter). | |||||
|
ZN435_HUMAN
|
||||||
| NC score | 0.004594 (rank : 51) | θ value | 0.47712 (rank : 14) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 774 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H4T2, Q9H6K2 | Gene names | ZNF435, ZSCAN16 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 435 (Zinc finger and SCAN domain-containing protein 16). | |||||
|
SLIK3_MOUSE
|
||||||
| NC score | 0.003946 (rank : 52) | θ value | 8.99809 (rank : 53) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q810B9 | Gene names | Slitrk3 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT and NTRK-like protein 3 precursor. | |||||
|
NRK_HUMAN
|
||||||
| NC score | 0.002513 (rank : 53) | θ value | 4.03905 (rank : 34) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7Z2Y5, Q32ND6, Q5H9K2, Q6ZMP2 | Gene names | NRK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nik-related protein kinase (EC 2.7.11.1). | |||||
|
ZN224_HUMAN
|
||||||
| NC score | 0.000533 (rank : 54) | θ value | 5.27518 (rank : 42) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 808 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NZL3, Q86V10, Q8IZC8, Q9UID9, Q9Y2P6 | Gene names | ZNF224, BMZF2, ZNF255 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 224 (Zinc finger protein 255) (Bone marrow zinc finger 2) (BMZF-2). | |||||