Please be patient as the page loads
|
PARP2_MOUSE
|
||||||
| SwissProt Accessions | O88554, Q99N29 | Gene names | Parp2, Adprt2, Adprtl2, Aspartl2 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 2 (EC 2.4.2.30) (PARP-2) (NAD(+) ADP- ribosyltransferase 2) (Poly[ADP-ribose] synthetase 2) (pADPRT-2) (mPARP-2). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
PARP2_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.991472 (rank : 2) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9UGN5, Q8TEU4, Q9NUV2, Q9UMR4, Q9Y6C8 | Gene names | PARP2, ADPRT2, ADPRTL2 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 2 (EC 2.4.2.30) (PARP-2) (NAD(+) ADP- ribosyltransferase 2) (Poly[ADP-ribose] synthetase 2) (pADPRT-2) (hPARP-2). | |||||
|
PARP2_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O88554, Q99N29 | Gene names | Parp2, Adprt2, Adprtl2, Aspartl2 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 2 (EC 2.4.2.30) (PARP-2) (NAD(+) ADP- ribosyltransferase 2) (Poly[ADP-ribose] synthetase 2) (pADPRT-2) (mPARP-2). | |||||
|
PARP1_HUMAN
|
||||||
| θ value | 9.32813e-106 (rank : 3) | NC score | 0.917143 (rank : 4) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P09874, Q8IUZ9 | Gene names | PARP1, ADPRT, PPOL | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 1 (EC 2.4.2.30) (PARP-1) (ADPRT) (NAD(+) ADP-ribosyltransferase 1) (Poly[ADP-ribose] synthetase 1). | |||||
|
PARP1_MOUSE
|
||||||
| θ value | 6.0463e-105 (rank : 4) | NC score | 0.907493 (rank : 5) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P11103, Q9JLX4, Q9QVQ3 | Gene names | Parp1, Adprp, Adprt, Adprt1 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 1 (EC 2.4.2.30) (PARP-1) (ADPRT) (NAD(+) ADP-ribosyltransferase 1) (Poly[ADP-ribose] synthetase 1) (msPARP). | |||||
|
PARP3_HUMAN
|
||||||
| θ value | 1.60597e-73 (rank : 5) | NC score | 0.923806 (rank : 3) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Y6F1, Q8NER9, Q96CG2, Q9UG81 | Gene names | PARP3, ADPRT3, ADPRTL3 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 3 (EC 2.4.2.30) (PARP-3) (NAD(+) ADP- ribosyltransferase 3) (Poly[ADP-ribose] synthetase 3) (pADPRT-3) (hPARP-3) (IRT1). | |||||
|
PARP4_HUMAN
|
||||||
| θ value | 5.07402e-19 (rank : 6) | NC score | 0.551828 (rank : 6) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9UKK3, O75903, Q14682, Q9H1M6 | Gene names | PARP4, ADPRTL1, KIAA0177, PARPL | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 4 (EC 2.4.2.30) (PARP-4) (Vault poly(ADP- ribose) polymerase) (VPARP) (193 kDa vault protein) (PARP- related/IalphaI-related H5/proline-rich) (PH5P). | |||||
|
PARP8_HUMAN
|
||||||
| θ value | 0.00228821 (rank : 7) | NC score | 0.188001 (rank : 7) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8N3A8, Q3KRB7, Q6DHZ1, Q9H754 | Gene names | PARP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 8 (EC 2.4.2.30) (PARP-8). | |||||
|
PARP8_MOUSE
|
||||||
| θ value | 0.00228821 (rank : 8) | NC score | 0.187701 (rank : 8) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q3UD82, Q3UDE4, Q3UDU5, Q8VCB5, Q9CYY3 | Gene names | Parp8, D13Ertd275e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 8 (EC 2.4.2.30) (PARP-8). | |||||
|
TNKS1_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 9) | NC score | 0.068156 (rank : 15) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 477 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O95271, O95272 | Gene names | TNKS, PARP5A, PARPL, TIN1, TINF1, TNKS1 | |||
|
Domain Architecture |

|
|||||
| Description | Tankyrase-1 (EC 2.4.2.30) (TANK1) (Tankyrase I) (TNKS-1) (TRF1- interacting ankyrin-related ADP-ribose polymerase). | |||||
|
PARP6_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 10) | NC score | 0.181736 (rank : 9) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q2NL67, Q9H7C5, Q9H9X6, Q9HAF3, Q9NPS6, Q9UFG4 | Gene names | PARP6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 6 (EC 2.4.2.30) (PARP-6). | |||||
|
PARP6_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 11) | NC score | 0.181633 (rank : 10) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6P6P7, Q8BVW1 | Gene names | Parp6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 6 (EC 2.4.2.30) (PARP-6). | |||||
|
TNKS2_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 12) | NC score | 0.063564 (rank : 16) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9H2K2, Q9H8F2, Q9HAS4 | Gene names | TNKS2, PARP5B, TANK2, TNKL | |||
|
Domain Architecture |

|
|||||
| Description | Tankyrase-2 (EC 2.4.2.30) (TANK2) (Tankyrase II) (TNKS-2) (TRF1- interacting ankyrin-related ADP-ribose polymerase 2) (Tankyrase-like protein) (Tankyrase-related protein). | |||||
|
TRAIP_HUMAN
|
||||||
| θ value | 0.125558 (rank : 13) | NC score | 0.042750 (rank : 19) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 780 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9BWF2, O00467 | Gene names | TRAIP, TRIP | |||
|
Domain Architecture |
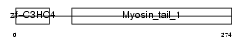
|
|||||
| Description | TRAF-interacting protein. | |||||
|
GCC2_HUMAN
|
||||||
| θ value | 0.21417 (rank : 14) | NC score | 0.027703 (rank : 30) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1558 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8IWJ2, O15045, Q8TDH3, Q9H2G8 | Gene names | GCC2, KIAA0336 | |||
|
Domain Architecture |

|
|||||
| Description | GRIP and coiled-coil domain-containing protein 2 (Golgi coiled coil protein GCC185) (CTCL tumor antigen se1-1) (CLL-associated antigen KW- 11) (NY-REN-53 antigen). | |||||
|
RBP24_HUMAN
|
||||||
| θ value | 0.21417 (rank : 15) | NC score | 0.027735 (rank : 29) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q4ZG46 | Gene names | RANBP2L4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 2-like 4 (RanBP2L4). | |||||
|
PARPT_HUMAN
|
||||||
| θ value | 0.365318 (rank : 16) | NC score | 0.073872 (rank : 12) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q7Z3E1, Q68CY9, Q86VP4, Q9Y4P7 | Gene names | TIPARP, PARP7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TCDD-inducible poly [ADP-ribose] polymerase (EC 2.4.2.30) (Poly [ADP- ribose] polymerase 7) (PARP-7). | |||||
|
PARPT_MOUSE
|
||||||
| θ value | 0.365318 (rank : 17) | NC score | 0.090260 (rank : 11) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8C1B2, Q3UD50, Q8C032 | Gene names | Tiparp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TCDD-inducible poly [ADP-ribose] polymerase (EC 2.4.2.30). | |||||
|
ABTAP_HUMAN
|
||||||
| θ value | 0.47712 (rank : 18) | NC score | 0.042049 (rank : 20) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9H501, Q86X92, Q9H9Q5, Q9HA35, Q9NX93, Q9P1S6 | Gene names | ABTAP, C20orf6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ABT1-associated protein. | |||||
|
AQR_MOUSE
|
||||||
| θ value | 0.813845 (rank : 19) | NC score | 0.026820 (rank : 31) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CFQ3, P97871, Q3U9N1, Q3ULE8, Q80TX8 | Gene names | Aqr, Kiaa0560 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intron-binding protein aquarius. | |||||
|
LIPB2_MOUSE
|
||||||
| θ value | 0.813845 (rank : 20) | NC score | 0.029643 (rank : 27) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 570 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O35711, Q7TMG4, Q8CBS6, Q99KX6 | Gene names | Ppfibp2, Cclp1 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-beta-2 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein-binding protein 2) (PTPRF-interacting protein-binding protein 2) (Coiled-coil-like protein 1). | |||||
|
TPM4_HUMAN
|
||||||
| θ value | 0.813845 (rank : 21) | NC score | 0.023561 (rank : 33) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 525 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P67936, P07226, Q15659, Q9BU85, Q9H8Q3 | Gene names | TPM4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tropomyosin alpha-4 chain (Tropomyosin-4) (TM30p1). | |||||
|
TRAIP_MOUSE
|
||||||
| θ value | 1.06291 (rank : 22) | NC score | 0.034767 (rank : 25) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 745 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8VIG6, O08854, Q922M8, Q9CPP4 | Gene names | Traip, Trip | |||
|
Domain Architecture |
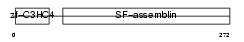
|
|||||
| Description | TRAF-interacting protein. | |||||
|
DNM3B_MOUSE
|
||||||
| θ value | 1.38821 (rank : 23) | NC score | 0.019272 (rank : 41) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O88509, O88510, O88511 | Gene names | Dnmt3b | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 3B (EC 2.1.1.37) (Dnmt3b) (DNA methyltransferase MmuIIIB) (DNA MTase MmuIIIB) (M.MmuIIIB). | |||||
|
LIPB2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 24) | NC score | 0.029070 (rank : 28) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 599 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8ND30, O75337, Q8WW26 | Gene names | PPFIBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-beta-2 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein-binding protein 2) (PTPRF-interacting protein-binding protein 2). | |||||
|
STML2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 25) | NC score | 0.039179 (rank : 24) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UJZ1, O60376, Q53G29, Q96FY2, Q9P042 | Gene names | STOML2, SLP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stomatin-like protein 2 (SLP-2) (EPB72-like 2). | |||||
|
STML2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 26) | NC score | 0.039187 (rank : 23) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99JB2, Q9DCG8 | Gene names | Stoml2, Slp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stomatin-like protein 2 (SLP-2). | |||||
|
UACA_HUMAN
|
||||||
| θ value | 1.38821 (rank : 27) | NC score | 0.020178 (rank : 40) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1622 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9BZF9, Q8N3B8, Q9HCL1, Q9NWC6 | Gene names | UACA, KIAA1561 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uveal autoantigen with coiled-coil domains and ankyrin repeats protein. | |||||
|
GA2L3_HUMAN
|
||||||
| θ value | 1.81305 (rank : 28) | NC score | 0.020695 (rank : 39) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q86XJ1 | Gene names | GAS2L3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GAS2-like protein 3 (Growth arrest-specific 2-like 3). | |||||
|
INVO_HUMAN
|
||||||
| θ value | 1.81305 (rank : 29) | NC score | 0.021362 (rank : 38) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 636 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P07476 | Gene names | IVL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Involucrin. | |||||
|
S39A6_MOUSE
|
||||||
| θ value | 1.81305 (rank : 30) | NC score | 0.021797 (rank : 37) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8C145, Q7TPP9, Q7TQE0, Q8R518 | Gene names | Slc39a6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc transporter SLC39A6 precursor (Solute carrier family 39 member 6) (Endoplasmic reticulum membrane-linked protein) (Ermelin). | |||||
|
SMC4_MOUSE
|
||||||
| θ value | 1.81305 (rank : 31) | NC score | 0.024319 (rank : 32) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 830 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8CG47, Q8BTS7, Q8BTY9, Q99K21 | Gene names | Smc4l1, Capc, Smc4 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosomes 4-like 1 protein (Chromosome- associated polypeptide C) (XCAP-C homolog). | |||||
|
FLOT2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 32) | NC score | 0.041228 (rank : 21) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14254 | Gene names | FLOT2, ESA1, M17S1 | |||
|
Domain Architecture |
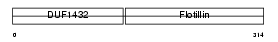
|
|||||
| Description | Flotillin-2 (Epidermal surface antigen) (ESA). | |||||
|
FLOT2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 33) | NC score | 0.041094 (rank : 22) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q60634, Q6NS75 | Gene names | Flot2, Esa1, M17s1 | |||
|
Domain Architecture |
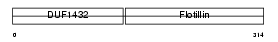
|
|||||
| Description | Flotillin-2 (Epidermal surface antigen) (ESA). | |||||
|
GUC2F_HUMAN
|
||||||
| θ value | 2.36792 (rank : 34) | NC score | 0.005725 (rank : 57) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 740 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P51841, Q9UJF1 | Gene names | GUCY2F, GUC2F, RETGC2 | |||
|
Domain Architecture |

|
|||||
| Description | Retinal guanylyl cyclase 2 precursor (EC 4.6.1.2) (Guanylate cyclase 2F, retinal) (RETGC-2) (Rod outer segment membrane guanylate cyclase 2) (ROS-GC2) (Guanylate cyclase F) (GC-F). | |||||
|
BRD8_HUMAN
|
||||||
| θ value | 3.0926 (rank : 35) | NC score | 0.015358 (rank : 44) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H0E9, O43178, Q15355, Q59GN0, Q969M9 | Gene names | BRD8, SMAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 8 (p120) (Skeletal muscle abundant protein) (Thyroid hormone receptor coactivating protein 120kDa) (TrCP120). | |||||
|
TPM4_MOUSE
|
||||||
| θ value | 3.0926 (rank : 36) | NC score | 0.022358 (rank : 35) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 703 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6IRU2 | Gene names | Tpm4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tropomyosin alpha-4 chain (Tropomyosin-4). | |||||
|
ABTAP_MOUSE
|
||||||
| θ value | 4.03905 (rank : 37) | NC score | 0.033812 (rank : 26) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q3V1V3 | Gene names | Abtap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ABT1-associated protein. | |||||
|
MYH8_HUMAN
|
||||||
| θ value | 4.03905 (rank : 38) | NC score | 0.012272 (rank : 53) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1610 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P13535, Q14910 | Gene names | MYH8 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-8 (Myosin heavy chain, skeletal muscle, perinatal) (MyHC- perinatal). | |||||
|
SMC4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 39) | NC score | 0.022453 (rank : 34) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 937 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9NTJ3, O95752, Q8NDL4, Q9UNT9 | Gene names | SMC4L1, CAPC, SMC4 | |||
|
Domain Architecture |
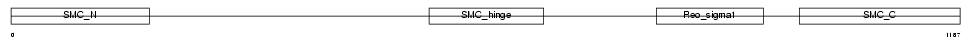
|
|||||
| Description | Structural maintenance of chromosomes 4-like 1 protein (Chromosome- associated polypeptide C) (hCAP-C) (XCAP-C homolog). | |||||
|
CN135_HUMAN
|
||||||
| θ value | 5.27518 (rank : 40) | NC score | 0.022126 (rank : 36) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q63HM2, Q9BQG8, Q9H9F2 | Gene names | C14orf135, FBP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf135 precursor (Hepatitis C virus F protein-binding protein 2) (HCV F protein-binding protein 2). | |||||
|
GUC2D_HUMAN
|
||||||
| θ value | 5.27518 (rank : 41) | NC score | 0.006119 (rank : 56) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 715 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q02846, Q6LEA7 | Gene names | GUCY2D, CORD6, GUC1A4, GUC2D, RETGC, RETGC1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinal guanylyl cyclase 1 precursor (EC 4.6.1.2) (Guanylate cyclase 2D, retinal) (RETGC-1) (Rod outer segment membrane guanylate cyclase) (ROS-GC). | |||||
|
MYO5C_HUMAN
|
||||||
| θ value | 5.27518 (rank : 42) | NC score | 0.012884 (rank : 51) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1206 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9NQX4 | Gene names | MYO5C | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5C (Myosin Vc). | |||||
|
GOGA2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 43) | NC score | 0.014171 (rank : 48) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 944 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q921M4 | Gene names | Golga2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgin subfamily A member 2 (Cis-Golgi matrix protein GM130). | |||||
|
OLR1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 44) | NC score | 0.012400 (rank : 52) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 596 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9EQ09 | Gene names | Olr1, Lox1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oxidized low-density lipoprotein receptor 1 (Ox-LDL receptor 1) (Lectin-type oxidized LDL receptor 1) (Lectin-like oxidized LDL receptor 1) (Lectin-like oxLDL receptor 1) (LOX-1) [Contains: Oxidized low-density lipoprotein receptor 1, soluble form]. | |||||
|
RENT2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 45) | NC score | 0.014372 (rank : 47) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 618 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9HAU5, Q8N8U1, Q9H1J2, Q9NWL1, Q9P2D9, Q9Y4M9 | Gene names | UPF2, KIAA1408, RENT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of nonsense transcripts 2 (Nonsense mRNA reducing factor 2) (Up-frameshift suppressor 2 homolog) (hUpf2). | |||||
|
USBP1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 46) | NC score | 0.015978 (rank : 42) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8N6Y0, Q8NBX7, Q96KH3, Q9BYI8 | Gene names | USHBP1, AIEBP, MCC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USH1C-binding protein 1 (Usher syndrome type-1C protein-binding protein 1) (MCC-2) (AIE-75 binding protein). | |||||
|
ASPH_HUMAN
|
||||||
| θ value | 8.99809 (rank : 47) | NC score | 0.013862 (rank : 49) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q12797 | Gene names | ASPH | |||
|
Domain Architecture |

|
|||||
| Description | Aspartyl/asparaginyl beta-hydroxylase (EC 1.14.11.16) (Aspartate beta- hydroxylase) (ASP beta-hydroxylase) (Peptide-aspartate beta- dioxygenase). | |||||
|
BRAP_MOUSE
|
||||||
| θ value | 8.99809 (rank : 48) | NC score | 0.015289 (rank : 45) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q99MP8, Q8CC00, Q8CHX1, Q99MP7, Q9CXX8 | Gene names | Brap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BRCA1-associated protein (EC 6.3.2.-) (BRAP2) (Impedes mitogenic signal propagation) (IMP). | |||||
|
JAD1A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 49) | NC score | 0.015129 (rank : 46) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P29375 | Gene names | JARID1A, RBBP2, RBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1A (Retinoblastoma-binding protein 2) (RBBP-2). | |||||
|
KTN1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 50) | NC score | 0.015829 (rank : 43) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1197 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q86UP2, Q13999, Q14707, Q15387, Q86W57 | Gene names | KTN1, CG1, KIAA0004 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinectin (Kinesin receptor) (CG-1 antigen). | |||||
|
STX1C_HUMAN
|
||||||
| θ value | 8.99809 (rank : 51) | NC score | 0.010969 (rank : 54) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 270 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P61266 | Gene names | STX1B2 | |||
|
Domain Architecture |

|
|||||
| Description | Syntaxin-1B2. | |||||
|
STX1C_MOUSE
|
||||||
| θ value | 8.99809 (rank : 52) | NC score | 0.010969 (rank : 55) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 270 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P61264, P32853 | Gene names | Stx1b2, Stx1b | |||
|
Domain Architecture |

|
|||||
| Description | Syntaxin-1B2 (Syntaxin 1B). | |||||
|
TPR_HUMAN
|
||||||
| θ value | 8.99809 (rank : 53) | NC score | 0.013389 (rank : 50) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
DNL3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 54) | NC score | 0.071049 (rank : 13) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P49916, Q16714 | Gene names | LIG3 | |||
|
Domain Architecture |
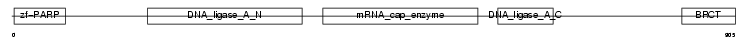
|
|||||
| Description | DNA ligase 3 (EC 6.5.1.1) (DNA ligase III) (Polydeoxyribonucleotide synthase [ATP] 3). | |||||
|
DNL3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 55) | NC score | 0.069153 (rank : 14) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P97386, P97385 | Gene names | Lig3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA ligase 3 (EC 6.5.1.1) (DNA ligase III) (Polydeoxyribonucleotide synthase [ATP] 3). | |||||
|
LHR2A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 56) | NC score | 0.050838 (rank : 18) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 1 | |
| SwissProt Accessions | O00534, Q6UN19, Q6UN20, Q9BVF8 | Gene names | LOH11CR2A, BCSC1 | |||
|
Domain Architecture |

|
|||||
| Description | Loss of heterozygosity 11 chromosomal region 2 gene A protein (Breast cancer suppressor candidate 1) (BCSC-1). | |||||
|
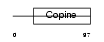
LHR2A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 57) | NC score | 0.054108 (rank : 17) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 1 | |
| SwissProt Accessions | Q99KC8, Q3TTU2, Q6UN22, Q8BHA8, Q9CTV9 | Gene names | Loh11cr2a | |||
|
Domain Architecture |
|
|||||
| Description | Loss of heterozygosity 11 chromosomal region 2 gene A protein homolog. | |||||
|
PARP2_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O88554, Q99N29 | Gene names | Parp2, Adprt2, Adprtl2, Aspartl2 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 2 (EC 2.4.2.30) (PARP-2) (NAD(+) ADP- ribosyltransferase 2) (Poly[ADP-ribose] synthetase 2) (pADPRT-2) (mPARP-2). | |||||
|
PARP2_HUMAN
|
||||||
| NC score | 0.991472 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9UGN5, Q8TEU4, Q9NUV2, Q9UMR4, Q9Y6C8 | Gene names | PARP2, ADPRT2, ADPRTL2 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 2 (EC 2.4.2.30) (PARP-2) (NAD(+) ADP- ribosyltransferase 2) (Poly[ADP-ribose] synthetase 2) (pADPRT-2) (hPARP-2). | |||||
|
PARP3_HUMAN
|
||||||
| NC score | 0.923806 (rank : 3) | θ value | 1.60597e-73 (rank : 5) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Y6F1, Q8NER9, Q96CG2, Q9UG81 | Gene names | PARP3, ADPRT3, ADPRTL3 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 3 (EC 2.4.2.30) (PARP-3) (NAD(+) ADP- ribosyltransferase 3) (Poly[ADP-ribose] synthetase 3) (pADPRT-3) (hPARP-3) (IRT1). | |||||
|
PARP1_HUMAN
|
||||||
| NC score | 0.917143 (rank : 4) | θ value | 9.32813e-106 (rank : 3) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P09874, Q8IUZ9 | Gene names | PARP1, ADPRT, PPOL | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 1 (EC 2.4.2.30) (PARP-1) (ADPRT) (NAD(+) ADP-ribosyltransferase 1) (Poly[ADP-ribose] synthetase 1). | |||||
|
PARP1_MOUSE
|
||||||
| NC score | 0.907493 (rank : 5) | θ value | 6.0463e-105 (rank : 4) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P11103, Q9JLX4, Q9QVQ3 | Gene names | Parp1, Adprp, Adprt, Adprt1 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 1 (EC 2.4.2.30) (PARP-1) (ADPRT) (NAD(+) ADP-ribosyltransferase 1) (Poly[ADP-ribose] synthetase 1) (msPARP). | |||||
|
PARP4_HUMAN
|
||||||
| NC score | 0.551828 (rank : 6) | θ value | 5.07402e-19 (rank : 6) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9UKK3, O75903, Q14682, Q9H1M6 | Gene names | PARP4, ADPRTL1, KIAA0177, PARPL | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 4 (EC 2.4.2.30) (PARP-4) (Vault poly(ADP- ribose) polymerase) (VPARP) (193 kDa vault protein) (PARP- related/IalphaI-related H5/proline-rich) (PH5P). | |||||
|
PARP8_HUMAN
|
||||||
| NC score | 0.188001 (rank : 7) | θ value | 0.00228821 (rank : 7) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8N3A8, Q3KRB7, Q6DHZ1, Q9H754 | Gene names | PARP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 8 (EC 2.4.2.30) (PARP-8). | |||||
|
PARP8_MOUSE
|
||||||
| NC score | 0.187701 (rank : 8) | θ value | 0.00228821 (rank : 8) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q3UD82, Q3UDE4, Q3UDU5, Q8VCB5, Q9CYY3 | Gene names | Parp8, D13Ertd275e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 8 (EC 2.4.2.30) (PARP-8). | |||||
|
PARP6_HUMAN
|
||||||
| NC score | 0.181736 (rank : 9) | θ value | 0.0330416 (rank : 10) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q2NL67, Q9H7C5, Q9H9X6, Q9HAF3, Q9NPS6, Q9UFG4 | Gene names | PARP6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 6 (EC 2.4.2.30) (PARP-6). | |||||
|
PARP6_MOUSE
|
||||||
| NC score | 0.181633 (rank : 10) | θ value | 0.0330416 (rank : 11) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6P6P7, Q8BVW1 | Gene names | Parp6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly [ADP-ribose] polymerase 6 (EC 2.4.2.30) (PARP-6). | |||||
|
PARPT_MOUSE
|
||||||
| NC score | 0.090260 (rank : 11) | θ value | 0.365318 (rank : 17) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8C1B2, Q3UD50, Q8C032 | Gene names | Tiparp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TCDD-inducible poly [ADP-ribose] polymerase (EC 2.4.2.30). | |||||
|
PARPT_HUMAN
|
||||||
| NC score | 0.073872 (rank : 12) | θ value | 0.365318 (rank : 16) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q7Z3E1, Q68CY9, Q86VP4, Q9Y4P7 | Gene names | TIPARP, PARP7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TCDD-inducible poly [ADP-ribose] polymerase (EC 2.4.2.30) (Poly [ADP- ribose] polymerase 7) (PARP-7). | |||||
|
DNL3_HUMAN
|
||||||
| NC score | 0.071049 (rank : 13) | θ value | θ > 10 (rank : 54) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P49916, Q16714 | Gene names | LIG3 | |||
|
Domain Architecture |
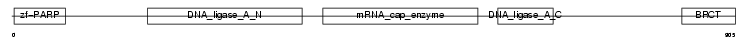
|
|||||
| Description | DNA ligase 3 (EC 6.5.1.1) (DNA ligase III) (Polydeoxyribonucleotide synthase [ATP] 3). | |||||
|
DNL3_MOUSE
|
||||||
| NC score | 0.069153 (rank : 14) | θ value | θ > 10 (rank : 55) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P97386, P97385 | Gene names | Lig3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA ligase 3 (EC 6.5.1.1) (DNA ligase III) (Polydeoxyribonucleotide synthase [ATP] 3). | |||||
|
TNKS1_HUMAN
|
||||||
| NC score | 0.068156 (rank : 15) | θ value | 0.0193708 (rank : 9) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 477 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O95271, O95272 | Gene names | TNKS, PARP5A, PARPL, TIN1, TINF1, TNKS1 | |||
|
Domain Architecture |

|
|||||
| Description | Tankyrase-1 (EC 2.4.2.30) (TANK1) (Tankyrase I) (TNKS-1) (TRF1- interacting ankyrin-related ADP-ribose polymerase). | |||||
|
TNKS2_HUMAN
|
||||||
| NC score | 0.063564 (rank : 16) | θ value | 0.0563607 (rank : 12) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9H2K2, Q9H8F2, Q9HAS4 | Gene names | TNKS2, PARP5B, TANK2, TNKL | |||
|
Domain Architecture |

|
|||||
| Description | Tankyrase-2 (EC 2.4.2.30) (TANK2) (Tankyrase II) (TNKS-2) (TRF1- interacting ankyrin-related ADP-ribose polymerase 2) (Tankyrase-like protein) (Tankyrase-related protein). | |||||
|
LHR2A_MOUSE
|
||||||
| NC score | 0.054108 (rank : 17) | θ value | θ > 10 (rank : 57) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 1 | |
| SwissProt Accessions | Q99KC8, Q3TTU2, Q6UN22, Q8BHA8, Q9CTV9 | Gene names | Loh11cr2a | |||
|
Domain Architecture |

|
|||||
| Description | Loss of heterozygosity 11 chromosomal region 2 gene A protein homolog. | |||||
|
LHR2A_HUMAN
|
||||||
| NC score | 0.050838 (rank : 18) | θ value | θ > 10 (rank : 56) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 1 | |
| SwissProt Accessions | O00534, Q6UN19, Q6UN20, Q9BVF8 | Gene names | LOH11CR2A, BCSC1 | |||
|
Domain Architecture |

|
|||||
| Description | Loss of heterozygosity 11 chromosomal region 2 gene A protein (Breast cancer suppressor candidate 1) (BCSC-1). | |||||
|
TRAIP_HUMAN
|
||||||
| NC score | 0.042750 (rank : 19) | θ value | 0.125558 (rank : 13) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 780 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9BWF2, O00467 | Gene names | TRAIP, TRIP | |||
|
Domain Architecture |

|
|||||
| Description | TRAF-interacting protein. | |||||
|
ABTAP_HUMAN
|
||||||
| NC score | 0.042049 (rank : 20) | θ value | 0.47712 (rank : 18) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9H501, Q86X92, Q9H9Q5, Q9HA35, Q9NX93, Q9P1S6 | Gene names | ABTAP, C20orf6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ABT1-associated protein. | |||||
|
FLOT2_HUMAN
|
||||||
| NC score | 0.041228 (rank : 21) | θ value | 2.36792 (rank : 32) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14254 | Gene names | FLOT2, ESA1, M17S1 | |||
|
Domain Architecture |

|
|||||
| Description | Flotillin-2 (Epidermal surface antigen) (ESA). | |||||
|
FLOT2_MOUSE
|
||||||
| NC score | 0.041094 (rank : 22) | θ value | 2.36792 (rank : 33) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q60634, Q6NS75 | Gene names | Flot2, Esa1, M17s1 | |||
|
Domain Architecture |

|
|||||
| Description | Flotillin-2 (Epidermal surface antigen) (ESA). | |||||
|
STML2_MOUSE
|
||||||
| NC score | 0.039187 (rank : 23) | θ value | 1.38821 (rank : 26) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99JB2, Q9DCG8 | Gene names | Stoml2, Slp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stomatin-like protein 2 (SLP-2). | |||||
|
STML2_HUMAN
|
||||||
| NC score | 0.039179 (rank : 24) | θ value | 1.38821 (rank : 25) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UJZ1, O60376, Q53G29, Q96FY2, Q9P042 | Gene names | STOML2, SLP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stomatin-like protein 2 (SLP-2) (EPB72-like 2). | |||||
|
TRAIP_MOUSE
|
||||||
| NC score | 0.034767 (rank : 25) | θ value | 1.06291 (rank : 22) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 745 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8VIG6, O08854, Q922M8, Q9CPP4 | Gene names | Traip, Trip | |||
|
Domain Architecture |

|
|||||
| Description | TRAF-interacting protein. | |||||
|
ABTAP_MOUSE
|
||||||
| NC score | 0.033812 (rank : 26) | θ value | 4.03905 (rank : 37) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q3V1V3 | Gene names | Abtap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ABT1-associated protein. | |||||
|
LIPB2_MOUSE
|
||||||
| NC score | 0.029643 (rank : 27) | θ value | 0.813845 (rank : 20) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 570 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O35711, Q7TMG4, Q8CBS6, Q99KX6 | Gene names | Ppfibp2, Cclp1 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-beta-2 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein-binding protein 2) (PTPRF-interacting protein-binding protein 2) (Coiled-coil-like protein 1). | |||||
|
LIPB2_HUMAN
|
||||||
| NC score | 0.029070 (rank : 28) | θ value | 1.38821 (rank : 24) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 599 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8ND30, O75337, Q8WW26 | Gene names | PPFIBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-beta-2 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein-binding protein 2) (PTPRF-interacting protein-binding protein 2). | |||||
|
RBP24_HUMAN
|
||||||
| NC score | 0.027735 (rank : 29) | θ value | 0.21417 (rank : 15) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q4ZG46 | Gene names | RANBP2L4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 2-like 4 (RanBP2L4). | |||||
|
GCC2_HUMAN
|
||||||
| NC score | 0.027703 (rank : 30) | θ value | 0.21417 (rank : 14) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1558 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8IWJ2, O15045, Q8TDH3, Q9H2G8 | Gene names | GCC2, KIAA0336 | |||
|
Domain Architecture |

|
|||||
| Description | GRIP and coiled-coil domain-containing protein 2 (Golgi coiled coil protein GCC185) (CTCL tumor antigen se1-1) (CLL-associated antigen KW- 11) (NY-REN-53 antigen). | |||||
|
AQR_MOUSE
|
||||||
| NC score | 0.026820 (rank : 31) | θ value | 0.813845 (rank : 19) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CFQ3, P97871, Q3U9N1, Q3ULE8, Q80TX8 | Gene names | Aqr, Kiaa0560 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intron-binding protein aquarius. | |||||
|
SMC4_MOUSE
|
||||||
| NC score | 0.024319 (rank : 32) | θ value | 1.81305 (rank : 31) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 830 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8CG47, Q8BTS7, Q8BTY9, Q99K21 | Gene names | Smc4l1, Capc, Smc4 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosomes 4-like 1 protein (Chromosome- associated polypeptide C) (XCAP-C homolog). | |||||
|
TPM4_HUMAN
|
||||||
| NC score | 0.023561 (rank : 33) | θ value | 0.813845 (rank : 21) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 525 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P67936, P07226, Q15659, Q9BU85, Q9H8Q3 | Gene names | TPM4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tropomyosin alpha-4 chain (Tropomyosin-4) (TM30p1). | |||||
|
SMC4_HUMAN
|
||||||
| NC score | 0.022453 (rank : 34) | θ value | 4.03905 (rank : 39) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 937 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9NTJ3, O95752, Q8NDL4, Q9UNT9 | Gene names | SMC4L1, CAPC, SMC4 | |||
|
Domain Architecture |
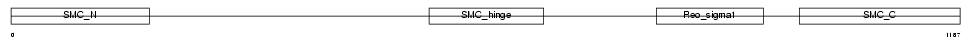
|
|||||
| Description | Structural maintenance of chromosomes 4-like 1 protein (Chromosome- associated polypeptide C) (hCAP-C) (XCAP-C homolog). | |||||
|
TPM4_MOUSE
|
||||||
| NC score | 0.022358 (rank : 35) | θ value | 3.0926 (rank : 36) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 703 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6IRU2 | Gene names | Tpm4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tropomyosin alpha-4 chain (Tropomyosin-4). | |||||
|
CN135_HUMAN
|
||||||
| NC score | 0.022126 (rank : 36) | θ value | 5.27518 (rank : 40) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q63HM2, Q9BQG8, Q9H9F2 | Gene names | C14orf135, FBP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf135 precursor (Hepatitis C virus F protein-binding protein 2) (HCV F protein-binding protein 2). | |||||
|
S39A6_MOUSE
|
||||||
| NC score | 0.021797 (rank : 37) | θ value | 1.81305 (rank : 30) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8C145, Q7TPP9, Q7TQE0, Q8R518 | Gene names | Slc39a6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc transporter SLC39A6 precursor (Solute carrier family 39 member 6) (Endoplasmic reticulum membrane-linked protein) (Ermelin). | |||||
|
INVO_HUMAN
|
||||||
| NC score | 0.021362 (rank : 38) | θ value | 1.81305 (rank : 29) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 636 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P07476 | Gene names | IVL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Involucrin. | |||||
|
GA2L3_HUMAN
|
||||||
| NC score | 0.020695 (rank : 39) | θ value | 1.81305 (rank : 28) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q86XJ1 | Gene names | GAS2L3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GAS2-like protein 3 (Growth arrest-specific 2-like 3). | |||||
|
UACA_HUMAN
|
||||||
| NC score | 0.020178 (rank : 40) | θ value | 1.38821 (rank : 27) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1622 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9BZF9, Q8N3B8, Q9HCL1, Q9NWC6 | Gene names | UACA, KIAA1561 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uveal autoantigen with coiled-coil domains and ankyrin repeats protein. | |||||
|
DNM3B_MOUSE
|
||||||
| NC score | 0.019272 (rank : 41) | θ value | 1.38821 (rank : 23) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O88509, O88510, O88511 | Gene names | Dnmt3b | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 3B (EC 2.1.1.37) (Dnmt3b) (DNA methyltransferase MmuIIIB) (DNA MTase MmuIIIB) (M.MmuIIIB). | |||||
|
USBP1_HUMAN
|
||||||
| NC score | 0.015978 (rank : 42) | θ value | 6.88961 (rank : 46) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8N6Y0, Q8NBX7, Q96KH3, Q9BYI8 | Gene names | USHBP1, AIEBP, MCC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USH1C-binding protein 1 (Usher syndrome type-1C protein-binding protein 1) (MCC-2) (AIE-75 binding protein). | |||||
|
KTN1_HUMAN
|
||||||
| NC score | 0.015829 (rank : 43) | θ value | 8.99809 (rank : 50) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1197 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q86UP2, Q13999, Q14707, Q15387, Q86W57 | Gene names | KTN1, CG1, KIAA0004 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinectin (Kinesin receptor) (CG-1 antigen). | |||||
|
BRD8_HUMAN
|
||||||
| NC score | 0.015358 (rank : 44) | θ value | 3.0926 (rank : 35) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H0E9, O43178, Q15355, Q59GN0, Q969M9 | Gene names | BRD8, SMAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 8 (p120) (Skeletal muscle abundant protein) (Thyroid hormone receptor coactivating protein 120kDa) (TrCP120). | |||||
|
BRAP_MOUSE
|
||||||
| NC score | 0.015289 (rank : 45) | θ value | 8.99809 (rank : 48) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q99MP8, Q8CC00, Q8CHX1, Q99MP7, Q9CXX8 | Gene names | Brap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BRCA1-associated protein (EC 6.3.2.-) (BRAP2) (Impedes mitogenic signal propagation) (IMP). | |||||
|
JAD1A_HUMAN
|
||||||
| NC score | 0.015129 (rank : 46) | θ value | 8.99809 (rank : 49) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P29375 | Gene names | JARID1A, RBBP2, RBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1A (Retinoblastoma-binding protein 2) (RBBP-2). | |||||
|
RENT2_HUMAN
|
||||||
| NC score | 0.014372 (rank : 47) | θ value | 6.88961 (rank : 45) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 618 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9HAU5, Q8N8U1, Q9H1J2, Q9NWL1, Q9P2D9, Q9Y4M9 | Gene names | UPF2, KIAA1408, RENT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of nonsense transcripts 2 (Nonsense mRNA reducing factor 2) (Up-frameshift suppressor 2 homolog) (hUpf2). | |||||
|
GOGA2_MOUSE
|
||||||
| NC score | 0.014171 (rank : 48) | θ value | 6.88961 (rank : 43) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 944 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q921M4 | Gene names | Golga2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgin subfamily A member 2 (Cis-Golgi matrix protein GM130). | |||||
|
ASPH_HUMAN
|
||||||
| NC score | 0.013862 (rank : 49) | θ value | 8.99809 (rank : 47) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q12797 | Gene names | ASPH | |||
|
Domain Architecture |

|
|||||
| Description | Aspartyl/asparaginyl beta-hydroxylase (EC 1.14.11.16) (Aspartate beta- hydroxylase) (ASP beta-hydroxylase) (Peptide-aspartate beta- dioxygenase). | |||||
|
TPR_HUMAN
|
||||||
| NC score | 0.013389 (rank : 50) | θ value | 8.99809 (rank : 53) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
MYO5C_HUMAN
|
||||||
| NC score | 0.012884 (rank : 51) | θ value | 5.27518 (rank : 42) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1206 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9NQX4 | Gene names | MYO5C | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5C (Myosin Vc). | |||||
|
OLR1_MOUSE
|
||||||
| NC score | 0.012400 (rank : 52) | θ value | 6.88961 (rank : 44) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 596 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9EQ09 | Gene names | Olr1, Lox1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oxidized low-density lipoprotein receptor 1 (Ox-LDL receptor 1) (Lectin-type oxidized LDL receptor 1) (Lectin-like oxidized LDL receptor 1) (Lectin-like oxLDL receptor 1) (LOX-1) [Contains: Oxidized low-density lipoprotein receptor 1, soluble form]. | |||||
|
MYH8_HUMAN
|
||||||
| NC score | 0.012272 (rank : 53) | θ value | 4.03905 (rank : 38) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1610 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P13535, Q14910 | Gene names | MYH8 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-8 (Myosin heavy chain, skeletal muscle, perinatal) (MyHC- perinatal). | |||||
|
STX1C_HUMAN
|
||||||
| NC score | 0.010969 (rank : 54) | θ value | 8.99809 (rank : 51) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 270 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P61266 | Gene names | STX1B2 | |||
|
Domain Architecture |

|
|||||
| Description | Syntaxin-1B2. | |||||
|
STX1C_MOUSE
|
||||||
| NC score | 0.010969 (rank : 55) | θ value | 8.99809 (rank : 52) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 270 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P61264, P32853 | Gene names | Stx1b2, Stx1b | |||
|
Domain Architecture |

|
|||||
| Description | Syntaxin-1B2 (Syntaxin 1B). | |||||
|
GUC2D_HUMAN
|
||||||
| NC score | 0.006119 (rank : 56) | θ value | 5.27518 (rank : 41) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 715 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q02846, Q6LEA7 | Gene names | GUCY2D, CORD6, GUC1A4, GUC2D, RETGC, RETGC1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinal guanylyl cyclase 1 precursor (EC 4.6.1.2) (Guanylate cyclase 2D, retinal) (RETGC-1) (Rod outer segment membrane guanylate cyclase) (ROS-GC). | |||||
|
GUC2F_HUMAN
|
||||||
| NC score | 0.005725 (rank : 57) | θ value | 2.36792 (rank : 34) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 740 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P51841, Q9UJF1 | Gene names | GUCY2F, GUC2F, RETGC2 | |||
|
Domain Architecture |

|
|||||
| Description | Retinal guanylyl cyclase 2 precursor (EC 4.6.1.2) (Guanylate cyclase 2F, retinal) (RETGC-2) (Rod outer segment membrane guanylate cyclase 2) (ROS-GC2) (Guanylate cyclase F) (GC-F). | |||||