Please be patient as the page loads
|
EGLN2_HUMAN
|
||||||
| SwissProt Accessions | Q96KS0, Q8WWY4, Q9BV14 | Gene names | EGLN2, EIT6 | |||
|
Domain Architecture |

|
|||||
| Description | Egl nine homolog 2 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl hydroxylase 1) (HIF-prolyl hydroxylase 1) (HIF-PH1) (HPH-3) (Prolyl hydroxylase domain-containing protein 1) (PHD1) (Estrogen-induced tag 6). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
EGLN2_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q96KS0, Q8WWY4, Q9BV14 | Gene names | EGLN2, EIT6 | |||
|
Domain Architecture |

|
|||||
| Description | Egl nine homolog 2 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl hydroxylase 1) (HIF-prolyl hydroxylase 1) (HIF-PH1) (HPH-3) (Prolyl hydroxylase domain-containing protein 1) (PHD1) (Estrogen-induced tag 6). | |||||
|
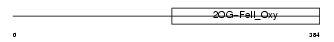
EGLN2_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.996205 (rank : 2) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q91YE2, Q8C6I4, Q8CIL9, Q8VHJ1, Q99MI0 | Gene names | Egln2 | |||
|
Domain Architecture |
|
|||||
| Description | Egl nine homolog 2 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl hydroxylase 1) (HIF-prolyl hydroxylase 1) (HIF-PH1) (Falkor). | |||||
|
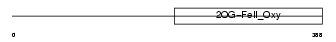
EGLN1_HUMAN
|
||||||
| θ value | 1.88829e-90 (rank : 3) | NC score | 0.938085 (rank : 5) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9GZT9, Q8N3M8, Q9BZS8, Q9BZT0 | Gene names | EGLN1, C1orf12 | |||
|
Domain Architecture |
|
|||||
| Description | Egl nine homolog 1 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl hydroxylase 2) (HIF-prolyl hydroxylase 2) (HIF-PH2) (HPH-2) (Prolyl hydroxylase domain-containing protein 2) (PHD2) (SM-20). | |||||
|
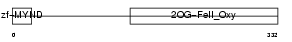
EGLN1_MOUSE
|
||||||
| θ value | 9.37149e-90 (rank : 4) | NC score | 0.926869 (rank : 6) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q91YE3, Q8VHJ2, Q922P3 | Gene names | Egln1 | |||
|
Domain Architecture |
|
|||||
| Description | Egl nine homolog 1 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl hydroxylase 2) (HIF-prolyl hydroxylase 2) (HIF-PH2) (HPH-2) (SM-20). | |||||
|
EGLN3_MOUSE
|
||||||
| θ value | 2.73302e-81 (rank : 5) | NC score | 0.976462 (rank : 3) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q91UZ4, Q3TCG8, Q8C8M6, Q8CCA8, Q8R5C7 | Gene names | Egln3 | |||
|
Domain Architecture |

|
|||||
| Description | Egl nine homolog 3 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl hydroxylase 3) (HIF-prolyl hydroxylase 3) (HIF-PH3) (SM-20). | |||||
|
EGLN3_HUMAN
|
||||||
| θ value | 6.08852e-81 (rank : 6) | NC score | 0.976252 (rank : 4) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9H6Z9 | Gene names | EGLN3 | |||
|
Domain Architecture |

|
|||||
| Description | Egl nine homolog 3 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl hydroxylase 3) (HIF-prolyl hydroxylase 3) (HIF-PH3) (HPH-1) (Prolyl hydroxylase domain-containing protein 3) (PHD3). | |||||
|
K1543_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 7) | NC score | 0.062358 (rank : 18) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9P1Y5, Q8NDF1 | Gene names | KIAA1543 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1543. | |||||
|
TTBK1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 8) | NC score | 0.009496 (rank : 23) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 1058 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5TCY1, Q2L6C6, Q8N444, Q96JH2 | Gene names | TTBK1, BDTK, KIAA1855 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tau-tubulin kinase 1 (EC 2.7.11.1) (Brain-derived tau kinase). | |||||
|
P3H1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 9) | NC score | 0.076681 (rank : 9) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q32P28, Q7KZR4, Q96BR8, Q96SK8, Q96SL5, Q96SN3, Q9H6K3, Q9HC86, Q9HC87 | Gene names | LEPRE1, GROS1, P3H1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 1 precursor (EC 1.14.11.7) (Leucine- and proline- enriched proteoglycan 1) (Leprecan-1) (Growth suppressor 1). | |||||
|
P3H2_MOUSE
|
||||||
| θ value | 0.813845 (rank : 10) | NC score | 0.079510 (rank : 8) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8CG71, Q8C673 | Gene names | Leprel1, P3h2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 2 precursor (EC 1.14.11.7) (Leprecan-like protein 1). | |||||
|
P3H1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 11) | NC score | 0.080928 (rank : 7) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q3V1T4, Q3TWX8, Q8BSV2, Q8CFL3, Q9CWK5, Q9QZT6, Q9QZT7 | Gene names | Lepre1, Gros1, P3h1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 1 precursor (EC 1.14.11.7) (Leucine- and proline- enriched proteoglycan 1) (Leprecan-1) (Growth suppressor 1). | |||||
|
RGS3_MOUSE
|
||||||
| θ value | 1.38821 (rank : 12) | NC score | 0.011024 (rank : 22) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
P3H2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 13) | NC score | 0.064079 (rank : 14) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8IVL5, Q9NVI2 | Gene names | LEPREL1, MLAT4, P3H2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 2 precursor (EC 1.14.11.7) (Leprecan-like protein 1) (Myxoid liposarcoma-associated protein 4). | |||||
|
DUT_HUMAN
|
||||||
| θ value | 3.0926 (rank : 14) | NC score | 0.032171 (rank : 19) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P33316, O14785, Q16708, Q16860, Q96Q81 | Gene names | DUT | |||
|
Domain Architecture |

|
|||||
| Description | Deoxyuridine 5'-triphosphate nucleotidohydrolase, mitochondrial precursor (EC 3.6.1.23) (dUTPase) (dUTP pyrophosphatase). | |||||
|
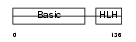
MYF6_HUMAN
|
||||||
| θ value | 4.03905 (rank : 15) | NC score | 0.014301 (rank : 20) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P23409, Q53X80, Q6FHI9 | Gene names | MYF6 | |||
|
Domain Architecture |
|
|||||
| Description | Myogenic factor 6 (Myf-6). | |||||
|
EPHB6_HUMAN
|
||||||
| θ value | 5.27518 (rank : 16) | NC score | 0.000855 (rank : 28) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 912 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O15197 | Gene names | EPHB6 | |||
|
Domain Architecture |

|
|||||
| Description | Ephrin type-B receptor 6 precursor (Tyrosine-protein kinase-defective receptor EPH-6) (HEP). | |||||
|
FA38A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 17) | NC score | 0.011466 (rank : 21) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q92508 | Gene names | FAM38A, KIAA0233 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM38A. | |||||
|
FGD3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 18) | NC score | 0.007988 (rank : 24) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5JSP0, Q7Z7D9, Q8N5G1 | Gene names | FGD3, ZFYVE5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 3 (Zinc finger FYVE domain-containing protein 5). | |||||
|
FXL19_MOUSE
|
||||||
| θ value | 8.99809 (rank : 19) | NC score | 0.007058 (rank : 26) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6PB97, Q7TSH0, Q8BIB4 | Gene names | Fbxl19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 19 (F-box and leucine-rich repeat protein 19). | |||||
|
MYPN_MOUSE
|
||||||
| θ value | 8.99809 (rank : 20) | NC score | 0.003766 (rank : 27) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 421 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5DTJ9, Q7TPW5, Q8BZ76 | Gene names | Mypn, Kiaa4170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myopalladin. | |||||
|
PSL1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 21) | NC score | 0.007411 (rank : 25) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8TCT7, O60365, Q567S3, Q8IUH9, Q9BUY6, Q9H3M4, Q9NPN2, Q9P1Z6 | Gene names | SPPL2B, KIAA1532, PSL1 | |||
|
Domain Architecture |

|
|||||
| Description | Signal peptide peptidase-like 2B (EC 3.4.23.-) (Protein SPP-like 2B) (Protein SPPL2b) (Intramembrane protease 4) (IMP4) (Presenilin-like protein 1). | |||||
|
PDCD2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 22) | NC score | 0.063587 (rank : 16) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q16342, Q9UH12 | Gene names | PDCD2, RP8, ZMYND7 | |||
|
Domain Architecture |
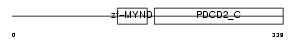
|
|||||
| Description | Programmed cell death protein 2 (Zinc finger protein Rp-8) (Zinc finger MYND domain-containing protein 7). | |||||
|
PDCD2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 23) | NC score | 0.070405 (rank : 10) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P46718, Q8BR06 | Gene names | Pdcd2, Rp8 | |||
|
Domain Architecture |
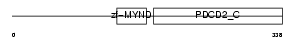
|
|||||
| Description | Programmed cell death protein 2 (Zinc finger protein Rp-8). | |||||
|
SMYD2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 24) | NC score | 0.063726 (rank : 15) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NRG4, Q4V765, Q5VSH9 | Gene names | SMYD2 | |||
|
Domain Architecture |

|
|||||
| Description | SET and MYND domain-containing protein 2 (HSKM-B). | |||||
|
SMYD2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 25) | NC score | 0.065141 (rank : 13) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8R5A0 | Gene names | Smyd2 | |||
|
Domain Architecture |
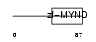
|
|||||
| Description | SET and MYND domain-containing protein 2. | |||||
|
ZMY10_HUMAN
|
||||||
| θ value | θ > 10 (rank : 26) | NC score | 0.067570 (rank : 12) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O75800, O14570, O75801, Q8N4R6, Q8NDN6 | Gene names | ZMYND10, BLU | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger MYND domain-containing protein 10 (BLu protein). | |||||
|
ZMY10_MOUSE
|
||||||
| θ value | θ > 10 (rank : 27) | NC score | 0.062874 (rank : 17) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99ML0, Q80W73 | Gene names | Zmynd10, Blu | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger MYND domain-containing protein 10 (BLu protein). | |||||
|
ZMY17_MOUSE
|
||||||
| θ value | θ > 10 (rank : 28) | NC score | 0.068041 (rank : 11) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9D5Z5 | Gene names | Zmynd17 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger MYND domain-containing protein 17. | |||||
|
EGLN2_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q96KS0, Q8WWY4, Q9BV14 | Gene names | EGLN2, EIT6 | |||
|
Domain Architecture |

|
|||||
| Description | Egl nine homolog 2 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl hydroxylase 1) (HIF-prolyl hydroxylase 1) (HIF-PH1) (HPH-3) (Prolyl hydroxylase domain-containing protein 1) (PHD1) (Estrogen-induced tag 6). | |||||
|
EGLN2_MOUSE
|
||||||
| NC score | 0.996205 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q91YE2, Q8C6I4, Q8CIL9, Q8VHJ1, Q99MI0 | Gene names | Egln2 | |||
|
Domain Architecture |

|
|||||
| Description | Egl nine homolog 2 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl hydroxylase 1) (HIF-prolyl hydroxylase 1) (HIF-PH1) (Falkor). | |||||
|
EGLN3_MOUSE
|
||||||
| NC score | 0.976462 (rank : 3) | θ value | 2.73302e-81 (rank : 5) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q91UZ4, Q3TCG8, Q8C8M6, Q8CCA8, Q8R5C7 | Gene names | Egln3 | |||
|
Domain Architecture |

|
|||||
| Description | Egl nine homolog 3 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl hydroxylase 3) (HIF-prolyl hydroxylase 3) (HIF-PH3) (SM-20). | |||||
|
EGLN3_HUMAN
|
||||||
| NC score | 0.976252 (rank : 4) | θ value | 6.08852e-81 (rank : 6) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9H6Z9 | Gene names | EGLN3 | |||
|
Domain Architecture |

|
|||||
| Description | Egl nine homolog 3 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl hydroxylase 3) (HIF-prolyl hydroxylase 3) (HIF-PH3) (HPH-1) (Prolyl hydroxylase domain-containing protein 3) (PHD3). | |||||
|
EGLN1_HUMAN
|
||||||
| NC score | 0.938085 (rank : 5) | θ value | 1.88829e-90 (rank : 3) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9GZT9, Q8N3M8, Q9BZS8, Q9BZT0 | Gene names | EGLN1, C1orf12 | |||
|
Domain Architecture |

|
|||||
| Description | Egl nine homolog 1 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl hydroxylase 2) (HIF-prolyl hydroxylase 2) (HIF-PH2) (HPH-2) (Prolyl hydroxylase domain-containing protein 2) (PHD2) (SM-20). | |||||
|
EGLN1_MOUSE
|
||||||
| NC score | 0.926869 (rank : 6) | θ value | 9.37149e-90 (rank : 4) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q91YE3, Q8VHJ2, Q922P3 | Gene names | Egln1 | |||
|
Domain Architecture |

|
|||||
| Description | Egl nine homolog 1 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl hydroxylase 2) (HIF-prolyl hydroxylase 2) (HIF-PH2) (HPH-2) (SM-20). | |||||
|
P3H1_MOUSE
|
||||||
| NC score | 0.080928 (rank : 7) | θ value | 1.06291 (rank : 11) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q3V1T4, Q3TWX8, Q8BSV2, Q8CFL3, Q9CWK5, Q9QZT6, Q9QZT7 | Gene names | Lepre1, Gros1, P3h1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 1 precursor (EC 1.14.11.7) (Leucine- and proline- enriched proteoglycan 1) (Leprecan-1) (Growth suppressor 1). | |||||
|
P3H2_MOUSE
|
||||||
| NC score | 0.079510 (rank : 8) | θ value | 0.813845 (rank : 10) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8CG71, Q8C673 | Gene names | Leprel1, P3h2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 2 precursor (EC 1.14.11.7) (Leprecan-like protein 1). | |||||
|
P3H1_HUMAN
|
||||||
| NC score | 0.076681 (rank : 9) | θ value | 0.813845 (rank : 9) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q32P28, Q7KZR4, Q96BR8, Q96SK8, Q96SL5, Q96SN3, Q9H6K3, Q9HC86, Q9HC87 | Gene names | LEPRE1, GROS1, P3H1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 1 precursor (EC 1.14.11.7) (Leucine- and proline- enriched proteoglycan 1) (Leprecan-1) (Growth suppressor 1). | |||||
|
PDCD2_MOUSE
|
||||||
| NC score | 0.070405 (rank : 10) | θ value | θ > 10 (rank : 23) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P46718, Q8BR06 | Gene names | Pdcd2, Rp8 | |||
|
Domain Architecture |

|
|||||
| Description | Programmed cell death protein 2 (Zinc finger protein Rp-8). | |||||
|
ZMY17_MOUSE
|
||||||
| NC score | 0.068041 (rank : 11) | θ value | θ > 10 (rank : 28) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9D5Z5 | Gene names | Zmynd17 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger MYND domain-containing protein 17. | |||||
|
ZMY10_HUMAN
|
||||||
| NC score | 0.067570 (rank : 12) | θ value | θ > 10 (rank : 26) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O75800, O14570, O75801, Q8N4R6, Q8NDN6 | Gene names | ZMYND10, BLU | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger MYND domain-containing protein 10 (BLu protein). | |||||
|
SMYD2_MOUSE
|
||||||
| NC score | 0.065141 (rank : 13) | θ value | θ > 10 (rank : 25) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8R5A0 | Gene names | Smyd2 | |||
|
Domain Architecture |

|
|||||
| Description | SET and MYND domain-containing protein 2. | |||||
|
P3H2_HUMAN
|
||||||
| NC score | 0.064079 (rank : 14) | θ value | 2.36792 (rank : 13) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8IVL5, Q9NVI2 | Gene names | LEPREL1, MLAT4, P3H2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 2 precursor (EC 1.14.11.7) (Leprecan-like protein 1) (Myxoid liposarcoma-associated protein 4). | |||||
|
SMYD2_HUMAN
|
||||||
| NC score | 0.063726 (rank : 15) | θ value | θ > 10 (rank : 24) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NRG4, Q4V765, Q5VSH9 | Gene names | SMYD2 | |||
|
Domain Architecture |

|
|||||
| Description | SET and MYND domain-containing protein 2 (HSKM-B). | |||||
|
PDCD2_HUMAN
|
||||||
| NC score | 0.063587 (rank : 16) | θ value | θ > 10 (rank : 22) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q16342, Q9UH12 | Gene names | PDCD2, RP8, ZMYND7 | |||
|
Domain Architecture |

|
|||||
| Description | Programmed cell death protein 2 (Zinc finger protein Rp-8) (Zinc finger MYND domain-containing protein 7). | |||||
|
ZMY10_MOUSE
|
||||||
| NC score | 0.062874 (rank : 17) | θ value | θ > 10 (rank : 27) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99ML0, Q80W73 | Gene names | Zmynd10, Blu | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger MYND domain-containing protein 10 (BLu protein). | |||||
|
K1543_HUMAN
|
||||||
| NC score | 0.062358 (rank : 18) | θ value | 0.00390308 (rank : 7) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9P1Y5, Q8NDF1 | Gene names | KIAA1543 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1543. | |||||
|
DUT_HUMAN
|
||||||
| NC score | 0.032171 (rank : 19) | θ value | 3.0926 (rank : 14) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P33316, O14785, Q16708, Q16860, Q96Q81 | Gene names | DUT | |||
|
Domain Architecture |

|
|||||
| Description | Deoxyuridine 5'-triphosphate nucleotidohydrolase, mitochondrial precursor (EC 3.6.1.23) (dUTPase) (dUTP pyrophosphatase). | |||||
|
MYF6_HUMAN
|
||||||
| NC score | 0.014301 (rank : 20) | θ value | 4.03905 (rank : 15) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P23409, Q53X80, Q6FHI9 | Gene names | MYF6 | |||
|
Domain Architecture |

|
|||||
| Description | Myogenic factor 6 (Myf-6). | |||||
|
FA38A_HUMAN
|
||||||
| NC score | 0.011466 (rank : 21) | θ value | 6.88961 (rank : 17) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q92508 | Gene names | FAM38A, KIAA0233 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM38A. | |||||
|
RGS3_MOUSE
|
||||||
| NC score | 0.011024 (rank : 22) | θ value | 1.38821 (rank : 12) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
TTBK1_HUMAN
|
||||||
| NC score | 0.009496 (rank : 23) | θ value | 0.62314 (rank : 8) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 1058 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5TCY1, Q2L6C6, Q8N444, Q96JH2 | Gene names | TTBK1, BDTK, KIAA1855 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tau-tubulin kinase 1 (EC 2.7.11.1) (Brain-derived tau kinase). | |||||
|
FGD3_HUMAN
|
||||||
| NC score | 0.007988 (rank : 24) | θ value | 6.88961 (rank : 18) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5JSP0, Q7Z7D9, Q8N5G1 | Gene names | FGD3, ZFYVE5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 3 (Zinc finger FYVE domain-containing protein 5). | |||||
|
PSL1_HUMAN
|
||||||
| NC score | 0.007411 (rank : 25) | θ value | 8.99809 (rank : 21) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8TCT7, O60365, Q567S3, Q8IUH9, Q9BUY6, Q9H3M4, Q9NPN2, Q9P1Z6 | Gene names | SPPL2B, KIAA1532, PSL1 | |||
|
Domain Architecture |

|
|||||
| Description | Signal peptide peptidase-like 2B (EC 3.4.23.-) (Protein SPP-like 2B) (Protein SPPL2b) (Intramembrane protease 4) (IMP4) (Presenilin-like protein 1). | |||||
|
FXL19_MOUSE
|
||||||
| NC score | 0.007058 (rank : 26) | θ value | 8.99809 (rank : 19) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6PB97, Q7TSH0, Q8BIB4 | Gene names | Fbxl19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 19 (F-box and leucine-rich repeat protein 19). | |||||
|
MYPN_MOUSE
|
||||||
| NC score | 0.003766 (rank : 27) | θ value | 8.99809 (rank : 20) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 421 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5DTJ9, Q7TPW5, Q8BZ76 | Gene names | Mypn, Kiaa4170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myopalladin. | |||||
|
EPHB6_HUMAN
|
||||||
| NC score | 0.000855 (rank : 28) | θ value | 5.27518 (rank : 16) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 912 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O15197 | Gene names | EPHB6 | |||
|
Domain Architecture |

|
|||||
| Description | Ephrin type-B receptor 6 precursor (Tyrosine-protein kinase-defective receptor EPH-6) (HEP). | |||||