Please be patient as the page loads
|
CGAT2_HUMAN
|
||||||
| SwissProt Accessions | Q8N6G5, Q6MZJ5, Q6MZP6, Q8TCH4, Q9P1I6 | Gene names | GALNACT2, CHGN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin beta-1,4-N-acetylgalactosaminyltransferase 2 (EC 2.4.1.174) (GalNAcT-2) (beta4GalNAcT-2). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
CGAT1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.982765 (rank : 4) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8TDX6, Q6P9G6, Q8IUF9, Q9NSQ7, Q9NUM9 | Gene names | GALNACT1, CHGN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin beta-1,4-N-acetylgalactosaminyltransferase 1 (EC 2.4.1.174) (beta4GalNAcT-1). | |||||
|
CGAT2_HUMAN
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q8N6G5, Q6MZJ5, Q6MZP6, Q8TCH4, Q9P1I6 | Gene names | GALNACT2, CHGN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin beta-1,4-N-acetylgalactosaminyltransferase 2 (EC 2.4.1.174) (GalNAcT-2) (beta4GalNAcT-2). | |||||
|
CGAT2_MOUSE
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.995528 (rank : 2) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8C1F4, Q6PAP6, Q8R5A2, Q9D2M1 | Gene names | Galnact2, Chgn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin beta-1,4-N-acetylgalactosaminyltransferase 2 (EC 2.4.1.174) (GalNAcT-2) (beta4GalNAcT-2). | |||||
|
CGAT1_MOUSE
|
||||||
| θ value | 5.52945e-183 (rank : 4) | NC score | 0.983558 (rank : 3) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8BJQ9, Q3UZX6, Q8BWV9, Q8BZU7, Q8C195, Q8R0M6 | Gene names | Galnact1, Chgn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin beta-1,4-N-acetylgalactosaminyltransferase 1 (EC 2.4.1.174) (beta4GalNAcT-1). | |||||
|
CHSS3_HUMAN
|
||||||
| θ value | 2.75849e-49 (rank : 5) | NC score | 0.818138 (rank : 5) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q70JA7, Q76L22, Q86Y52 | Gene names | CSS3, CHSY2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate synthase 3 (EC 2.4.1.175) (Glucuronosyl-N- acetylgalactosaminyl-proteoglycan 4-beta-N- acetylgalactosaminyltransferase II) (Chondroitin synthase 2) (N- acetylgalactosaminyl-proteoglycan 3-beta-glucuronosyltransferase II) (EC 2.4.1.226) (Chondroitin glucuronyltransferase II) (N- acetylgalactosaminyltransferase II) (Carbohydrate synthase 2). | |||||
|
CHSS1_HUMAN
|
||||||
| θ value | 8.30557e-46 (rank : 6) | NC score | 0.809579 (rank : 6) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q86X52, Q6UX38, Q7LFU5, Q9Y2J5 | Gene names | CHSY1, CHSY, CSS1, KIAA0990 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate synthase 1 (EC 2.4.1.175) (Glucuronosyl-N- acetylgalactosaminyl-proteoglycan 4-beta-N- acetylgalactosaminyltransferase 1) (Chondroitin sulfate synthase 1) (N-acetylgalactosaminyl-proteoglycan 3-beta-glucuronosyltransferase 1) (EC 2.4.1.226) (Chondroitin glucuronyltransferase II) (N- acetylgalactosaminyltransferase II). | |||||
|
CHSS1_MOUSE
|
||||||
| θ value | 9.18288e-45 (rank : 7) | NC score | 0.804713 (rank : 7) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6ZQ11 | Gene names | Chsy1, Kiaa0990 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate synthase 1 (EC 2.4.1.175) (Glucuronosyl-N- acetylgalactosaminyl-proteoglycan 4-beta-N- acetylgalactosaminyltransferase) (Chondroitin sulfate synthase 1) (N- acetylgalactosaminyl-proteoglycan 3-beta-glucuronosyltransferase) (EC 2.4.1.226) (Chondroitin glucuronyltransferase II) (N- acetylgalactosaminyltransferase II). | |||||
|
B4GN3_HUMAN
|
||||||
| θ value | 5.25075e-16 (rank : 8) | NC score | 0.567510 (rank : 9) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q6L9W6, Q6ZNC1, Q8N7T6 | Gene names | B4GALNT3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 2 (EC 2.4.1.244) (NGalNAc-T2) (Beta- 1,4-N-acetylgalactosaminyltransferase III) (Beta4GalNAc-T3) (Beta4GalNAcT3). | |||||
|
B4GN4_MOUSE
|
||||||
| θ value | 1.16975e-15 (rank : 9) | NC score | 0.562996 (rank : 10) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q766D5, Q8CHV2 | Gene names | B4galnt4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 1 (EC 2.4.1.244) (NGalNAc-T1) (Beta- 1,4-N-acetylgalactosaminyltransferase IV) (Beta4GalNAc-T4) (Beta4GalNAcT4). | |||||
|
B4GN3_MOUSE
|
||||||
| θ value | 1.52774e-15 (rank : 10) | NC score | 0.577507 (rank : 8) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q6L8S8 | Gene names | B4galnt3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 2 (EC 2.4.1.244) (NGalNAc-T2) (Beta- 1,4-N-acetylgalactosaminyltransferase III) (Beta4GalNAc-T3) (Beta4GalNAcT3). | |||||
|
B4GN4_HUMAN
|
||||||
| θ value | 8.38298e-14 (rank : 11) | NC score | 0.560313 (rank : 11) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q76KP1, Q96LV2 | Gene names | B4GALNT4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 1 (EC 2.4.1.244) (NGalNAc-T1) (Beta- 1,4-N-acetylgalactosaminyltransferase IV) (Beta4GalNAc-T4) (Beta4GalNAcT4). | |||||
|
CHSS2_HUMAN
|
||||||
| θ value | 1.02475e-11 (rank : 12) | NC score | 0.526527 (rank : 12) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8IZ52, Q6UXD6, Q7L4G1, Q9H0F8, Q9H618 | Gene names | CSS2, CHPF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate synthase 2 (EC 2.4.1.175) (Glucuronosyl-N- acetylgalactosaminyl-proteoglycan 4-beta-N- acetylgalactosaminyltransferase II) (N-acetylgalactosaminyl- proteoglycan 3-beta-glucuronosyltransferase II) (EC 2.4.1.226) (Chondroitin glucuronyltransferase II) (N- acetylgalactosaminyltransferase) (Chondroitin-polymerizing factor). | |||||
|
CHSS2_MOUSE
|
||||||
| θ value | 1.47974e-10 (rank : 13) | NC score | 0.520474 (rank : 13) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6IQX7 | Gene names | Css2, D1Bwg1363e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate synthase 2 (EC 2.4.1.175) (Glucuronosyl-N- acetylgalactosaminyl-proteoglycan 4-beta-N- acetylgalactosaminyltransferase II) (N-acetylgalactosaminyl- proteoglycan 3-beta-glucuronosyltransferase II) (EC 2.4.1.226) (Chondroitin glucuronyltransferase II) (N- acetylgalactosaminyltransferase). | |||||
|
CHGUT_HUMAN
|
||||||
| θ value | 7.59969e-07 (rank : 14) | NC score | 0.476460 (rank : 14) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9P2E5, Q6P2I4, Q6UXD2 | Gene names | CSGLCAT, KIAA1402 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate glucuronyltransferase (EC 2.4.1.226) (N- acetylgalactosaminyl-proteoglycan 3-beta-glucuronosyltransferase) (Chondroitin glucuronyltransferase II) (CSGlcA-T). | |||||
|
EEA1_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 15) | NC score | 0.030734 (rank : 30) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1796 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q15075, Q14221 | Gene names | EEA1, ZFYVE2 | |||
|
Domain Architecture |

|
|||||
| Description | Early endosome antigen 1 (Endosome-associated protein p162) (Zinc finger FYVE domain-containing protein 2). | |||||
|
ITSN2_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 16) | NC score | 0.021405 (rank : 43) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1371 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9Z0R6, Q9Z0R5 | Gene names | Itsn2, Ese2, Sh3d1B | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-2 (SH3 domain-containing protein 1B) (EH and SH3 domains protein 2) (EH domain and SH3 domain regulator of endocytosis 2). | |||||
|
B4GT1_MOUSE
|
||||||
| θ value | 0.21417 (rank : 17) | NC score | 0.101490 (rank : 19) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P15535 | Gene names | B4galt1, Ggtb, Ggtb2 | |||
|
Domain Architecture |
|
|||||
| Description | Beta-1,4-galactosyltransferase 1 (EC 2.4.1.-) (Beta-1,4-GalTase 1) (Beta4Gal-T1) (b4Gal-T1) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 1) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 1) [Includes: Lactose synthase A protein (EC 2.4.1.22); N-acetyllactosamine synthase (EC 2.4.1.90) (Nal synthetase); Beta-N-acetylglucosaminylglycopeptide beta-1,4- galactosyltransferase (EC 2.4.1.38); Beta-N-acetylglucosaminyl- glycolipid beta-1,4-galactosyltransferase (EC 2.4.1.-)]. | |||||
|
CCD39_HUMAN
|
||||||
| θ value | 0.21417 (rank : 18) | NC score | 0.030909 (rank : 29) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 868 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9UFE4 | Gene names | CCDC39 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 39. | |||||
|
MYO5C_HUMAN
|
||||||
| θ value | 0.21417 (rank : 19) | NC score | 0.024285 (rank : 35) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1206 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9NQX4 | Gene names | MYO5C | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5C (Myosin Vc). | |||||
|
CK5P2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 20) | NC score | 0.029876 (rank : 32) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1433 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q96SN8, Q7Z3L4, Q7Z3U1, Q7Z7I6, Q9BSW0, Q9H6J6, Q9HCD9, Q9NV90, Q9UIW9 | Gene names | CDK5RAP2, CEP215, KIAA1633 | |||
|
Domain Architecture |

|
|||||
| Description | CDK5 regulatory subunit-associated protein 2 (CDK5 activator-binding protein C48) (Centrosome-associated protein 215). | |||||
|
CUTL2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 21) | NC score | 0.035889 (rank : 27) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O14529 | Gene names | CUTL2, KIAA0293 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 2 (Homeobox protein Cux-2) (Cut-like 2). | |||||
|
B4GT1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 22) | NC score | 0.084223 (rank : 20) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P15291, Q12909, Q12910, Q12911, Q14456, Q14509, Q14523 | Gene names | B4GALT1, GGTB2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-1,4-galactosyltransferase 1 (EC 2.4.1.-) (Beta-1,4-GalTase 1) (Beta4Gal-T1) (b4Gal-T1) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 1) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 1) [Includes: Lactose synthase A protein (EC 2.4.1.22); N-acetyllactosamine synthase (EC 2.4.1.90) (Nal synthetase); Beta-N-acetylglucosaminylglycopeptide beta-1,4- galactosyltransferase (EC 2.4.1.38); Beta-N-acetylglucosaminyl- glycolipid beta-1,4-galactosyltransferase (EC 2.4.1.-)]. | |||||
|
CP250_HUMAN
|
||||||
| θ value | 0.62314 (rank : 23) | NC score | 0.022136 (rank : 40) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1805 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9BV73, O14812, O60588, Q9H450 | Gene names | CEP250, CEP2, CNAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1). | |||||
|
CT096_HUMAN
|
||||||
| θ value | 0.813845 (rank : 24) | NC score | 0.044505 (rank : 26) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 438 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9NUD7, Q8N840, Q8NAX5 | Gene names | C20orf96 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C20orf96. | |||||
|
B4GT2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 25) | NC score | 0.116829 (rank : 16) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Z2Y2, Q3TMP2 | Gene names | B4galt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-1,4-galactosyltransferase 2 (EC 2.4.1.-) (Beta-1,4-GalTase 2) (Beta4Gal-T2) (b4Gal-T2) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 2) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 2) [Includes: Lactose synthase A protein (EC 2.4.1.22); N-acetyllactosamine synthase (EC 2.4.1.90) (Nal synthetase); Beta-N-acetylglucosaminylglycopeptide beta-1,4- galactosyltransferase (EC 2.4.1.38); Beta-N-acetylglucosaminyl- glycolipid beta-1,4-galactosyltransferase (EC 2.4.1.-)]. | |||||
|
B4GT4_HUMAN
|
||||||
| θ value | 1.38821 (rank : 26) | NC score | 0.075948 (rank : 22) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O60513, Q68D68, Q9BSW3, Q9C078 | Gene names | B4GALT4 | |||
|
Domain Architecture |
|
|||||
| Description | Beta-1,4-galactosyltransferase 4 (EC 2.4.1.-) (Beta-1,4-GalTase 4) (Beta4Gal-T4) (b4Gal-T4) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 4) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 4) [Includes: N-acetyllactosamine synthase (EC 2.4.1.90) (Nal synthetase); Beta-N-acetylglucosaminyl-glycolipid beta-1,4-galactosyltransferase (EC 2.4.1.-)]. | |||||
|
SMC1A_MOUSE
|
||||||
| θ value | 1.38821 (rank : 27) | NC score | 0.020232 (rank : 45) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1384 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9CU62, Q3V480, Q9CUX9, Q9D959, Q9WTU1 | Gene names | Smc1l1, Sb1.8, Smc1, Smcb | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Chromosome segregation protein SmcB) (Sb1.8). | |||||
|
B4GT4_MOUSE
|
||||||
| θ value | 1.81305 (rank : 28) | NC score | 0.081238 (rank : 21) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9JJ04, Q8BR54, Q9QY12 | Gene names | B4galt4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-1,4-galactosyltransferase 4 (EC 2.4.1.-) (Beta-1,4-GalTase 4) (Beta4Gal-T4) (b4Gal-T4) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 4) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 4) [Includes: N-acetyllactosamine synthase (EC 2.4.1.90) (Nal synthetase); Beta-N-acetylglucosaminyl-glycolipid beta-1,4-galactosyltransferase (EC 2.4.1.-)]. | |||||
|
WWC3_HUMAN
|
||||||
| θ value | 1.81305 (rank : 29) | NC score | 0.034134 (rank : 28) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9ULE0, Q659C1, Q9BTQ1 | Gene names | WWC3, KIAA1280 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WWC family member 3. | |||||
|
B4GT2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 30) | NC score | 0.112910 (rank : 17) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O60909, O60511, Q5T4X5, Q5T4Y5, Q9BUP6, Q9NSY7 | Gene names | B4GALT2 | |||
|
Domain Architecture |
|
|||||
| Description | Beta-1,4-galactosyltransferase 2 (EC 2.4.1.-) (Beta-1,4-GalTase 2) (Beta4Gal-T2) (b4Gal-T2) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 2) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 2) [Includes: Lactose synthase A protein (EC 2.4.1.22); N-acetyllactosamine synthase (EC 2.4.1.90) (Nal synthetase); Beta-N-acetylglucosaminylglycopeptide beta-1,4- galactosyltransferase (EC 2.4.1.38); Beta-N-acetylglucosaminyl- glycolipid beta-1,4-galactosyltransferase (EC 2.4.1.-)]. | |||||
|
FYCO1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 31) | NC score | 0.026186 (rank : 34) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1059 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8VDC1, Q3V385, Q7TMT0, Q8BJN2 | Gene names | Fyco1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE and coiled-coil domain-containing protein 1. | |||||
|
APXL_HUMAN
|
||||||
| θ value | 3.0926 (rank : 32) | NC score | 0.023238 (rank : 37) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 553 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q13796 | Gene names | APXL | |||
|
Domain Architecture |
|
|||||
| Description | Apical-like protein (Protein APXL). | |||||
|
BCN1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 33) | NC score | 0.029904 (rank : 31) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q14457, O75595, Q9UNA8 | Gene names | BECN1, GT197 | |||
|
Domain Architecture |

|
|||||
| Description | Beclin-1 (Coiled-coil myosin-like BCL2-interacting protein) (Protein GT197). | |||||
|
CAR11_HUMAN
|
||||||
| θ value | 3.0926 (rank : 34) | NC score | 0.021669 (rank : 42) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9BXL7, Q548H3 | Gene names | CARD11, CARMA1 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 11 (CARD-containing MAGUK protein 3) (Carma 1). | |||||
|
LIPB1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 35) | NC score | 0.022664 (rank : 39) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 727 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8C8U0, Q69ZN5, Q6GQV3, Q80VB4, Q9CUT7 | Gene names | Ppfibp1, Kiaa1230 | |||
|
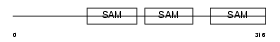
Domain Architecture |
|
|||||
| Description | Liprin-beta-1 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein-binding protein 1) (PTPRF-interacting protein-binding protein 1). | |||||
|
MYH11_MOUSE
|
||||||
| θ value | 3.0926 (rank : 36) | NC score | 0.022784 (rank : 38) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1777 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O08638, O08639, Q62462, Q64195 | Gene names | Myh11 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
GFAP_MOUSE
|
||||||
| θ value | 4.03905 (rank : 37) | NC score | 0.012946 (rank : 56) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 725 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P03995, Q7TQ30, Q925K3 | Gene names | Gfap | |||
|
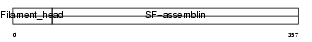
Domain Architecture |
|
|||||
| Description | Glial fibrillary acidic protein, astrocyte (GFAP). | |||||
|
MYH11_HUMAN
|
||||||
| θ value | 4.03905 (rank : 38) | NC score | 0.021777 (rank : 41) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1806 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P35749, O00396, O94944, P78422 | Gene names | MYH11, KIAA0866 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
TFAP4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 39) | NC score | 0.018734 (rank : 50) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q01664, O60409 | Gene names | TFAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor AP-4 (Activating enhancer-binding protein 4). | |||||
|
TTC28_MOUSE
|
||||||
| θ value | 4.03905 (rank : 40) | NC score | 0.012092 (rank : 57) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80XJ3, Q6P9J6, Q80TL5, Q8BV10, Q8C0F2 | Gene names | Ttc28, Kiaa1043 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tetratricopeptide repeat protein 28 (TPR repeat protein 28). | |||||
|
ANR35_HUMAN
|
||||||
| θ value | 6.88961 (rank : 41) | NC score | 0.010095 (rank : 61) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 784 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8N283, Q3MJ10, Q96LS3 | Gene names | ANKRD35 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 35. | |||||
|
GCC1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 42) | NC score | 0.020841 (rank : 44) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 887 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q96CN9, Q9H6N7 | Gene names | GCC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GRIP and coiled-coil domain-containing protein 1 (Golgi coiled coil protein 1). | |||||
|
GP110_MOUSE
|
||||||
| θ value | 6.88961 (rank : 43) | NC score | 0.005227 (rank : 63) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8VEC3, Q8BMV8 | Gene names | Gpr110 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-protein coupled receptor 110 precursor. | |||||
|
HOOK2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 44) | NC score | 0.026222 (rank : 33) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 714 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q96ED9, O60562 | Gene names | HOOK2 | |||
|
Domain Architecture |

|
|||||
| Description | Hook homolog 2 (h-hook2) (hHK2). | |||||
|
MACF1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 45) | NC score | 0.015592 (rank : 55) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1341 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9UPN3, O75053, Q5VW20, Q8WXY2, Q9H540, Q9UKP0, Q9ULG9 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoforms 1/2/3/5 (Actin cross-linking family protein 7) (Macrophin-1) (Trabeculin-alpha) (620 kDa actin-binding protein) (ABP620). | |||||
|
MACF4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 46) | NC score | 0.018002 (rank : 51) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1321 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q96PK2, Q8WXY1 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoform 4. | |||||
|
MMS19_HUMAN
|
||||||
| θ value | 6.88961 (rank : 47) | NC score | 0.011282 (rank : 59) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96T76, Q5T455, Q66K82, Q7L4W8, Q969Z1, Q96DF1, Q96MR1, Q96RK5, Q96SK1, Q9BUE2, Q9BYS9 | Gene names | MMS19L, MMS19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MMS19-like protein (hMMS19) (MET18 homolog). | |||||
|
MYH10_HUMAN
|
||||||
| θ value | 6.88961 (rank : 48) | NC score | 0.019833 (rank : 48) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P35580, Q16087 | Gene names | MYH10 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
MYH10_MOUSE
|
||||||
| θ value | 6.88961 (rank : 49) | NC score | 0.019940 (rank : 46) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1612 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q61879, Q5SV63 | Gene names | Myh10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
ZBT20_HUMAN
|
||||||
| θ value | 6.88961 (rank : 50) | NC score | -0.001937 (rank : 64) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 835 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9HC78 | Gene names | ZBTB20, DPZF, ZNF288 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 20 (Zinc finger protein 288) (Dendritic-derived BTB/POZ zinc finger protein). | |||||
|
CENPE_HUMAN
|
||||||
| θ value | 8.99809 (rank : 51) | NC score | 0.015648 (rank : 54) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 2060 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q02224 | Gene names | CENPE | |||
|
Domain Architecture |

|
|||||
| Description | Centromeric protein E (CENP-E). | |||||
|
CEP72_MOUSE
|
||||||
| θ value | 8.99809 (rank : 52) | NC score | 0.015724 (rank : 53) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9D3R3, Q8BM53, Q9D0B7 | Gene names | Cep72 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 72 kDa (Cep72 protein). | |||||
|
CP135_HUMAN
|
||||||
| θ value | 8.99809 (rank : 53) | NC score | 0.019905 (rank : 47) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1290 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q66GS9, O75130, Q9H8H7 | Gene names | CEP135, CEP4, KIAA0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||
|
KIFC3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 54) | NC score | 0.008981 (rank : 62) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 619 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9BVG8, O75299, Q8IUT3, Q96HW6 | Gene names | KIFC3 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIFC3. | |||||
|
KTN1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 55) | NC score | 0.019179 (rank : 49) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1324 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q61595, Q8BG49, Q8BHF4, Q8BHM8, Q8C9Y5, Q8CG51, Q8CG52, Q8CG53, Q8CG54, Q8CG55, Q8CG56, Q8CG57, Q8CG58, Q8CG59, Q8CG60, Q8CG61, Q8CG62, Q8CG63 | Gene names | Ktn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinectin. | |||||
|
MYH9_HUMAN
|
||||||
| θ value | 8.99809 (rank : 56) | NC score | 0.017934 (rank : 52) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P35579, O60805 | Gene names | MYH9 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
RBP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 57) | NC score | 0.010860 (rank : 60) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 521 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q15311 | Gene names | RALBP1, RLIP1, RLIP76 | |||
|
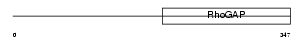
Domain Architecture |
|
|||||
| Description | RalA-binding protein 1 (RalBP1) (Ral-interacting protein 1) (76 kDa Ral-interacting protein) (Dinitrophenyl S-glutathione ATPase) (DNP-SG ATPase). | |||||
|
RFIP4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 58) | NC score | 0.023953 (rank : 36) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q86YS3, Q8N829, Q8NDT7, Q969D8 | Gene names | RAB11FIP4, KIAA1821 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab11 family-interacting protein 4 (Rab11-FIP4). | |||||
|
VIME_HUMAN
|
||||||
| θ value | 8.99809 (rank : 59) | NC score | 0.011868 (rank : 58) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 706 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P08670, Q15867, Q15868, Q15869, Q6LER9, Q8N850, Q96ML2, Q9NTM3 | Gene names | VIM | |||
|
Domain Architecture |

|
|||||
| Description | Vimentin. | |||||
|
B3GLT_HUMAN
|
||||||
| θ value | θ > 10 (rank : 60) | NC score | 0.108890 (rank : 18) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6Y288, Q5W0H2, Q6NUI3 | Gene names | B3GALTL, B3GTL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-1,3-glucosyltransferase (EC 2.4.1.-) (Beta3Glc-T) (Beta-3- glycosyltransferase-like). | |||||
|
B3GLT_MOUSE
|
||||||
| θ value | θ > 10 (rank : 61) | NC score | 0.117428 (rank : 15) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BHT6, Q0PCR7, Q149S5 | Gene names | B3galtl, Gm1057 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-1,3-glucosyltransferase (EC 2.4.1.-) (Beta3Glc-T) (Beta-3- glycosyltransferase-like). | |||||
|
LFNG_HUMAN
|
||||||
| θ value | θ > 10 (rank : 62) | NC score | 0.051247 (rank : 25) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NES3, O00589, Q96C39, Q9UJW5 | Gene names | LFNG | |||
|
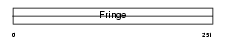
Domain Architecture |
|
|||||
| Description | Beta-1,3-N-acetylglucosaminyltransferase lunatic fringe (EC 2.4.1.222) (O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase). | |||||
|
LFNG_MOUSE
|
||||||
| θ value | θ > 10 (rank : 63) | NC score | 0.059339 (rank : 24) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O09010, Q3U659, Q8K3F1, Q9DC10 | Gene names | Lfng | |||
|
Domain Architecture |

|
|||||
| Description | Beta-1,3-N-acetylglucosaminyltransferase lunatic fringe (EC 2.4.1.222) (O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase). | |||||
|
RFNG_MOUSE
|
||||||
| θ value | θ > 10 (rank : 64) | NC score | 0.060173 (rank : 23) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O09009 | Gene names | Rfng | |||
|
Domain Architecture |
|
|||||
| Description | Beta-1,3-N-acetylglucosaminyltransferase radical fringe (EC 2.4.1.222) (O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase). | |||||
|
CGAT2_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q8N6G5, Q6MZJ5, Q6MZP6, Q8TCH4, Q9P1I6 | Gene names | GALNACT2, CHGN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin beta-1,4-N-acetylgalactosaminyltransferase 2 (EC 2.4.1.174) (GalNAcT-2) (beta4GalNAcT-2). | |||||
|
CGAT2_MOUSE
|
||||||
| NC score | 0.995528 (rank : 2) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8C1F4, Q6PAP6, Q8R5A2, Q9D2M1 | Gene names | Galnact2, Chgn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin beta-1,4-N-acetylgalactosaminyltransferase 2 (EC 2.4.1.174) (GalNAcT-2) (beta4GalNAcT-2). | |||||
|
CGAT1_MOUSE
|
||||||
| NC score | 0.983558 (rank : 3) | θ value | 5.52945e-183 (rank : 4) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8BJQ9, Q3UZX6, Q8BWV9, Q8BZU7, Q8C195, Q8R0M6 | Gene names | Galnact1, Chgn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin beta-1,4-N-acetylgalactosaminyltransferase 1 (EC 2.4.1.174) (beta4GalNAcT-1). | |||||
|
CGAT1_HUMAN
|
||||||
| NC score | 0.982765 (rank : 4) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8TDX6, Q6P9G6, Q8IUF9, Q9NSQ7, Q9NUM9 | Gene names | GALNACT1, CHGN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin beta-1,4-N-acetylgalactosaminyltransferase 1 (EC 2.4.1.174) (beta4GalNAcT-1). | |||||
|
CHSS3_HUMAN
|
||||||
| NC score | 0.818138 (rank : 5) | θ value | 2.75849e-49 (rank : 5) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q70JA7, Q76L22, Q86Y52 | Gene names | CSS3, CHSY2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate synthase 3 (EC 2.4.1.175) (Glucuronosyl-N- acetylgalactosaminyl-proteoglycan 4-beta-N- acetylgalactosaminyltransferase II) (Chondroitin synthase 2) (N- acetylgalactosaminyl-proteoglycan 3-beta-glucuronosyltransferase II) (EC 2.4.1.226) (Chondroitin glucuronyltransferase II) (N- acetylgalactosaminyltransferase II) (Carbohydrate synthase 2). | |||||
|
CHSS1_HUMAN
|
||||||
| NC score | 0.809579 (rank : 6) | θ value | 8.30557e-46 (rank : 6) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q86X52, Q6UX38, Q7LFU5, Q9Y2J5 | Gene names | CHSY1, CHSY, CSS1, KIAA0990 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate synthase 1 (EC 2.4.1.175) (Glucuronosyl-N- acetylgalactosaminyl-proteoglycan 4-beta-N- acetylgalactosaminyltransferase 1) (Chondroitin sulfate synthase 1) (N-acetylgalactosaminyl-proteoglycan 3-beta-glucuronosyltransferase 1) (EC 2.4.1.226) (Chondroitin glucuronyltransferase II) (N- acetylgalactosaminyltransferase II). | |||||
|
CHSS1_MOUSE
|
||||||
| NC score | 0.804713 (rank : 7) | θ value | 9.18288e-45 (rank : 7) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6ZQ11 | Gene names | Chsy1, Kiaa0990 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate synthase 1 (EC 2.4.1.175) (Glucuronosyl-N- acetylgalactosaminyl-proteoglycan 4-beta-N- acetylgalactosaminyltransferase) (Chondroitin sulfate synthase 1) (N- acetylgalactosaminyl-proteoglycan 3-beta-glucuronosyltransferase) (EC 2.4.1.226) (Chondroitin glucuronyltransferase II) (N- acetylgalactosaminyltransferase II). | |||||
|
B4GN3_MOUSE
|
||||||
| NC score | 0.577507 (rank : 8) | θ value | 1.52774e-15 (rank : 10) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q6L8S8 | Gene names | B4galnt3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 2 (EC 2.4.1.244) (NGalNAc-T2) (Beta- 1,4-N-acetylgalactosaminyltransferase III) (Beta4GalNAc-T3) (Beta4GalNAcT3). | |||||
|
B4GN3_HUMAN
|
||||||
| NC score | 0.567510 (rank : 9) | θ value | 5.25075e-16 (rank : 8) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q6L9W6, Q6ZNC1, Q8N7T6 | Gene names | B4GALNT3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 2 (EC 2.4.1.244) (NGalNAc-T2) (Beta- 1,4-N-acetylgalactosaminyltransferase III) (Beta4GalNAc-T3) (Beta4GalNAcT3). | |||||
|
B4GN4_MOUSE
|
||||||
| NC score | 0.562996 (rank : 10) | θ value | 1.16975e-15 (rank : 9) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q766D5, Q8CHV2 | Gene names | B4galnt4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 1 (EC 2.4.1.244) (NGalNAc-T1) (Beta- 1,4-N-acetylgalactosaminyltransferase IV) (Beta4GalNAc-T4) (Beta4GalNAcT4). | |||||
|
B4GN4_HUMAN
|
||||||
| NC score | 0.560313 (rank : 11) | θ value | 8.38298e-14 (rank : 11) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q76KP1, Q96LV2 | Gene names | B4GALNT4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 1 (EC 2.4.1.244) (NGalNAc-T1) (Beta- 1,4-N-acetylgalactosaminyltransferase IV) (Beta4GalNAc-T4) (Beta4GalNAcT4). | |||||
|
CHSS2_HUMAN
|
||||||
| NC score | 0.526527 (rank : 12) | θ value | 1.02475e-11 (rank : 12) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8IZ52, Q6UXD6, Q7L4G1, Q9H0F8, Q9H618 | Gene names | CSS2, CHPF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate synthase 2 (EC 2.4.1.175) (Glucuronosyl-N- acetylgalactosaminyl-proteoglycan 4-beta-N- acetylgalactosaminyltransferase II) (N-acetylgalactosaminyl- proteoglycan 3-beta-glucuronosyltransferase II) (EC 2.4.1.226) (Chondroitin glucuronyltransferase II) (N- acetylgalactosaminyltransferase) (Chondroitin-polymerizing factor). | |||||
|
CHSS2_MOUSE
|
||||||
| NC score | 0.520474 (rank : 13) | θ value | 1.47974e-10 (rank : 13) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6IQX7 | Gene names | Css2, D1Bwg1363e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate synthase 2 (EC 2.4.1.175) (Glucuronosyl-N- acetylgalactosaminyl-proteoglycan 4-beta-N- acetylgalactosaminyltransferase II) (N-acetylgalactosaminyl- proteoglycan 3-beta-glucuronosyltransferase II) (EC 2.4.1.226) (Chondroitin glucuronyltransferase II) (N- acetylgalactosaminyltransferase). | |||||
|
CHGUT_HUMAN
|
||||||
| NC score | 0.476460 (rank : 14) | θ value | 7.59969e-07 (rank : 14) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9P2E5, Q6P2I4, Q6UXD2 | Gene names | CSGLCAT, KIAA1402 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate glucuronyltransferase (EC 2.4.1.226) (N- acetylgalactosaminyl-proteoglycan 3-beta-glucuronosyltransferase) (Chondroitin glucuronyltransferase II) (CSGlcA-T). | |||||
|
B3GLT_MOUSE
|
||||||
| NC score | 0.117428 (rank : 15) | θ value | θ > 10 (rank : 61) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BHT6, Q0PCR7, Q149S5 | Gene names | B3galtl, Gm1057 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-1,3-glucosyltransferase (EC 2.4.1.-) (Beta3Glc-T) (Beta-3- glycosyltransferase-like). | |||||
|
B4GT2_MOUSE
|
||||||
| NC score | 0.116829 (rank : 16) | θ value | 1.06291 (rank : 25) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Z2Y2, Q3TMP2 | Gene names | B4galt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-1,4-galactosyltransferase 2 (EC 2.4.1.-) (Beta-1,4-GalTase 2) (Beta4Gal-T2) (b4Gal-T2) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 2) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 2) [Includes: Lactose synthase A protein (EC 2.4.1.22); N-acetyllactosamine synthase (EC 2.4.1.90) (Nal synthetase); Beta-N-acetylglucosaminylglycopeptide beta-1,4- galactosyltransferase (EC 2.4.1.38); Beta-N-acetylglucosaminyl- glycolipid beta-1,4-galactosyltransferase (EC 2.4.1.-)]. | |||||
|
B4GT2_HUMAN
|
||||||
| NC score | 0.112910 (rank : 17) | θ value | 2.36792 (rank : 30) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O60909, O60511, Q5T4X5, Q5T4Y5, Q9BUP6, Q9NSY7 | Gene names | B4GALT2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-1,4-galactosyltransferase 2 (EC 2.4.1.-) (Beta-1,4-GalTase 2) (Beta4Gal-T2) (b4Gal-T2) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 2) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 2) [Includes: Lactose synthase A protein (EC 2.4.1.22); N-acetyllactosamine synthase (EC 2.4.1.90) (Nal synthetase); Beta-N-acetylglucosaminylglycopeptide beta-1,4- galactosyltransferase (EC 2.4.1.38); Beta-N-acetylglucosaminyl- glycolipid beta-1,4-galactosyltransferase (EC 2.4.1.-)]. | |||||
|
B3GLT_HUMAN
|
||||||
| NC score | 0.108890 (rank : 18) | θ value | θ > 10 (rank : 60) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6Y288, Q5W0H2, Q6NUI3 | Gene names | B3GALTL, B3GTL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-1,3-glucosyltransferase (EC 2.4.1.-) (Beta3Glc-T) (Beta-3- glycosyltransferase-like). | |||||
|
B4GT1_MOUSE
|
||||||
| NC score | 0.101490 (rank : 19) | θ value | 0.21417 (rank : 17) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P15535 | Gene names | B4galt1, Ggtb, Ggtb2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-1,4-galactosyltransferase 1 (EC 2.4.1.-) (Beta-1,4-GalTase 1) (Beta4Gal-T1) (b4Gal-T1) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 1) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 1) [Includes: Lactose synthase A protein (EC 2.4.1.22); N-acetyllactosamine synthase (EC 2.4.1.90) (Nal synthetase); Beta-N-acetylglucosaminylglycopeptide beta-1,4- galactosyltransferase (EC 2.4.1.38); Beta-N-acetylglucosaminyl- glycolipid beta-1,4-galactosyltransferase (EC 2.4.1.-)]. | |||||
|
B4GT1_HUMAN
|
||||||
| NC score | 0.084223 (rank : 20) | θ value | 0.62314 (rank : 22) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P15291, Q12909, Q12910, Q12911, Q14456, Q14509, Q14523 | Gene names | B4GALT1, GGTB2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-1,4-galactosyltransferase 1 (EC 2.4.1.-) (Beta-1,4-GalTase 1) (Beta4Gal-T1) (b4Gal-T1) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 1) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 1) [Includes: Lactose synthase A protein (EC 2.4.1.22); N-acetyllactosamine synthase (EC 2.4.1.90) (Nal synthetase); Beta-N-acetylglucosaminylglycopeptide beta-1,4- galactosyltransferase (EC 2.4.1.38); Beta-N-acetylglucosaminyl- glycolipid beta-1,4-galactosyltransferase (EC 2.4.1.-)]. | |||||
|
B4GT4_MOUSE
|
||||||
| NC score | 0.081238 (rank : 21) | θ value | 1.81305 (rank : 28) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9JJ04, Q8BR54, Q9QY12 | Gene names | B4galt4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-1,4-galactosyltransferase 4 (EC 2.4.1.-) (Beta-1,4-GalTase 4) (Beta4Gal-T4) (b4Gal-T4) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 4) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 4) [Includes: N-acetyllactosamine synthase (EC 2.4.1.90) (Nal synthetase); Beta-N-acetylglucosaminyl-glycolipid beta-1,4-galactosyltransferase (EC 2.4.1.-)]. | |||||
|
B4GT4_HUMAN
|
||||||
| NC score | 0.075948 (rank : 22) | θ value | 1.38821 (rank : 26) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O60513, Q68D68, Q9BSW3, Q9C078 | Gene names | B4GALT4 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-1,4-galactosyltransferase 4 (EC 2.4.1.-) (Beta-1,4-GalTase 4) (Beta4Gal-T4) (b4Gal-T4) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 4) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 4) [Includes: N-acetyllactosamine synthase (EC 2.4.1.90) (Nal synthetase); Beta-N-acetylglucosaminyl-glycolipid beta-1,4-galactosyltransferase (EC 2.4.1.-)]. | |||||
|
RFNG_MOUSE
|
||||||
| NC score | 0.060173 (rank : 23) | θ value | θ > 10 (rank : 64) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O09009 | Gene names | Rfng | |||
|
Domain Architecture |

|
|||||
| Description | Beta-1,3-N-acetylglucosaminyltransferase radical fringe (EC 2.4.1.222) (O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase). | |||||
|
LFNG_MOUSE
|
||||||
| NC score | 0.059339 (rank : 24) | θ value | θ > 10 (rank : 63) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O09010, Q3U659, Q8K3F1, Q9DC10 | Gene names | Lfng | |||
|
Domain Architecture |

|
|||||
| Description | Beta-1,3-N-acetylglucosaminyltransferase lunatic fringe (EC 2.4.1.222) (O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase). | |||||
|
LFNG_HUMAN
|
||||||
| NC score | 0.051247 (rank : 25) | θ value | θ > 10 (rank : 62) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NES3, O00589, Q96C39, Q9UJW5 | Gene names | LFNG | |||
|
Domain Architecture |

|
|||||
| Description | Beta-1,3-N-acetylglucosaminyltransferase lunatic fringe (EC 2.4.1.222) (O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase). | |||||
|
CT096_HUMAN
|
||||||
| NC score | 0.044505 (rank : 26) | θ value | 0.813845 (rank : 24) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 438 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9NUD7, Q8N840, Q8NAX5 | Gene names | C20orf96 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C20orf96. | |||||
|
CUTL2_HUMAN
|
||||||
| NC score | 0.035889 (rank : 27) | θ value | 0.47712 (rank : 21) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O14529 | Gene names | CUTL2, KIAA0293 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 2 (Homeobox protein Cux-2) (Cut-like 2). | |||||
|
WWC3_HUMAN
|
||||||
| NC score | 0.034134 (rank : 28) | θ value | 1.81305 (rank : 29) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9ULE0, Q659C1, Q9BTQ1 | Gene names | WWC3, KIAA1280 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WWC family member 3. | |||||
|
CCD39_HUMAN
|
||||||
| NC score | 0.030909 (rank : 29) | θ value | 0.21417 (rank : 18) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 868 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9UFE4 | Gene names | CCDC39 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 39. | |||||
|
EEA1_HUMAN
|
||||||
| NC score | 0.030734 (rank : 30) | θ value | 0.0563607 (rank : 15) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1796 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q15075, Q14221 | Gene names | EEA1, ZFYVE2 | |||
|
Domain Architecture |

|
|||||
| Description | Early endosome antigen 1 (Endosome-associated protein p162) (Zinc finger FYVE domain-containing protein 2). | |||||
|
BCN1_HUMAN
|
||||||
| NC score | 0.029904 (rank : 31) | θ value | 3.0926 (rank : 33) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q14457, O75595, Q9UNA8 | Gene names | BECN1, GT197 | |||
|
Domain Architecture |

|
|||||
| Description | Beclin-1 (Coiled-coil myosin-like BCL2-interacting protein) (Protein GT197). | |||||
|
CK5P2_HUMAN
|
||||||
| NC score | 0.029876 (rank : 32) | θ value | 0.365318 (rank : 20) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1433 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q96SN8, Q7Z3L4, Q7Z3U1, Q7Z7I6, Q9BSW0, Q9H6J6, Q9HCD9, Q9NV90, Q9UIW9 | Gene names | CDK5RAP2, CEP215, KIAA1633 | |||
|
Domain Architecture |

|
|||||
| Description | CDK5 regulatory subunit-associated protein 2 (CDK5 activator-binding protein C48) (Centrosome-associated protein 215). | |||||
|
HOOK2_HUMAN
|
||||||
| NC score | 0.026222 (rank : 33) | θ value | 6.88961 (rank : 44) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 714 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q96ED9, O60562 | Gene names | HOOK2 | |||
|
Domain Architecture |

|
|||||
| Description | Hook homolog 2 (h-hook2) (hHK2). | |||||
|
FYCO1_MOUSE
|
||||||
| NC score | 0.026186 (rank : 34) | θ value | 2.36792 (rank : 31) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1059 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8VDC1, Q3V385, Q7TMT0, Q8BJN2 | Gene names | Fyco1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE and coiled-coil domain-containing protein 1. | |||||
|
MYO5C_HUMAN
|
||||||
| NC score | 0.024285 (rank : 35) | θ value | 0.21417 (rank : 19) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1206 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9NQX4 | Gene names | MYO5C | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5C (Myosin Vc). | |||||
|
RFIP4_HUMAN
|
||||||
| NC score | 0.023953 (rank : 36) | θ value | 8.99809 (rank : 58) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q86YS3, Q8N829, Q8NDT7, Q969D8 | Gene names | RAB11FIP4, KIAA1821 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab11 family-interacting protein 4 (Rab11-FIP4). | |||||
|
APXL_HUMAN
|
||||||
| NC score | 0.023238 (rank : 37) | θ value | 3.0926 (rank : 32) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 553 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q13796 | Gene names | APXL | |||
|
Domain Architecture |

|
|||||
| Description | Apical-like protein (Protein APXL). | |||||
|
MYH11_MOUSE
|
||||||
| NC score | 0.022784 (rank : 38) | θ value | 3.0926 (rank : 36) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1777 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O08638, O08639, Q62462, Q64195 | Gene names | Myh11 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
LIPB1_MOUSE
|
||||||
| NC score | 0.022664 (rank : 39) | θ value | 3.0926 (rank : 35) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 727 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8C8U0, Q69ZN5, Q6GQV3, Q80VB4, Q9CUT7 | Gene names | Ppfibp1, Kiaa1230 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-beta-1 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein-binding protein 1) (PTPRF-interacting protein-binding protein 1). | |||||
|
CP250_HUMAN
|
||||||
| NC score | 0.022136 (rank : 40) | θ value | 0.62314 (rank : 23) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1805 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9BV73, O14812, O60588, Q9H450 | Gene names | CEP250, CEP2, CNAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1). | |||||
|
MYH11_HUMAN
|
||||||
| NC score | 0.021777 (rank : 41) | θ value | 4.03905 (rank : 38) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1806 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P35749, O00396, O94944, P78422 | Gene names | MYH11, KIAA0866 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
CAR11_HUMAN
|
||||||
| NC score | 0.021669 (rank : 42) | θ value | 3.0926 (rank : 34) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9BXL7, Q548H3 | Gene names | CARD11, CARMA1 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 11 (CARD-containing MAGUK protein 3) (Carma 1). | |||||
|
ITSN2_MOUSE
|
||||||
| NC score | 0.021405 (rank : 43) | θ value | 0.0563607 (rank : 16) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1371 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9Z0R6, Q9Z0R5 | Gene names | Itsn2, Ese2, Sh3d1B | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-2 (SH3 domain-containing protein 1B) (EH and SH3 domains protein 2) (EH domain and SH3 domain regulator of endocytosis 2). | |||||
|
GCC1_HUMAN
|
||||||
| NC score | 0.020841 (rank : 44) | θ value | 6.88961 (rank : 42) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 887 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q96CN9, Q9H6N7 | Gene names | GCC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GRIP and coiled-coil domain-containing protein 1 (Golgi coiled coil protein 1). | |||||
|
SMC1A_MOUSE
|
||||||
| NC score | 0.020232 (rank : 45) | θ value | 1.38821 (rank : 27) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1384 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9CU62, Q3V480, Q9CUX9, Q9D959, Q9WTU1 | Gene names | Smc1l1, Sb1.8, Smc1, Smcb | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Chromosome segregation protein SmcB) (Sb1.8). | |||||
|
MYH10_MOUSE
|
||||||
| NC score | 0.019940 (rank : 46) | θ value | 6.88961 (rank : 49) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1612 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q61879, Q5SV63 | Gene names | Myh10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
CP135_HUMAN
|
||||||
| NC score | 0.019905 (rank : 47) | θ value | 8.99809 (rank : 53) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1290 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q66GS9, O75130, Q9H8H7 | Gene names | CEP135, CEP4, KIAA0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||
|
MYH10_HUMAN
|
||||||
| NC score | 0.019833 (rank : 48) | θ value | 6.88961 (rank : 48) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P35580, Q16087 | Gene names | MYH10 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
KTN1_MOUSE
|
||||||
| NC score | 0.019179 (rank : 49) | θ value | 8.99809 (rank : 55) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1324 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q61595, Q8BG49, Q8BHF4, Q8BHM8, Q8C9Y5, Q8CG51, Q8CG52, Q8CG53, Q8CG54, Q8CG55, Q8CG56, Q8CG57, Q8CG58, Q8CG59, Q8CG60, Q8CG61, Q8CG62, Q8CG63 | Gene names | Ktn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinectin. | |||||
|
TFAP4_HUMAN
|
||||||
| NC score | 0.018734 (rank : 50) | θ value | 4.03905 (rank : 39) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q01664, O60409 | Gene names | TFAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor AP-4 (Activating enhancer-binding protein 4). | |||||
|
MACF4_HUMAN
|
||||||
| NC score | 0.018002 (rank : 51) | θ value | 6.88961 (rank : 46) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1321 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q96PK2, Q8WXY1 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoform 4. | |||||
|
MYH9_HUMAN
|
||||||
| NC score | 0.017934 (rank : 52) | θ value | 8.99809 (rank : 56) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P35579, O60805 | Gene names | MYH9 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
CEP72_MOUSE
|
||||||
| NC score | 0.015724 (rank : 53) | θ value | 8.99809 (rank : 52) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9D3R3, Q8BM53, Q9D0B7 | Gene names | Cep72 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 72 kDa (Cep72 protein). | |||||
|
CENPE_HUMAN
|
||||||
| NC score | 0.015648 (rank : 54) | θ value | 8.99809 (rank : 51) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 2060 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q02224 | Gene names | CENPE | |||
|
Domain Architecture |

|
|||||
| Description | Centromeric protein E (CENP-E). | |||||
|
MACF1_HUMAN
|
||||||
| NC score | 0.015592 (rank : 55) | θ value | 6.88961 (rank : 45) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 1341 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9UPN3, O75053, Q5VW20, Q8WXY2, Q9H540, Q9UKP0, Q9ULG9 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoforms 1/2/3/5 (Actin cross-linking family protein 7) (Macrophin-1) (Trabeculin-alpha) (620 kDa actin-binding protein) (ABP620). | |||||
|
GFAP_MOUSE
|
||||||
| NC score | 0.012946 (rank : 56) | θ value | 4.03905 (rank : 37) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 725 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P03995, Q7TQ30, Q925K3 | Gene names | Gfap | |||
|
Domain Architecture |

|
|||||
| Description | Glial fibrillary acidic protein, astrocyte (GFAP). | |||||
|
TTC28_MOUSE
|
||||||
| NC score | 0.012092 (rank : 57) | θ value | 4.03905 (rank : 40) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80XJ3, Q6P9J6, Q80TL5, Q8BV10, Q8C0F2 | Gene names | Ttc28, Kiaa1043 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tetratricopeptide repeat protein 28 (TPR repeat protein 28). | |||||
|
VIME_HUMAN
|
||||||
| NC score | 0.011868 (rank : 58) | θ value | 8.99809 (rank : 59) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 706 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P08670, Q15867, Q15868, Q15869, Q6LER9, Q8N850, Q96ML2, Q9NTM3 | Gene names | VIM | |||
|
Domain Architecture |

|
|||||
| Description | Vimentin. | |||||
|
MMS19_HUMAN
|
||||||
| NC score | 0.011282 (rank : 59) | θ value | 6.88961 (rank : 47) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96T76, Q5T455, Q66K82, Q7L4W8, Q969Z1, Q96DF1, Q96MR1, Q96RK5, Q96SK1, Q9BUE2, Q9BYS9 | Gene names | MMS19L, MMS19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MMS19-like protein (hMMS19) (MET18 homolog). | |||||
|
RBP1_HUMAN
|
||||||
| NC score | 0.010860 (rank : 60) | θ value | 8.99809 (rank : 57) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 521 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q15311 | Gene names | RALBP1, RLIP1, RLIP76 | |||
|
Domain Architecture |

|
|||||
| Description | RalA-binding protein 1 (RalBP1) (Ral-interacting protein 1) (76 kDa Ral-interacting protein) (Dinitrophenyl S-glutathione ATPase) (DNP-SG ATPase). | |||||
|
ANR35_HUMAN
|
||||||
| NC score | 0.010095 (rank : 61) | θ value | 6.88961 (rank : 41) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 784 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8N283, Q3MJ10, Q96LS3 | Gene names | ANKRD35 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 35. | |||||
|
KIFC3_HUMAN
|
||||||
| NC score | 0.008981 (rank : 62) | θ value | 8.99809 (rank : 54) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 619 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9BVG8, O75299, Q8IUT3, Q96HW6 | Gene names | KIFC3 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIFC3. | |||||
|
GP110_MOUSE
|
||||||
| NC score | 0.005227 (rank : 63) | θ value | 6.88961 (rank : 43) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8VEC3, Q8BMV8 | Gene names | Gpr110 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-protein coupled receptor 110 precursor. | |||||
|
ZBT20_HUMAN
|
||||||
| NC score | -0.001937 (rank : 64) | θ value | 6.88961 (rank : 50) | |||
| Query Neighborhood Hits | 59 | Target Neighborhood Hits | 835 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9HC78 | Gene names | ZBTB20, DPZF, ZNF288 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 20 (Zinc finger protein 288) (Dendritic-derived BTB/POZ zinc finger protein). | |||||