Please be patient as the page loads
|
ELK3_HUMAN
|
||||||
| SwissProt Accessions | P41970, Q6FG57, Q6GU29, Q9UD17 | Gene names | ELK3, NET, SAP2 | |||
|
Domain Architecture |
|
|||||
| Description | ETS domain-containing protein Elk-3 (ETS-related protein NET) (ETS- related protein ERP) (SRF accessory protein 2) (SAP-2). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
ELK3_HUMAN
|
||||||
| θ value | 2.67656e-153 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | P41970, Q6FG57, Q6GU29, Q9UD17 | Gene names | ELK3, NET, SAP2 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-3 (ETS-related protein NET) (ETS- related protein ERP) (SRF accessory protein 2) (SAP-2). | |||||
|
ELK3_MOUSE
|
||||||
| θ value | 3.38586e-148 (rank : 2) | NC score | 0.997678 (rank : 2) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P41971, P97747, Q62346 | Gene names | Elk3, Erp, Net | |||
|
Domain Architecture |
|
|||||
| Description | ETS domain-containing protein Elk-3 (ETS-related protein NET) (ETS- related protein ERP). | |||||
|
ELK4_HUMAN
|
||||||
| θ value | 7.19216e-82 (rank : 3) | NC score | 0.971380 (rank : 4) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P28324, P28323 | Gene names | ELK4, SAP1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-4 (Serum response factor accessory protein 1) (SAP-1). | |||||
|
ELK4_MOUSE
|
||||||
| θ value | 2.17053e-70 (rank : 4) | NC score | 0.979405 (rank : 3) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P41158 | Gene names | Elk4, Sap1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-4 (Serum response factor accessory protein 1) (SAP-1). | |||||
|
ELK1_MOUSE
|
||||||
| θ value | 2.50074e-42 (rank : 5) | NC score | 0.956722 (rank : 5) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P41969 | Gene names | Elk1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-1. | |||||
|
ELK1_HUMAN
|
||||||
| θ value | 1.2411e-41 (rank : 6) | NC score | 0.955805 (rank : 6) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P19419, O75606, O95058, Q969X8, Q9UJM4 | Gene names | ELK1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-1. | |||||
|
ETV4_HUMAN
|
||||||
| θ value | 1.62847e-25 (rank : 7) | NC score | 0.879472 (rank : 19) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P43268, Q96AW9 | Gene names | ETV4, E1AF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 4 (Adenovirus E1A enhancer-binding protein) (E1A-F). | |||||
|
ETV4_MOUSE
|
||||||
| θ value | 1.62847e-25 (rank : 8) | NC score | 0.880055 (rank : 18) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P28322 | Gene names | Etv4, Pea-3, Pea3 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 4 (Polyomavirus enhancer activator 3) (Protein PEA3). | |||||
|
ETV1_HUMAN
|
||||||
| θ value | 1.05554e-24 (rank : 9) | NC score | 0.877698 (rank : 21) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P50549, O75849, Q9UQ71, Q9Y636 | Gene names | ETV1, ER81 | |||
|
Domain Architecture |
|
|||||
| Description | ETS translocation variant 1 (ER81 protein). | |||||
|
ETV1_MOUSE
|
||||||
| θ value | 1.05554e-24 (rank : 10) | NC score | 0.878500 (rank : 20) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P41164 | Gene names | Etv1, Er81, Etsrp81 | |||
|
Domain Architecture |
|
|||||
| Description | ETS translocation variant 1 (ER81 protein). | |||||
|
ETV5_HUMAN
|
||||||
| θ value | 1.05554e-24 (rank : 11) | NC score | 0.871632 (rank : 24) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P41161 | Gene names | ETV5, ERM | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 5 (Ets-related protein ERM). | |||||
|
ETV5_MOUSE
|
||||||
| θ value | 2.35151e-24 (rank : 12) | NC score | 0.852120 (rank : 30) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9CXC9, Q3TG49, Q8C0F3, Q9JHB1 | Gene names | Etv5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 5. | |||||
|
FLI1_MOUSE
|
||||||
| θ value | 4.01107e-24 (rank : 13) | NC score | 0.885584 (rank : 15) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P26323 | Gene names | Fli1, Fli-1 | |||
|
Domain Architecture |
|
|||||
| Description | Friend leukemia integration 1 transcription factor (Retroviral integration site protein Fli-1). | |||||
|
FLI1_HUMAN
|
||||||
| θ value | 5.23862e-24 (rank : 14) | NC score | 0.883884 (rank : 16) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q01543, Q14319, Q92480, Q9UE07 | Gene names | FLI1 | |||
|
Domain Architecture |
|
|||||
| Description | Friend leukemia integration 1 transcription factor (Fli-1 proto- oncogene) (ERGB transcription factor). | |||||
|
ETS1_HUMAN
|
||||||
| θ value | 5.79196e-23 (rank : 15) | NC score | 0.893726 (rank : 11) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P14921, Q14278, Q16080 | Gene names | ETS1 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-1 protein (p54). | |||||
|
ETS1_MOUSE
|
||||||
| θ value | 5.79196e-23 (rank : 16) | NC score | 0.893600 (rank : 13) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P27577, Q61403 | Gene names | Ets1, Ets-1 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-1 protein (p54). | |||||
|
ETS2_HUMAN
|
||||||
| θ value | 7.56453e-23 (rank : 17) | NC score | 0.893853 (rank : 10) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P15036 | Gene names | ETS2 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-2 protein. | |||||
|
ETS2_MOUSE
|
||||||
| θ value | 7.56453e-23 (rank : 18) | NC score | 0.892512 (rank : 14) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P15037 | Gene names | Ets2 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-2 protein. | |||||
|
GABPA_HUMAN
|
||||||
| θ value | 3.75424e-22 (rank : 19) | NC score | 0.893680 (rank : 12) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q06546, Q12939 | Gene names | GABPA, E4TF1A | |||
|
Domain Architecture |
|
|||||
| Description | GA-binding protein alpha chain (GABP-subunit alpha) (Transcription factor E4TF1-60) (Nuclear respiratory factor 2 subunit alpha). | |||||
|
GABPA_MOUSE
|
||||||
| θ value | 3.75424e-22 (rank : 20) | NC score | 0.895153 (rank : 9) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q00422 | Gene names | Gabpa, E4tf1a | |||
|
Domain Architecture |
|
|||||
| Description | GA-binding protein alpha chain (GABP-subunit alpha). | |||||
|
ERF_HUMAN
|
||||||
| θ value | 4.58923e-20 (rank : 21) | NC score | 0.868943 (rank : 25) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P50548, Q9UPI7 | Gene names | ERF | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing transcription factor ERF (Ets2 repressor factor). | |||||
|
ERF_MOUSE
|
||||||
| θ value | 4.58923e-20 (rank : 22) | NC score | 0.868409 (rank : 26) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P70459 | Gene names | Erf | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing transcription factor ERF. | |||||
|
ERG_HUMAN
|
||||||
| θ value | 4.58923e-20 (rank : 23) | NC score | 0.876407 (rank : 22) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P11308, Q16113 | Gene names | ERG | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ERG (Transforming protein ERG). | |||||
|
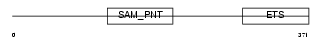
ERG_MOUSE
|
||||||
| θ value | 4.58923e-20 (rank : 24) | NC score | 0.875771 (rank : 23) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P81270, Q8C5L4, Q920K7, Q920K8, Q920K9 | Gene names | Erg, Erg-3 | |||
|
Domain Architecture |
|
|||||
| Description | Transcriptional regulator ERG. | |||||
|
ETV2_HUMAN
|
||||||
| θ value | 2.27762e-19 (rank : 25) | NC score | 0.906436 (rank : 7) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O00321, Q9UEA0 | Gene names | ETV2, ER71, ETSRP71 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 2 (Ets-related protein 71). | |||||
|
ETV3_HUMAN
|
||||||
| θ value | 2.27762e-19 (rank : 26) | NC score | 0.837334 (rank : 37) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P41162, Q8TAC8, Q9BX30 | Gene names | ETV3, METS, PE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
ETV3_MOUSE
|
||||||
| θ value | 2.27762e-19 (rank : 27) | NC score | 0.863251 (rank : 29) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8R4Z4, Q9QZW1 | Gene names | Etv3, Mets, Pe1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
ETV2_MOUSE
|
||||||
| θ value | 5.07402e-19 (rank : 28) | NC score | 0.903775 (rank : 8) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P41163 | Gene names | Etv2, Er71, Etsrp71 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 2 (Ets-related protein 71). | |||||
|
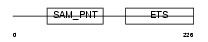
ETV7_HUMAN
|
||||||
| θ value | 9.56915e-18 (rank : 29) | NC score | 0.882976 (rank : 17) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9Y603, Q9NZ65, Q9NZ66, Q9NZ68, Q9NZR8, Q9UNJ7, Q9Y5K4, Q9Y604 | Gene names | ETV7, TEL2, TELB, TREF | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor ETV7 (Transcription factor Tel-2) (ETS-related protein Tel2) (Tel-related Ets factor). | |||||
|
ELF4_HUMAN
|
||||||
| θ value | 3.63628e-17 (rank : 30) | NC score | 0.839167 (rank : 35) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q99607, O60435 | Gene names | ELF4, ELFR, MEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-4 (E74-like factor 4) (Myeloid Elf-1-like factor). | |||||
|
ELF4_MOUSE
|
||||||
| θ value | 3.63628e-17 (rank : 31) | NC score | 0.817109 (rank : 41) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9Z2U4 | Gene names | Elf4, Mef | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-4 (E74-like factor 4) (Myeloid Elf-1-like factor). | |||||
|
ELF2_HUMAN
|
||||||
| θ value | 4.74913e-17 (rank : 32) | NC score | 0.843099 (rank : 33) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q15723, Q15724, Q15725, Q6P1K5 | Gene names | ELF2, NERF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-2 (E74-like factor 2) (New ETS- related factor). | |||||
|
ELF1_HUMAN
|
||||||
| θ value | 6.20254e-17 (rank : 33) | NC score | 0.837569 (rank : 36) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P32519, Q9UDE1 | Gene names | ELF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-1 (E74-like factor 1). | |||||
|
ELF2_MOUSE
|
||||||
| θ value | 6.20254e-17 (rank : 34) | NC score | 0.843949 (rank : 32) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9JHC9, Q6NST2, Q8BTX8, Q9JHC7, Q9JHC8, Q9JHD0 | Gene names | Elf2, Nerf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-2 (E74-like factor 2) (New ETS- related factor). | |||||
|
ETV6_MOUSE
|
||||||
| θ value | 8.10077e-17 (rank : 35) | NC score | 0.844233 (rank : 31) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P97360 | Gene names | Etv6, Tel, Tel1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor ETV6 (ETS-related protein Tel1) (Tel) (ETS translocation variant 6). | |||||
|
ETV6_HUMAN
|
||||||
| θ value | 2.35696e-16 (rank : 36) | NC score | 0.840539 (rank : 34) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P41212, Q9UMF6 | Gene names | ETV6, TEL, TEL1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor ETV6 (ETS-related protein Tel1) (Tel) (ETS translocation variant 6). | |||||
|
ELF1_MOUSE
|
||||||
| θ value | 3.07829e-16 (rank : 37) | NC score | 0.830196 (rank : 38) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q60775 | Gene names | Elf1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS-related transcription factor Elf-1 (E74-like factor 1). | |||||
|
SPDEF_HUMAN
|
||||||
| θ value | 5.8054e-15 (rank : 38) | NC score | 0.867767 (rank : 27) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O95238 | Gene names | SPDEF, PDEF, PSE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SAM pointed domain-containing Ets transcription factor (Prostate- derived Ets factor) (Prostate epithelium-specific Ets transcription factor) (Prostate-specific Ets). | |||||
|
SPDEF_MOUSE
|
||||||
| θ value | 7.58209e-15 (rank : 39) | NC score | 0.865603 (rank : 28) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9WTP3 | Gene names | Spdef, Pdef, Pse | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SAM pointed domain-containing Ets transcription factor (Prostate- derived Ets factor) (Prostate epithelium-specific Ets transcription factor) (Prostate-specific Ets). | |||||
|
ELF5_HUMAN
|
||||||
| θ value | 1.02475e-11 (rank : 40) | NC score | 0.823232 (rank : 40) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9UKW6, O95175, Q8N2K9, Q96QY3, Q9UKW5 | Gene names | ELF5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-5 (E74-like factor 5) (Epithelium-specific Ets transcription factor 2) (ESE-2) (Epithelium- restricted ESE-1-related Ets factor). | |||||
|
ELF5_MOUSE
|
||||||
| θ value | 1.02475e-11 (rank : 41) | NC score | 0.824152 (rank : 39) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8VDK3, Q921H5, Q9Z2K6 | Gene names | Elf5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-5 (E74-like factor 5). | |||||
|
SPIB_HUMAN
|
||||||
| θ value | 0.00035302 (rank : 42) | NC score | 0.612981 (rank : 44) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q01892, Q15359 | Gene names | SPIB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor Spi-B. | |||||
|
SPI1_HUMAN
|
||||||
| θ value | 0.000602161 (rank : 43) | NC score | 0.621906 (rank : 43) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P17947 | Gene names | SPI1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor PU.1 (31 kDa transforming protein). | |||||
|
SPI1_MOUSE
|
||||||
| θ value | 0.000602161 (rank : 44) | NC score | 0.623219 (rank : 42) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P17433, Q99L57 | Gene names | Spi1, Sfpi-1, Sfpi1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor PU.1 (31 kDa transforming protein) (SFFV proviral integration 1 protein). | |||||
|
SPIC_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 45) | NC score | 0.573763 (rank : 46) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8N5J4 | Gene names | SPIC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor Spi-C. | |||||
|
SPIC_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 46) | NC score | 0.539531 (rank : 47) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q6P3D7, Q9Z0Y0 | Gene names | Spic | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor Spi-C (Pu.1-related factor) (Prf). | |||||
|
SPIB_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 47) | NC score | 0.592320 (rank : 45) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O35906, O35907, O35909, O55199 | Gene names | Spib | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor Spi-B. | |||||
|
PTN22_MOUSE
|
||||||
| θ value | 0.125558 (rank : 48) | NC score | 0.013505 (rank : 54) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 179 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P29352, Q7TMP9 | Gene names | Ptpn22, Ptpn8 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 22 (EC 3.1.3.48) (Hematopoietic cell protein-tyrosine phosphatase 70Z-PEP). | |||||
|
ACD_HUMAN
|
||||||
| θ value | 1.38821 (rank : 49) | NC score | 0.032938 (rank : 48) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96AP0, Q562H5, Q9H8F9 | Gene names | ACD, PIP1, PTOP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Adrenocortical dysplasia protein homolog (POT1 and TIN2-interacting protein). | |||||
|
BCAS1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 50) | NC score | 0.029108 (rank : 49) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80YN3, Q9CVA1 | Gene names | Bcas1, Nabc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Breast carcinoma amplified sequence 1 homolog (Novel amplified in breast cancer 1 homolog). | |||||
|
UROL1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 51) | NC score | 0.015017 (rank : 52) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5DID3, Q5DID1, Q5DID2 | Gene names | Umodl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uromodulin-like 1 precursor (Olfactorin). | |||||
|
CRSP2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 52) | NC score | 0.017769 (rank : 51) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O60244, Q9UNB3 | Gene names | CRSP2, ARC150, DRIP150, EXLM1, TRAP170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CRSP complex subunit 2 (Cofactor required for Sp1 transcriptional activation subunit 2) (Transcriptional coactivator CRSP150) (Vitamin D3 receptor-interacting protein complex 150 kDa component) (DRIP150) (Thyroid hormone receptor-associated protein complex 170 kDa component) (Trap170) (Activator-recruited cofactor 150 kDa component) (ARC150). | |||||
|
HEY2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 53) | NC score | 0.014468 (rank : 53) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QUS4, Q3TZ99, Q8CD44 | Gene names | Hey2, Chf1, Herp, Herp1, Hesr2, Hrt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES-related repressor protein 2). | |||||
|
ALEX_HUMAN
|
||||||
| θ value | 5.27518 (rank : 54) | NC score | 0.023574 (rank : 50) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P84996 | Gene names | GNAS, GNAS1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ALEX (Alternative gene product encoded by XL-exon). | |||||
|
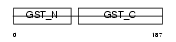
GSTA4_MOUSE
|
||||||
| θ value | 5.27518 (rank : 55) | NC score | 0.005166 (rank : 57) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P24472, Q9CQ81, Q9CTY7, Q9CY87, Q9D038 | Gene names | Gsta4, Gsta | |||
|
Domain Architecture |
|
|||||
| Description | Glutathione S-transferase A4 (EC 2.5.1.18) (GST A4-4) (GSTA4-4) (Glutathione S-transferase 5.7) (GST 5.7) (GST class-alpha member 4). | |||||
|
NFAC1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 56) | NC score | 0.008666 (rank : 56) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O88942, O70345 | Gene names | Nfatc1, Nfat2, Nfatc | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor of activated T-cells, cytoplasmic 1 (NFAT transcription complex cytosolic component) (NF-ATc1) (NF-ATc). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 57) | NC score | 0.002083 (rank : 61) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
CD45_MOUSE
|
||||||
| θ value | 6.88961 (rank : 58) | NC score | 0.004917 (rank : 58) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P06800, Q61812, Q61813, Q61814, Q61815, Q78EF1 | Gene names | Ptprc, Ly-5 | |||
|
Domain Architecture |

|
|||||
| Description | Leukocyte common antigen precursor (EC 3.1.3.48) (L-CA) (Lymphocyte common antigen Ly-5) (CD45 antigen) (T200). | |||||
|
CTGE4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 59) | NC score | 0.003648 (rank : 60) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 393 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8IX94, O95046 | Gene names | CTAGE4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cutaneous T-cell lymphoma-associated antigen 4 (cTAGE-4 protein). | |||||
|
LCP2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 60) | NC score | 0.004641 (rank : 59) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 531 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q13094 | Gene names | LCP2 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphocyte cytosolic protein 2 (SH2 domain-containing leukocyte protein of 76 kDa) (SLP-76 tyrosine phosphoprotein) (SLP76). | |||||
|
BCL9_HUMAN
|
||||||
| θ value | 8.99809 (rank : 61) | NC score | 0.011207 (rank : 55) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1008 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O00512 | Gene names | BCL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9) (Legless homolog). | |||||
|
CSPG2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 62) | NC score | 0.000618 (rank : 63) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 711 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q62059, Q62058, Q9CUU0 | Gene names | Cspg2 | |||
|
Domain Architecture |

|
|||||
| Description | Versican core protein precursor (Large fibroblast proteoglycan) (Chondroitin sulfate proteoglycan core protein 2) (PG-M). | |||||
|
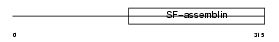
DMN_HUMAN
|
||||||
| θ value | 8.99809 (rank : 63) | NC score | -0.000161 (rank : 64) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O15061 | Gene names | DMN, KIAA0353 | |||
|
Domain Architecture |
|
|||||
| Description | Desmuslin. | |||||
|
SOX6_MOUSE
|
||||||
| θ value | 8.99809 (rank : 64) | NC score | 0.001159 (rank : 62) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P40645, Q62250, Q9QWS5 | Gene names | Sox6, Sox-6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-6 (SOX-LZ). | |||||
|
ELK3_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 2.67656e-153 (rank : 1) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | P41970, Q6FG57, Q6GU29, Q9UD17 | Gene names | ELK3, NET, SAP2 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-3 (ETS-related protein NET) (ETS- related protein ERP) (SRF accessory protein 2) (SAP-2). | |||||
|
ELK3_MOUSE
|
||||||
| NC score | 0.997678 (rank : 2) | θ value | 3.38586e-148 (rank : 2) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P41971, P97747, Q62346 | Gene names | Elk3, Erp, Net | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-3 (ETS-related protein NET) (ETS- related protein ERP). | |||||
|
ELK4_MOUSE
|
||||||
| NC score | 0.979405 (rank : 3) | θ value | 2.17053e-70 (rank : 4) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P41158 | Gene names | Elk4, Sap1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-4 (Serum response factor accessory protein 1) (SAP-1). | |||||
|
ELK4_HUMAN
|
||||||
| NC score | 0.971380 (rank : 4) | θ value | 7.19216e-82 (rank : 3) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P28324, P28323 | Gene names | ELK4, SAP1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-4 (Serum response factor accessory protein 1) (SAP-1). | |||||
|
ELK1_MOUSE
|
||||||
| NC score | 0.956722 (rank : 5) | θ value | 2.50074e-42 (rank : 5) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P41969 | Gene names | Elk1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-1. | |||||
|
ELK1_HUMAN
|
||||||
| NC score | 0.955805 (rank : 6) | θ value | 1.2411e-41 (rank : 6) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P19419, O75606, O95058, Q969X8, Q9UJM4 | Gene names | ELK1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-1. | |||||
|
ETV2_HUMAN
|
||||||
| NC score | 0.906436 (rank : 7) | θ value | 2.27762e-19 (rank : 25) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O00321, Q9UEA0 | Gene names | ETV2, ER71, ETSRP71 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 2 (Ets-related protein 71). | |||||
|
ETV2_MOUSE
|
||||||
| NC score | 0.903775 (rank : 8) | θ value | 5.07402e-19 (rank : 28) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P41163 | Gene names | Etv2, Er71, Etsrp71 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 2 (Ets-related protein 71). | |||||
|
GABPA_MOUSE
|
||||||
| NC score | 0.895153 (rank : 9) | θ value | 3.75424e-22 (rank : 20) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q00422 | Gene names | Gabpa, E4tf1a | |||
|
Domain Architecture |

|
|||||
| Description | GA-binding protein alpha chain (GABP-subunit alpha). | |||||
|
ETS2_HUMAN
|
||||||
| NC score | 0.893853 (rank : 10) | θ value | 7.56453e-23 (rank : 17) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P15036 | Gene names | ETS2 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-2 protein. | |||||
|
ETS1_HUMAN
|
||||||
| NC score | 0.893726 (rank : 11) | θ value | 5.79196e-23 (rank : 15) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P14921, Q14278, Q16080 | Gene names | ETS1 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-1 protein (p54). | |||||
|
GABPA_HUMAN
|
||||||
| NC score | 0.893680 (rank : 12) | θ value | 3.75424e-22 (rank : 19) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q06546, Q12939 | Gene names | GABPA, E4TF1A | |||
|
Domain Architecture |

|
|||||
| Description | GA-binding protein alpha chain (GABP-subunit alpha) (Transcription factor E4TF1-60) (Nuclear respiratory factor 2 subunit alpha). | |||||
|
ETS1_MOUSE
|
||||||
| NC score | 0.893600 (rank : 13) | θ value | 5.79196e-23 (rank : 16) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P27577, Q61403 | Gene names | Ets1, Ets-1 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-1 protein (p54). | |||||
|
ETS2_MOUSE
|
||||||
| NC score | 0.892512 (rank : 14) | θ value | 7.56453e-23 (rank : 18) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P15037 | Gene names | Ets2 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-2 protein. | |||||
|
FLI1_MOUSE
|
||||||
| NC score | 0.885584 (rank : 15) | θ value | 4.01107e-24 (rank : 13) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P26323 | Gene names | Fli1, Fli-1 | |||
|
Domain Architecture |

|
|||||
| Description | Friend leukemia integration 1 transcription factor (Retroviral integration site protein Fli-1). | |||||
|
FLI1_HUMAN
|
||||||
| NC score | 0.883884 (rank : 16) | θ value | 5.23862e-24 (rank : 14) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q01543, Q14319, Q92480, Q9UE07 | Gene names | FLI1 | |||
|
Domain Architecture |

|
|||||
| Description | Friend leukemia integration 1 transcription factor (Fli-1 proto- oncogene) (ERGB transcription factor). | |||||
|
ETV7_HUMAN
|
||||||
| NC score | 0.882976 (rank : 17) | θ value | 9.56915e-18 (rank : 29) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9Y603, Q9NZ65, Q9NZ66, Q9NZ68, Q9NZR8, Q9UNJ7, Q9Y5K4, Q9Y604 | Gene names | ETV7, TEL2, TELB, TREF | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor ETV7 (Transcription factor Tel-2) (ETS-related protein Tel2) (Tel-related Ets factor). | |||||
|
ETV4_MOUSE
|
||||||
| NC score | 0.880055 (rank : 18) | θ value | 1.62847e-25 (rank : 8) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P28322 | Gene names | Etv4, Pea-3, Pea3 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 4 (Polyomavirus enhancer activator 3) (Protein PEA3). | |||||
|
ETV4_HUMAN
|
||||||
| NC score | 0.879472 (rank : 19) | θ value | 1.62847e-25 (rank : 7) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P43268, Q96AW9 | Gene names | ETV4, E1AF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 4 (Adenovirus E1A enhancer-binding protein) (E1A-F). | |||||
|
ETV1_MOUSE
|
||||||
| NC score | 0.878500 (rank : 20) | θ value | 1.05554e-24 (rank : 10) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P41164 | Gene names | Etv1, Er81, Etsrp81 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 1 (ER81 protein). | |||||
|
ETV1_HUMAN
|
||||||
| NC score | 0.877698 (rank : 21) | θ value | 1.05554e-24 (rank : 9) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P50549, O75849, Q9UQ71, Q9Y636 | Gene names | ETV1, ER81 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 1 (ER81 protein). | |||||
|
ERG_HUMAN
|
||||||
| NC score | 0.876407 (rank : 22) | θ value | 4.58923e-20 (rank : 23) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P11308, Q16113 | Gene names | ERG | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ERG (Transforming protein ERG). | |||||
|
ERG_MOUSE
|
||||||
| NC score | 0.875771 (rank : 23) | θ value | 4.58923e-20 (rank : 24) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P81270, Q8C5L4, Q920K7, Q920K8, Q920K9 | Gene names | Erg, Erg-3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ERG. | |||||
|
ETV5_HUMAN
|
||||||
| NC score | 0.871632 (rank : 24) | θ value | 1.05554e-24 (rank : 11) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P41161 | Gene names | ETV5, ERM | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 5 (Ets-related protein ERM). | |||||
|
ERF_HUMAN
|
||||||
| NC score | 0.868943 (rank : 25) | θ value | 4.58923e-20 (rank : 21) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P50548, Q9UPI7 | Gene names | ERF | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing transcription factor ERF (Ets2 repressor factor). | |||||
|
ERF_MOUSE
|
||||||
| NC score | 0.868409 (rank : 26) | θ value | 4.58923e-20 (rank : 22) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P70459 | Gene names | Erf | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing transcription factor ERF. | |||||
|
SPDEF_HUMAN
|
||||||
| NC score | 0.867767 (rank : 27) | θ value | 5.8054e-15 (rank : 38) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O95238 | Gene names | SPDEF, PDEF, PSE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SAM pointed domain-containing Ets transcription factor (Prostate- derived Ets factor) (Prostate epithelium-specific Ets transcription factor) (Prostate-specific Ets). | |||||
|
SPDEF_MOUSE
|
||||||
| NC score | 0.865603 (rank : 28) | θ value | 7.58209e-15 (rank : 39) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9WTP3 | Gene names | Spdef, Pdef, Pse | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SAM pointed domain-containing Ets transcription factor (Prostate- derived Ets factor) (Prostate epithelium-specific Ets transcription factor) (Prostate-specific Ets). | |||||
|
ETV3_MOUSE
|
||||||
| NC score | 0.863251 (rank : 29) | θ value | 2.27762e-19 (rank : 27) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8R4Z4, Q9QZW1 | Gene names | Etv3, Mets, Pe1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
ETV5_MOUSE
|
||||||
| NC score | 0.852120 (rank : 30) | θ value | 2.35151e-24 (rank : 12) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9CXC9, Q3TG49, Q8C0F3, Q9JHB1 | Gene names | Etv5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 5. | |||||
|
ETV6_MOUSE
|
||||||
| NC score | 0.844233 (rank : 31) | θ value | 8.10077e-17 (rank : 35) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P97360 | Gene names | Etv6, Tel, Tel1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor ETV6 (ETS-related protein Tel1) (Tel) (ETS translocation variant 6). | |||||
|
ELF2_MOUSE
|
||||||
| NC score | 0.843949 (rank : 32) | θ value | 6.20254e-17 (rank : 34) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9JHC9, Q6NST2, Q8BTX8, Q9JHC7, Q9JHC8, Q9JHD0 | Gene names | Elf2, Nerf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-2 (E74-like factor 2) (New ETS- related factor). | |||||
|
ELF2_HUMAN
|
||||||
| NC score | 0.843099 (rank : 33) | θ value | 4.74913e-17 (rank : 32) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q15723, Q15724, Q15725, Q6P1K5 | Gene names | ELF2, NERF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-2 (E74-like factor 2) (New ETS- related factor). | |||||
|
ETV6_HUMAN
|
||||||
| NC score | 0.840539 (rank : 34) | θ value | 2.35696e-16 (rank : 36) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P41212, Q9UMF6 | Gene names | ETV6, TEL, TEL1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor ETV6 (ETS-related protein Tel1) (Tel) (ETS translocation variant 6). | |||||
|
ELF4_HUMAN
|
||||||
| NC score | 0.839167 (rank : 35) | θ value | 3.63628e-17 (rank : 30) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q99607, O60435 | Gene names | ELF4, ELFR, MEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-4 (E74-like factor 4) (Myeloid Elf-1-like factor). | |||||
|
ELF1_HUMAN
|
||||||
| NC score | 0.837569 (rank : 36) | θ value | 6.20254e-17 (rank : 33) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P32519, Q9UDE1 | Gene names | ELF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-1 (E74-like factor 1). | |||||
|
ETV3_HUMAN
|
||||||
| NC score | 0.837334 (rank : 37) | θ value | 2.27762e-19 (rank : 26) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P41162, Q8TAC8, Q9BX30 | Gene names | ETV3, METS, PE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
ELF1_MOUSE
|
||||||
| NC score | 0.830196 (rank : 38) | θ value | 3.07829e-16 (rank : 37) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q60775 | Gene names | Elf1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS-related transcription factor Elf-1 (E74-like factor 1). | |||||
|
ELF5_MOUSE
|
||||||
| NC score | 0.824152 (rank : 39) | θ value | 1.02475e-11 (rank : 41) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8VDK3, Q921H5, Q9Z2K6 | Gene names | Elf5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-5 (E74-like factor 5). | |||||
|
ELF5_HUMAN
|
||||||
| NC score | 0.823232 (rank : 40) | θ value | 1.02475e-11 (rank : 40) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9UKW6, O95175, Q8N2K9, Q96QY3, Q9UKW5 | Gene names | ELF5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-5 (E74-like factor 5) (Epithelium-specific Ets transcription factor 2) (ESE-2) (Epithelium- restricted ESE-1-related Ets factor). | |||||
|
ELF4_MOUSE
|
||||||
| NC score | 0.817109 (rank : 41) | θ value | 3.63628e-17 (rank : 31) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9Z2U4 | Gene names | Elf4, Mef | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-4 (E74-like factor 4) (Myeloid Elf-1-like factor). | |||||
|
SPI1_MOUSE
|
||||||
| NC score | 0.623219 (rank : 42) | θ value | 0.000602161 (rank : 44) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P17433, Q99L57 | Gene names | Spi1, Sfpi-1, Sfpi1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor PU.1 (31 kDa transforming protein) (SFFV proviral integration 1 protein). | |||||
|
SPI1_HUMAN
|
||||||
| NC score | 0.621906 (rank : 43) | θ value | 0.000602161 (rank : 43) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P17947 | Gene names | SPI1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor PU.1 (31 kDa transforming protein). | |||||
|
SPIB_HUMAN
|
||||||
| NC score | 0.612981 (rank : 44) | θ value | 0.00035302 (rank : 42) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q01892, Q15359 | Gene names | SPIB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor Spi-B. | |||||
|
SPIB_MOUSE
|
||||||
| NC score | 0.592320 (rank : 45) | θ value | 0.0431538 (rank : 47) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O35906, O35907, O35909, O55199 | Gene names | Spib | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor Spi-B. | |||||
|
SPIC_HUMAN
|
||||||
| NC score | 0.573763 (rank : 46) | θ value | 0.00509761 (rank : 45) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8N5J4 | Gene names | SPIC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor Spi-C. | |||||
|
SPIC_MOUSE
|
||||||
| NC score | 0.539531 (rank : 47) | θ value | 0.0252991 (rank : 46) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q6P3D7, Q9Z0Y0 | Gene names | Spic | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor Spi-C (Pu.1-related factor) (Prf). | |||||
|
ACD_HUMAN
|
||||||
| NC score | 0.032938 (rank : 48) | θ value | 1.38821 (rank : 49) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96AP0, Q562H5, Q9H8F9 | Gene names | ACD, PIP1, PTOP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Adrenocortical dysplasia protein homolog (POT1 and TIN2-interacting protein). | |||||
|
BCAS1_MOUSE
|
||||||
| NC score | 0.029108 (rank : 49) | θ value | 2.36792 (rank : 50) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80YN3, Q9CVA1 | Gene names | Bcas1, Nabc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Breast carcinoma amplified sequence 1 homolog (Novel amplified in breast cancer 1 homolog). | |||||
|
ALEX_HUMAN
|
||||||
| NC score | 0.023574 (rank : 50) | θ value | 5.27518 (rank : 54) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P84996 | Gene names | GNAS, GNAS1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ALEX (Alternative gene product encoded by XL-exon). | |||||
|
CRSP2_HUMAN
|
||||||
| NC score | 0.017769 (rank : 51) | θ value | 3.0926 (rank : 52) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O60244, Q9UNB3 | Gene names | CRSP2, ARC150, DRIP150, EXLM1, TRAP170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CRSP complex subunit 2 (Cofactor required for Sp1 transcriptional activation subunit 2) (Transcriptional coactivator CRSP150) (Vitamin D3 receptor-interacting protein complex 150 kDa component) (DRIP150) (Thyroid hormone receptor-associated protein complex 170 kDa component) (Trap170) (Activator-recruited cofactor 150 kDa component) (ARC150). | |||||
|
UROL1_MOUSE
|
||||||
| NC score | 0.015017 (rank : 52) | θ value | 2.36792 (rank : 51) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5DID3, Q5DID1, Q5DID2 | Gene names | Umodl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uromodulin-like 1 precursor (Olfactorin). | |||||
|
HEY2_MOUSE
|
||||||
| NC score | 0.014468 (rank : 53) | θ value | 4.03905 (rank : 53) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QUS4, Q3TZ99, Q8CD44 | Gene names | Hey2, Chf1, Herp, Herp1, Hesr2, Hrt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES-related repressor protein 2). | |||||
|
PTN22_MOUSE
|
||||||
| NC score | 0.013505 (rank : 54) | θ value | 0.125558 (rank : 48) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 179 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P29352, Q7TMP9 | Gene names | Ptpn22, Ptpn8 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 22 (EC 3.1.3.48) (Hematopoietic cell protein-tyrosine phosphatase 70Z-PEP). | |||||
|
BCL9_HUMAN
|
||||||
| NC score | 0.011207 (rank : 55) | θ value | 8.99809 (rank : 61) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1008 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O00512 | Gene names | BCL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9) (Legless homolog). | |||||
|
NFAC1_MOUSE
|
||||||
| NC score | 0.008666 (rank : 56) | θ value | 5.27518 (rank : 56) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O88942, O70345 | Gene names | Nfatc1, Nfat2, Nfatc | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor of activated T-cells, cytoplasmic 1 (NFAT transcription complex cytosolic component) (NF-ATc1) (NF-ATc). | |||||
|
GSTA4_MOUSE
|
||||||
| NC score | 0.005166 (rank : 57) | θ value | 5.27518 (rank : 55) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P24472, Q9CQ81, Q9CTY7, Q9CY87, Q9D038 | Gene names | Gsta4, Gsta | |||
|
Domain Architecture |

|
|||||
| Description | Glutathione S-transferase A4 (EC 2.5.1.18) (GST A4-4) (GSTA4-4) (Glutathione S-transferase 5.7) (GST 5.7) (GST class-alpha member 4). | |||||
|
CD45_MOUSE
|
||||||
| NC score | 0.004917 (rank : 58) | θ value | 6.88961 (rank : 58) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P06800, Q61812, Q61813, Q61814, Q61815, Q78EF1 | Gene names | Ptprc, Ly-5 | |||
|
Domain Architecture |

|
|||||
| Description | Leukocyte common antigen precursor (EC 3.1.3.48) (L-CA) (Lymphocyte common antigen Ly-5) (CD45 antigen) (T200). | |||||
|
LCP2_HUMAN
|
||||||
| NC score | 0.004641 (rank : 59) | θ value | 6.88961 (rank : 60) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 531 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q13094 | Gene names | LCP2 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphocyte cytosolic protein 2 (SH2 domain-containing leukocyte protein of 76 kDa) (SLP-76 tyrosine phosphoprotein) (SLP76). | |||||
|
CTGE4_HUMAN
|
||||||
| NC score | 0.003648 (rank : 60) | θ value | 6.88961 (rank : 59) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 393 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8IX94, O95046 | Gene names | CTAGE4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cutaneous T-cell lymphoma-associated antigen 4 (cTAGE-4 protein). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.002083 (rank : 61) | θ value | 5.27518 (rank : 57) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
SOX6_MOUSE
|
||||||
| NC score | 0.001159 (rank : 62) | θ value | 8.99809 (rank : 64) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P40645, Q62250, Q9QWS5 | Gene names | Sox6, Sox-6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-6 (SOX-LZ). | |||||
|
CSPG2_MOUSE
|
||||||
| NC score | 0.000618 (rank : 63) | θ value | 8.99809 (rank : 62) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 711 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q62059, Q62058, Q9CUU0 | Gene names | Cspg2 | |||
|
Domain Architecture |

|
|||||
| Description | Versican core protein precursor (Large fibroblast proteoglycan) (Chondroitin sulfate proteoglycan core protein 2) (PG-M). | |||||
|
DMN_HUMAN
|
||||||
| NC score | -0.000161 (rank : 64) | θ value | 8.99809 (rank : 63) | |||
| Query Neighborhood Hits | 64 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O15061 | Gene names | DMN, KIAA0353 | |||
|
Domain Architecture |

|
|||||
| Description | Desmuslin. | |||||