Please be patient as the page loads
|
I17RA_HUMAN
|
||||||
| SwissProt Accessions | Q96F46, O43844 | Gene names | IL17RA, IL17R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interleukin-17 receptor A precursor (IL-17 receptor). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
I17RA_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q96F46, O43844 | Gene names | IL17RA, IL17R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interleukin-17 receptor A precursor (IL-17 receptor). | |||||
|
I17RA_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.987260 (rank : 2) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q60943 | Gene names | IL17ra, Il17r | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interleukin-17 receptor A precursor (IL-17 receptor). | |||||
|
I17RB_HUMAN
|
||||||
| θ value | 3.61944e-33 (rank : 3) | NC score | 0.728568 (rank : 3) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NRM6, Q9BPZ0, Q9NRL4, Q9NRM5 | Gene names | IL17RB, EVI27, IL17BR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interleukin-17 receptor B precursor (IL-17 receptor B) (IL-17RB) (Interleukin-17B receptor) (IL-17B receptor) (IL-17 receptor homolog 1) (IL-17Rh1) (IL17Rh1) (Cytokine receptor CRL4). | |||||
|
I17RB_MOUSE
|
||||||
| θ value | 4.89182e-30 (rank : 4) | NC score | 0.705037 (rank : 4) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9JIP3, Q9JIP2 | Gene names | Il17rb, Evi27, Il17br | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interleukin-17 receptor B precursor (IL-17 receptor B) (IL-17RB) (Interleukin-17B receptor) (IL-17B receptor) (IL-17 receptor homolog 1) (IL-17Rh1) (IL17Rh1) (IL-17ER). | |||||
|
I17RD_HUMAN
|
||||||
| θ value | 7.0802e-21 (rank : 5) | NC score | 0.569717 (rank : 5) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NFM7, Q58EZ7, Q6RVF4, Q6UWI5, Q8N113, Q8NFS0, Q9UFA0 | Gene names | IL17RD, IL17RLM, SEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interleukin-17 receptor D precursor (IL-17 receptor D) (IL-17RD) (Interleukin-17D receptor) (IL-17D receptor) (IL17Rhom) (Interleukin- 17 receptor-like protein) (Sef homolog) (hSef). | |||||
|
I17RD_MOUSE
|
||||||
| θ value | 6.20254e-17 (rank : 6) | NC score | 0.541600 (rank : 6) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8JZL1, Q8K447, Q8R5J8 | Gene names | Il17rd, Il17rlm, Sef | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interleukin-17 receptor D precursor (IL-17 receptor D) (IL-17RD) (Interleukin-17D receptor) (IL-17D receptor) (Interleukin-17 receptor- like protein) (Sef homolog) (mSef). | |||||
|
I17RC_HUMAN
|
||||||
| θ value | 1.17247e-07 (rank : 7) | NC score | 0.296538 (rank : 7) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NAC3, Q6UVY3, Q6UWD4, Q8NFS1, Q9BR97 | Gene names | IL17RC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interleukin-17 receptor C precursor (IL-17 receptor C) (IL-17RC) (Interleukin-17 receptor-like protein) (IL-17RL) (Interleukin-17 receptor homolog) (IL17Rhom). | |||||
|
I17RC_MOUSE
|
||||||
| θ value | 4.92598e-06 (rank : 8) | NC score | 0.286558 (rank : 8) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8K4C2, Q99J43 | Gene names | Il17rc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interleukin-17 receptor C precursor (IL-17 receptor C) (IL-17RC) (Interleukin-17 receptor-like protein) (IL-17RL). | |||||
|
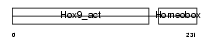
HXB9_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 9) | NC score | 0.022534 (rank : 14) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P20615 | Gene names | Hoxb9, Hox-2.5, Hoxb-9 | |||
|
Domain Architecture |
|
|||||
| Description | Homeobox protein Hox-B9 (Hox-2.5). | |||||
|
ZN447_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 10) | NC score | 0.015065 (rank : 22) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 899 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TBC5, Q9BRK7, Q9H9A0 | Gene names | ZNF447 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 447. | |||||
|
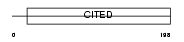
CITE1_MOUSE
|
||||||
| θ value | 0.21417 (rank : 11) | NC score | 0.062812 (rank : 9) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P97769 | Gene names | Cited1, Msg1 | |||
|
Domain Architecture |
|
|||||
| Description | Cbp/p300-interacting transactivator 1 (Melanocyte-specific protein 1). | |||||
|
SIAH2_HUMAN
|
||||||
| θ value | 0.21417 (rank : 12) | NC score | 0.037065 (rank : 10) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O43255, O43270 | Gene names | SIAH2 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin ligase SIAH2 (EC 6.3.2.-) (Seven in absentia homolog 2) (Siah-2) (hSiah2). | |||||
|
ANR38_MOUSE
|
||||||
| θ value | 0.62314 (rank : 13) | NC score | 0.020616 (rank : 17) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 512 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6P9J5, Q8BV03 | Gene names | Ankrd38 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 38. | |||||
|
ATS20_MOUSE
|
||||||
| θ value | 0.813845 (rank : 14) | NC score | 0.011230 (rank : 26) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P59511 | Gene names | Adamts20 | |||
|
Domain Architecture |

|
|||||
| Description | ADAMTS-20 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase with thrombospondin motifs 20) (ADAM-TS 20) (ADAM-TS20). | |||||
|
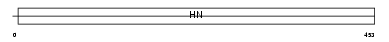
NEUR4_HUMAN
|
||||||
| θ value | 0.813845 (rank : 15) | NC score | 0.029840 (rank : 11) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8WWR8, Q96D64 | Gene names | NEU4 | |||
|
Domain Architecture |
|
|||||
| Description | Sialidase-4 (EC 3.2.1.18) (N-acetyl-alpha-neuraminidase 4). | |||||
|
TIF1B_HUMAN
|
||||||
| θ value | 1.06291 (rank : 16) | NC score | 0.018823 (rank : 19) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q13263, O00677, Q7Z632, Q93040, Q96IM1 | Gene names | TRIM28, KAP1, RNF96, TIF1B | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-beta (TIF1-beta) (Tripartite motif-containing protein 28) (Nuclear corepressor KAP-1) (KRAB- associated protein 1) (KAP-1) (KRAB-interacting protein 1) (KRIP-1) (RING finger protein 96). | |||||
|
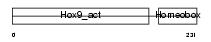
HXB9_HUMAN
|
||||||
| θ value | 2.36792 (rank : 17) | NC score | 0.016086 (rank : 21) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P17482, Q9H1I1 | Gene names | HOXB9, HOX2E | |||
|
Domain Architecture |
|
|||||
| Description | Homeobox protein Hox-B9 (Hox-2E) (Hox-2.5). | |||||
|
UB7I1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 18) | NC score | 0.029310 (rank : 12) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P58283, Q3U493, Q3UE56, Q3UGM3, Q68FN0, Q6P1H8, Q6PWY5, Q8BN27, Q8C1U3 | Gene names | Ubce7ip1, Triad3, Uip83, Zin | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin ligase TRIAD3 (EC 6.3.2.-) (Ubiquitin-conjugating enzyme 7-interacting protein 1) (UbcM4-interacting protein 83) (Triad domain- containing protein 3). | |||||
|
PSB10_MOUSE
|
||||||
| θ value | 3.0926 (rank : 19) | NC score | 0.017546 (rank : 20) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O35955, O08687, Q99KC5 | Gene names | Psmb10, Lmp10, Mecl1 | |||
|
Domain Architecture |
|
|||||
| Description | Proteasome subunit beta type 10 precursor (EC 3.4.25.1) (Proteasome MECl-1) (Macropain subunit MECl-1) (Multicatalytic endopeptidase complex subunit MECl-1). | |||||
|
CARD6_HUMAN
|
||||||
| θ value | 4.03905 (rank : 20) | NC score | 0.025252 (rank : 13) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BX69 | Gene names | CARD6 | |||
|
Domain Architecture |
|
|||||
| Description | Caspase recruitment domain-containing protein 6. | |||||
|
CEBPD_MOUSE
|
||||||
| θ value | 4.03905 (rank : 21) | NC score | 0.018838 (rank : 18) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q00322 | Gene names | Cebpd, Crp3 | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein delta (C/EBP delta) (C/EBP-related protein 3). | |||||
|
MILK2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 22) | NC score | 0.011273 (rank : 25) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8IY33, Q7RTP4, Q7Z655, Q8TEQ4, Q9H5F9 | Gene names | ||||
|
Domain Architecture |
|
|||||
| Description | MICAL-like protein 2. | |||||
|
ETV3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 23) | NC score | 0.010773 (rank : 29) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P41162, Q8TAC8, Q9BX30 | Gene names | ETV3, METS, PE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
PURA1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 24) | NC score | 0.014571 (rank : 24) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P28650, Q8CHQ1 | Gene names | Adssl1, Adss1 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylosuccinate synthetase isozyme 1 (EC 6.3.4.4) (Adenylosuccinate synthetase, muscle isozyme) (IMP--aspartate ligase 1) (AdSS 1) (AMPSase 1). | |||||
|
TNFC_MOUSE
|
||||||
| θ value | 5.27518 (rank : 25) | NC score | 0.021290 (rank : 16) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P41155 | Gene names | Ltb, Tnfc, Tnfsf3 | |||
|
Domain Architecture |
|
|||||
| Description | Lymphotoxin-beta (LT-beta) (Tumor necrosis factor C) (TNF-C) (Tumor necrosis factor ligand superfamily member 3). | |||||
|
B4GT1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 26) | NC score | 0.008604 (rank : 33) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P15535 | Gene names | B4galt1, Ggtb, Ggtb2 | |||
|
Domain Architecture |
|
|||||
| Description | Beta-1,4-galactosyltransferase 1 (EC 2.4.1.-) (Beta-1,4-GalTase 1) (Beta4Gal-T1) (b4Gal-T1) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 1) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 1) [Includes: Lactose synthase A protein (EC 2.4.1.22); N-acetyllactosamine synthase (EC 2.4.1.90) (Nal synthetase); Beta-N-acetylglucosaminylglycopeptide beta-1,4- galactosyltransferase (EC 2.4.1.38); Beta-N-acetylglucosaminyl- glycolipid beta-1,4-galactosyltransferase (EC 2.4.1.-)]. | |||||
|
CSTFT_HUMAN
|
||||||
| θ value | 6.88961 (rank : 27) | NC score | 0.010285 (rank : 30) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H0L4, O75174, Q53HK6, Q7LGE8, Q8N6T1 | Gene names | CSTF2T, KIAA0689 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage stimulation factor 64 kDa subunit, tau variant (CSTF 64 kDa subunit, tau variant) (CF-1 64 kDa subunit, tau variant) (TauCstF-64). | |||||
|
CSTFT_MOUSE
|
||||||
| θ value | 6.88961 (rank : 28) | NC score | 0.009911 (rank : 31) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C7E9, Q6A016, Q8BHH7, Q8C3W7, Q8R2Y1, Q9EPU3 | Gene names | Cstf2t, Kiaa0689 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage stimulation factor 64 kDa subunit, tau variant (CSTF 64 kDa subunit, tau variant) (CF-1 64 kDa subunit, tau variant) (TauCstF-64). | |||||
|
GP152_HUMAN
|
||||||
| θ value | 6.88961 (rank : 29) | NC score | 0.005739 (rank : 35) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 636 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8TDT2, Q86SM0 | Gene names | GPR152, PGR5 | |||
|
Domain Architecture |

|
|||||
| Description | Probable G-protein coupled receptor 152 (G-protein coupled receptor PGR5). | |||||
|
HDAC6_HUMAN
|
||||||
| θ value | 6.88961 (rank : 30) | NC score | 0.008565 (rank : 34) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UBN7 | Gene names | HDAC6 | |||
|
Domain Architecture |

|
|||||
| Description | Histone deacetylase 6 (HD6). | |||||
|
RINI_HUMAN
|
||||||
| θ value | 6.88961 (rank : 31) | NC score | 0.008747 (rank : 32) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P13489 | Gene names | RNH1, PRI, RNH | |||
|
Domain Architecture |
|
|||||
| Description | Ribonuclease inhibitor (Ribonuclease/angiogenin inhibitor 1) (RAI) (Placental ribonuclease inhibitor) (RNase inhibitor) (RI). | |||||
|
HD_HUMAN
|
||||||
| θ value | 8.99809 (rank : 32) | NC score | 0.010984 (rank : 28) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P42858, Q9UQB7 | Gene names | HD, IT15 | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin (Huntington disease protein) (HD protein). | |||||
|
HEMGN_MOUSE
|
||||||
| θ value | 8.99809 (rank : 33) | NC score | 0.014787 (rank : 23) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9ERZ0, Q3TAE6, Q80YT2, Q8CF29 | Gene names | Hemgn, Ndr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hemogen (Hemopoietic gene protein) (Negative differentiation regulator protein) (mNDR). | |||||
|
LGR5_MOUSE
|
||||||
| θ value | 8.99809 (rank : 34) | NC score | 0.002205 (rank : 36) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 484 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Z1P4 | Gene names | Lgr5, Fex, Gpr49 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeat-containing G-protein coupled receptor 5 precursor (G-protein coupled receptor 49) (Orphan G-protein coupled receptor FEX). | |||||
|
TNR27_MOUSE
|
||||||
| θ value | 8.99809 (rank : 35) | NC score | 0.011134 (rank : 27) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BX35, Q8BM50 | Gene names | Eda2r, Tnfrsf27, Xedar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor necrosis factor receptor superfamily member 27 (X-linked ectodysplasin-A2 receptor) (EDA-A2 receptor). | |||||
|
UB7I1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 36) | NC score | 0.022223 (rank : 15) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NWF9, Q6Y691, Q75ML7, Q7Z2H7, Q7Z7C1, Q8NHW7, Q9NYT1 | Gene names | UBCE7IP1, ZIN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin ligase TRIAD3 (EC 6.3.2.-) (Ubiquitin-conjugating enzyme 7-interacting protein 1) (Zinc finger protein inhibiting NF-kappa-B) (Triad domain-containing protein 3). | |||||
|
I17RA_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q96F46, O43844 | Gene names | IL17RA, IL17R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interleukin-17 receptor A precursor (IL-17 receptor). | |||||
|
I17RA_MOUSE
|
||||||
| NC score | 0.987260 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q60943 | Gene names | IL17ra, Il17r | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interleukin-17 receptor A precursor (IL-17 receptor). | |||||
|
I17RB_HUMAN
|
||||||
| NC score | 0.728568 (rank : 3) | θ value | 3.61944e-33 (rank : 3) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NRM6, Q9BPZ0, Q9NRL4, Q9NRM5 | Gene names | IL17RB, EVI27, IL17BR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interleukin-17 receptor B precursor (IL-17 receptor B) (IL-17RB) (Interleukin-17B receptor) (IL-17B receptor) (IL-17 receptor homolog 1) (IL-17Rh1) (IL17Rh1) (Cytokine receptor CRL4). | |||||
|
I17RB_MOUSE
|
||||||
| NC score | 0.705037 (rank : 4) | θ value | 4.89182e-30 (rank : 4) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9JIP3, Q9JIP2 | Gene names | Il17rb, Evi27, Il17br | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interleukin-17 receptor B precursor (IL-17 receptor B) (IL-17RB) (Interleukin-17B receptor) (IL-17B receptor) (IL-17 receptor homolog 1) (IL-17Rh1) (IL17Rh1) (IL-17ER). | |||||
|
I17RD_HUMAN
|
||||||
| NC score | 0.569717 (rank : 5) | θ value | 7.0802e-21 (rank : 5) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NFM7, Q58EZ7, Q6RVF4, Q6UWI5, Q8N113, Q8NFS0, Q9UFA0 | Gene names | IL17RD, IL17RLM, SEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interleukin-17 receptor D precursor (IL-17 receptor D) (IL-17RD) (Interleukin-17D receptor) (IL-17D receptor) (IL17Rhom) (Interleukin- 17 receptor-like protein) (Sef homolog) (hSef). | |||||
|
I17RD_MOUSE
|
||||||
| NC score | 0.541600 (rank : 6) | θ value | 6.20254e-17 (rank : 6) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8JZL1, Q8K447, Q8R5J8 | Gene names | Il17rd, Il17rlm, Sef | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interleukin-17 receptor D precursor (IL-17 receptor D) (IL-17RD) (Interleukin-17D receptor) (IL-17D receptor) (Interleukin-17 receptor- like protein) (Sef homolog) (mSef). | |||||
|
I17RC_HUMAN
|
||||||
| NC score | 0.296538 (rank : 7) | θ value | 1.17247e-07 (rank : 7) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NAC3, Q6UVY3, Q6UWD4, Q8NFS1, Q9BR97 | Gene names | IL17RC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interleukin-17 receptor C precursor (IL-17 receptor C) (IL-17RC) (Interleukin-17 receptor-like protein) (IL-17RL) (Interleukin-17 receptor homolog) (IL17Rhom). | |||||
|
I17RC_MOUSE
|
||||||
| NC score | 0.286558 (rank : 8) | θ value | 4.92598e-06 (rank : 8) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8K4C2, Q99J43 | Gene names | Il17rc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interleukin-17 receptor C precursor (IL-17 receptor C) (IL-17RC) (Interleukin-17 receptor-like protein) (IL-17RL). | |||||
|
CITE1_MOUSE
|
||||||
| NC score | 0.062812 (rank : 9) | θ value | 0.21417 (rank : 11) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P97769 | Gene names | Cited1, Msg1 | |||
|
Domain Architecture |

|
|||||
| Description | Cbp/p300-interacting transactivator 1 (Melanocyte-specific protein 1). | |||||
|
SIAH2_HUMAN
|
||||||
| NC score | 0.037065 (rank : 10) | θ value | 0.21417 (rank : 12) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O43255, O43270 | Gene names | SIAH2 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin ligase SIAH2 (EC 6.3.2.-) (Seven in absentia homolog 2) (Siah-2) (hSiah2). | |||||
|
NEUR4_HUMAN
|
||||||
| NC score | 0.029840 (rank : 11) | θ value | 0.813845 (rank : 15) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8WWR8, Q96D64 | Gene names | NEU4 | |||
|
Domain Architecture |

|
|||||
| Description | Sialidase-4 (EC 3.2.1.18) (N-acetyl-alpha-neuraminidase 4). | |||||
|
UB7I1_MOUSE
|
||||||
| NC score | 0.029310 (rank : 12) | θ value | 2.36792 (rank : 18) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P58283, Q3U493, Q3UE56, Q3UGM3, Q68FN0, Q6P1H8, Q6PWY5, Q8BN27, Q8C1U3 | Gene names | Ubce7ip1, Triad3, Uip83, Zin | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin ligase TRIAD3 (EC 6.3.2.-) (Ubiquitin-conjugating enzyme 7-interacting protein 1) (UbcM4-interacting protein 83) (Triad domain- containing protein 3). | |||||
|
CARD6_HUMAN
|
||||||
| NC score | 0.025252 (rank : 13) | θ value | 4.03905 (rank : 20) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BX69 | Gene names | CARD6 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 6. | |||||
|
HXB9_MOUSE
|
||||||
| NC score | 0.022534 (rank : 14) | θ value | 0.0961366 (rank : 9) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P20615 | Gene names | Hoxb9, Hox-2.5, Hoxb-9 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-B9 (Hox-2.5). | |||||
|
UB7I1_HUMAN
|
||||||
| NC score | 0.022223 (rank : 15) | θ value | 8.99809 (rank : 36) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NWF9, Q6Y691, Q75ML7, Q7Z2H7, Q7Z7C1, Q8NHW7, Q9NYT1 | Gene names | UBCE7IP1, ZIN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin ligase TRIAD3 (EC 6.3.2.-) (Ubiquitin-conjugating enzyme 7-interacting protein 1) (Zinc finger protein inhibiting NF-kappa-B) (Triad domain-containing protein 3). | |||||
|
TNFC_MOUSE
|
||||||
| NC score | 0.021290 (rank : 16) | θ value | 5.27518 (rank : 25) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P41155 | Gene names | Ltb, Tnfc, Tnfsf3 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphotoxin-beta (LT-beta) (Tumor necrosis factor C) (TNF-C) (Tumor necrosis factor ligand superfamily member 3). | |||||
|
ANR38_MOUSE
|
||||||
| NC score | 0.020616 (rank : 17) | θ value | 0.62314 (rank : 13) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 512 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6P9J5, Q8BV03 | Gene names | Ankrd38 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 38. | |||||
|
CEBPD_MOUSE
|
||||||
| NC score | 0.018838 (rank : 18) | θ value | 4.03905 (rank : 21) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q00322 | Gene names | Cebpd, Crp3 | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein delta (C/EBP delta) (C/EBP-related protein 3). | |||||
|
TIF1B_HUMAN
|
||||||
| NC score | 0.018823 (rank : 19) | θ value | 1.06291 (rank : 16) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q13263, O00677, Q7Z632, Q93040, Q96IM1 | Gene names | TRIM28, KAP1, RNF96, TIF1B | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-beta (TIF1-beta) (Tripartite motif-containing protein 28) (Nuclear corepressor KAP-1) (KRAB- associated protein 1) (KAP-1) (KRAB-interacting protein 1) (KRIP-1) (RING finger protein 96). | |||||
|
PSB10_MOUSE
|
||||||
| NC score | 0.017546 (rank : 20) | θ value | 3.0926 (rank : 19) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O35955, O08687, Q99KC5 | Gene names | Psmb10, Lmp10, Mecl1 | |||
|
Domain Architecture |

|
|||||
| Description | Proteasome subunit beta type 10 precursor (EC 3.4.25.1) (Proteasome MECl-1) (Macropain subunit MECl-1) (Multicatalytic endopeptidase complex subunit MECl-1). | |||||
|
HXB9_HUMAN
|
||||||
| NC score | 0.016086 (rank : 21) | θ value | 2.36792 (rank : 17) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P17482, Q9H1I1 | Gene names | HOXB9, HOX2E | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-B9 (Hox-2E) (Hox-2.5). | |||||
|
ZN447_HUMAN
|
||||||
| NC score | 0.015065 (rank : 22) | θ value | 0.0961366 (rank : 10) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 899 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TBC5, Q9BRK7, Q9H9A0 | Gene names | ZNF447 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 447. | |||||
|
HEMGN_MOUSE
|
||||||
| NC score | 0.014787 (rank : 23) | θ value | 8.99809 (rank : 33) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9ERZ0, Q3TAE6, Q80YT2, Q8CF29 | Gene names | Hemgn, Ndr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hemogen (Hemopoietic gene protein) (Negative differentiation regulator protein) (mNDR). | |||||
|
PURA1_MOUSE
|
||||||
| NC score | 0.014571 (rank : 24) | θ value | 5.27518 (rank : 24) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P28650, Q8CHQ1 | Gene names | Adssl1, Adss1 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylosuccinate synthetase isozyme 1 (EC 6.3.4.4) (Adenylosuccinate synthetase, muscle isozyme) (IMP--aspartate ligase 1) (AdSS 1) (AMPSase 1). | |||||
|
MILK2_HUMAN
|
||||||
| NC score | 0.011273 (rank : 25) | θ value | 4.03905 (rank : 22) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8IY33, Q7RTP4, Q7Z655, Q8TEQ4, Q9H5F9 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | MICAL-like protein 2. | |||||
|
ATS20_MOUSE
|
||||||
| NC score | 0.011230 (rank : 26) | θ value | 0.813845 (rank : 14) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P59511 | Gene names | Adamts20 | |||
|
Domain Architecture |

|
|||||
| Description | ADAMTS-20 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase with thrombospondin motifs 20) (ADAM-TS 20) (ADAM-TS20). | |||||
|
TNR27_MOUSE
|
||||||
| NC score | 0.011134 (rank : 27) | θ value | 8.99809 (rank : 35) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BX35, Q8BM50 | Gene names | Eda2r, Tnfrsf27, Xedar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor necrosis factor receptor superfamily member 27 (X-linked ectodysplasin-A2 receptor) (EDA-A2 receptor). | |||||
|
HD_HUMAN
|
||||||
| NC score | 0.010984 (rank : 28) | θ value | 8.99809 (rank : 32) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P42858, Q9UQB7 | Gene names | HD, IT15 | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin (Huntington disease protein) (HD protein). | |||||
|
ETV3_HUMAN
|
||||||
| NC score | 0.010773 (rank : 29) | θ value | 5.27518 (rank : 23) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P41162, Q8TAC8, Q9BX30 | Gene names | ETV3, METS, PE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
CSTFT_HUMAN
|
||||||
| NC score | 0.010285 (rank : 30) | θ value | 6.88961 (rank : 27) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H0L4, O75174, Q53HK6, Q7LGE8, Q8N6T1 | Gene names | CSTF2T, KIAA0689 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage stimulation factor 64 kDa subunit, tau variant (CSTF 64 kDa subunit, tau variant) (CF-1 64 kDa subunit, tau variant) (TauCstF-64). | |||||
|
CSTFT_MOUSE
|
||||||
| NC score | 0.009911 (rank : 31) | θ value | 6.88961 (rank : 28) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C7E9, Q6A016, Q8BHH7, Q8C3W7, Q8R2Y1, Q9EPU3 | Gene names | Cstf2t, Kiaa0689 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage stimulation factor 64 kDa subunit, tau variant (CSTF 64 kDa subunit, tau variant) (CF-1 64 kDa subunit, tau variant) (TauCstF-64). | |||||
|
RINI_HUMAN
|
||||||
| NC score | 0.008747 (rank : 32) | θ value | 6.88961 (rank : 31) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P13489 | Gene names | RNH1, PRI, RNH | |||
|
Domain Architecture |

|
|||||
| Description | Ribonuclease inhibitor (Ribonuclease/angiogenin inhibitor 1) (RAI) (Placental ribonuclease inhibitor) (RNase inhibitor) (RI). | |||||
|
B4GT1_MOUSE
|
||||||
| NC score | 0.008604 (rank : 33) | θ value | 6.88961 (rank : 26) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P15535 | Gene names | B4galt1, Ggtb, Ggtb2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-1,4-galactosyltransferase 1 (EC 2.4.1.-) (Beta-1,4-GalTase 1) (Beta4Gal-T1) (b4Gal-T1) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 1) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 1) [Includes: Lactose synthase A protein (EC 2.4.1.22); N-acetyllactosamine synthase (EC 2.4.1.90) (Nal synthetase); Beta-N-acetylglucosaminylglycopeptide beta-1,4- galactosyltransferase (EC 2.4.1.38); Beta-N-acetylglucosaminyl- glycolipid beta-1,4-galactosyltransferase (EC 2.4.1.-)]. | |||||
|
HDAC6_HUMAN
|
||||||
| NC score | 0.008565 (rank : 34) | θ value | 6.88961 (rank : 30) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UBN7 | Gene names | HDAC6 | |||
|
Domain Architecture |

|
|||||
| Description | Histone deacetylase 6 (HD6). | |||||
|
GP152_HUMAN
|
||||||
| NC score | 0.005739 (rank : 35) | θ value | 6.88961 (rank : 29) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 636 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8TDT2, Q86SM0 | Gene names | GPR152, PGR5 | |||
|
Domain Architecture |

|
|||||
| Description | Probable G-protein coupled receptor 152 (G-protein coupled receptor PGR5). | |||||
|
LGR5_MOUSE
|
||||||
| NC score | 0.002205 (rank : 36) | θ value | 8.99809 (rank : 34) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 484 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Z1P4 | Gene names | Lgr5, Fex, Gpr49 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeat-containing G-protein coupled receptor 5 precursor (G-protein coupled receptor 49) (Orphan G-protein coupled receptor FEX). | |||||