Please be patient as the page loads
|
ZSWM5_MOUSE
|
||||||
| SwissProt Accessions | Q80TC6 | Gene names | Zswim5, Kiaa1511 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger SWIM domain-containing protein 5. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
ZSWM4_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.976844 (rank : 4) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H7M6 | Gene names | ZSWIM4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger SWIM domain-containing protein 4. | |||||
|
ZSWM4_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.991319 (rank : 3) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8C7B8, Q8BPN9 | Gene names | Zswim4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger SWIM domain-containing protein 4. | |||||
|
ZSWM5_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.996895 (rank : 2) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9P217, Q5SXQ9 | Gene names | ZSWIM5, KIAA1511 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger SWIM domain-containing protein 5. | |||||
|
ZSWM5_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q80TC6 | Gene names | Zswim5, Kiaa1511 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger SWIM domain-containing protein 5. | |||||
|
CO9A1_MOUSE
|
||||||
| θ value | 0.125558 (rank : 5) | NC score | 0.026484 (rank : 8) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 477 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q05722, Q61269, Q61270, Q61433, Q61940 | Gene names | Col9a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(IX) chain precursor. | |||||
|
CO9A1_HUMAN
|
||||||
| θ value | 0.163984 (rank : 6) | NC score | 0.030461 (rank : 5) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P20849, Q13699, Q13700, Q5TF52, Q6P467, Q96BM8, Q99225, Q9H151, Q9H152, Q9Y6P2, Q9Y6P3 | Gene names | COL9A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(IX) chain precursor. | |||||
|
CO5A2_HUMAN
|
||||||
| θ value | 0.21417 (rank : 7) | NC score | 0.019863 (rank : 10) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 760 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P05997 | Gene names | COL5A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(V) chain precursor. | |||||
|
RABX5_HUMAN
|
||||||
| θ value | 0.813845 (rank : 8) | NC score | 0.028426 (rank : 6) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UJ41 | Gene names | RABGEF1, RABEX5 | |||
|
Domain Architecture |

|
|||||
| Description | Rab5 GDP/GTP exchange factor (Rabex-5) (Rabaptin-5-associated exchange factor for Rab5). | |||||
|
CTDP1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 9) | NC score | 0.026822 (rank : 7) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y5B0, Q7Z644, Q96BZ1, Q9Y6F5 | Gene names | CTDP1, FCP1 | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase II subunit A C-terminal domain phosphatase (EC 3.1.3.16) (TFIIF-associating CTD phosphatase). | |||||
|
RPGF2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 10) | NC score | 0.024413 (rank : 9) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y4G8 | Gene names | RAPGEF2, KIAA0313, NRAPGEP, PDZGEF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rap guanine nucleotide exchange factor 2 (Neural RAP guanine nucleotide exchange protein) (nRap GEP) (PDZ domain-containing guanine nucleotide exchange factor 1) (PDZ-GEF1) (RA-GEF). | |||||
|
TS1R1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 11) | NC score | 0.011720 (rank : 17) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99PG6, Q6NS58, Q923J9, Q925I5, Q99PG5 | Gene names | Tas1r1, Gpr70, T1r1, Tr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Taste receptor type 1 member 1 precursor (G-protein coupled receptor 70). | |||||
|
FHOD1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 12) | NC score | 0.017141 (rank : 13) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6P9Q4, Q8BMK2 | Gene names | Fhod1, Fhos1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FH1/FH2 domain-containing protein (Formin homolog overexpressed in spleen) (FHOS) (Formin homology 2 domain-containing protein 1). | |||||
|
PDZD2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 13) | NC score | 0.010416 (rank : 22) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O15018, Q9BXD4 | Gene names | PDZD2, AIPC, KIAA0300, PDZK3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 2 (PDZ domain-containing protein 3) (Activated in prostate cancer protein). | |||||
|
PER3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 14) | NC score | 0.017260 (rank : 12) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P56645, Q5H8X4, Q969K6, Q96S77, Q96S78, Q9C0J3, Q9NSP9, Q9UGU8 | Gene names | PER3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (hPER3). | |||||
|
SREC_HUMAN
|
||||||
| θ value | 3.0926 (rank : 15) | NC score | 0.005146 (rank : 27) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14162, O43701 | Gene names | SCARF1, KIAA0149, SREC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endothelial cells scavenger receptor precursor (Acetyl LDL receptor) (Scavenger receptor class F member 1). | |||||
|
GEMI5_HUMAN
|
||||||
| θ value | 4.03905 (rank : 16) | NC score | 0.009592 (rank : 24) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8TEQ6, Q8WWV4, Q969W4, Q9H9K3, Q9UFI5 | Gene names | GEMIN5 | |||
|
Domain Architecture |

|
|||||
| Description | Gem-associated protein 5 (Gemin5). | |||||
|
PIAS3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 17) | NC score | 0.017613 (rank : 11) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y6X2, Q9UFI3 | Gene names | PIAS3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein inhibitor of activated STAT protein 3. | |||||
|
PKHG4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 18) | NC score | 0.013446 (rank : 16) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q58EX7, Q4G0J8, Q4H485, Q56A69, Q9H7K4, Q9UFW0 | Gene names | PLEKHG4, PRTPHN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Puratrophin-1 (Pleckstrin homology domain-containing family G member 4) (Purkinje cell atrophy-associated protein 1). | |||||
|
B4GT1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 19) | NC score | 0.013687 (rank : 15) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P15291, Q12909, Q12910, Q12911, Q14456, Q14509, Q14523 | Gene names | B4GALT1, GGTB2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-1,4-galactosyltransferase 1 (EC 2.4.1.-) (Beta-1,4-GalTase 1) (Beta4Gal-T1) (b4Gal-T1) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 1) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 1) [Includes: Lactose synthase A protein (EC 2.4.1.22); N-acetyllactosamine synthase (EC 2.4.1.90) (Nal synthetase); Beta-N-acetylglucosaminylglycopeptide beta-1,4- galactosyltransferase (EC 2.4.1.38); Beta-N-acetylglucosaminyl- glycolipid beta-1,4-galactosyltransferase (EC 2.4.1.-)]. | |||||
|
CO4A2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 20) | NC score | 0.016131 (rank : 14) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 492 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P08122, Q61375 | Gene names | Col4a2 | |||
|
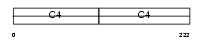
Domain Architecture |
|
|||||
| Description | Collagen alpha-2(IV) chain precursor. | |||||
|
RIMB1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 21) | NC score | 0.005749 (rank : 26) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1479 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O95153, O75111, Q8N5W3 | Gene names | BZRAP1, KIAA0612, RBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripheral-type benzodiazepine receptor-associated protein 1 (PRAX-1) (Peripheral benzodiazepine receptor-interacting protein) (PBR-IP) (RIM-binding protein 1) (RIM-BP1). | |||||
|
ANR47_MOUSE
|
||||||
| θ value | 6.88961 (rank : 22) | NC score | 0.004858 (rank : 28) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Z1P7 | Gene names | Ankrd47, D17Ertd288e, Ng28 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 47. | |||||
|
KPCG_HUMAN
|
||||||
| θ value | 6.88961 (rank : 23) | NC score | 0.001231 (rank : 30) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1126 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P05129 | Gene names | PRKCG, PKCG | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C gamma type (EC 2.7.11.13) (PKC-gamma). | |||||
|
NKX25_MOUSE
|
||||||
| θ value | 6.88961 (rank : 24) | NC score | 0.003428 (rank : 29) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P42582, P97335 | Gene names | Nkx2-5, Csx, Nkx-2.5, Nkx2e | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Nkx-2.5 (Homeobox protein NK-2 homolog E) (Cardiac- specific homeobox) (Homeobox protein CSX). | |||||
|
PO121_MOUSE
|
||||||
| θ value | 6.88961 (rank : 25) | NC score | 0.009615 (rank : 23) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8K3Z9, Q7TSH5 | Gene names | Pom121, Nup121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear envelope pore membrane protein POM 121 (Pore membrane protein of 121 kDa). | |||||
|
SEPT3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 26) | NC score | 0.010887 (rank : 20) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UH03, Q6IBZ6, Q8N3P3 | Gene names | SEPT3, SEP3 | |||
|
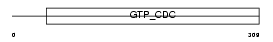
Domain Architecture |
|
|||||
| Description | Neuronal-specific septin-3. | |||||
|
SEPT3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 27) | NC score | 0.011248 (rank : 19) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z1S5 | Gene names | Sept3, Sep3 | |||
|
Domain Architecture |
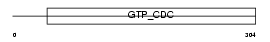
|
|||||
| Description | Neuronal-specific septin-3. | |||||
|
KPCG_MOUSE
|
||||||
| θ value | 8.99809 (rank : 28) | NC score | 0.000830 (rank : 31) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1123 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P63318, P05697 | Gene names | Prkcg, Pkcc, Pkcg, Prkcc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein kinase C gamma type (EC 2.7.11.13) (PKC-gamma). | |||||
|
LSD1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 29) | NC score | 0.008195 (rank : 25) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O60341, Q5TH94, Q5TH95, Q86VT7, Q8IXK4, Q8NDP6, Q8TAZ3, Q96AW4 | Gene names | AOF2, KIAA0601, LSD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lysine-specific histone demethylase 1 (EC 1.-.-.-) (Flavin-containing amine oxidase domain-containing protein 2) (BRAF35-HDAC complex protein BHC110). | |||||
|
PIAS3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 30) | NC score | 0.010575 (rank : 21) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O54714, Q80WF8, Q8R598 | Gene names | Pias3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein inhibitor of activated STAT protein 3. | |||||
|
RM37_MOUSE
|
||||||
| θ value | 8.99809 (rank : 31) | NC score | 0.011552 (rank : 18) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q921S7 | Gene names | Mrpl37 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 39S ribosomal protein L37, mitochondrial precursor (L37mt) (MRP-L37). | |||||
|
ZSWM5_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q80TC6 | Gene names | Zswim5, Kiaa1511 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger SWIM domain-containing protein 5. | |||||
|
ZSWM5_HUMAN
|
||||||
| NC score | 0.996895 (rank : 2) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9P217, Q5SXQ9 | Gene names | ZSWIM5, KIAA1511 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger SWIM domain-containing protein 5. | |||||
|
ZSWM4_MOUSE
|
||||||
| NC score | 0.991319 (rank : 3) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8C7B8, Q8BPN9 | Gene names | Zswim4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger SWIM domain-containing protein 4. | |||||
|
ZSWM4_HUMAN
|
||||||
| NC score | 0.976844 (rank : 4) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H7M6 | Gene names | ZSWIM4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger SWIM domain-containing protein 4. | |||||
|
CO9A1_HUMAN
|
||||||
| NC score | 0.030461 (rank : 5) | θ value | 0.163984 (rank : 6) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P20849, Q13699, Q13700, Q5TF52, Q6P467, Q96BM8, Q99225, Q9H151, Q9H152, Q9Y6P2, Q9Y6P3 | Gene names | COL9A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(IX) chain precursor. | |||||
|
RABX5_HUMAN
|
||||||
| NC score | 0.028426 (rank : 6) | θ value | 0.813845 (rank : 8) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UJ41 | Gene names | RABGEF1, RABEX5 | |||
|
Domain Architecture |

|
|||||
| Description | Rab5 GDP/GTP exchange factor (Rabex-5) (Rabaptin-5-associated exchange factor for Rab5). | |||||
|
CTDP1_HUMAN
|
||||||
| NC score | 0.026822 (rank : 7) | θ value | 1.38821 (rank : 9) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y5B0, Q7Z644, Q96BZ1, Q9Y6F5 | Gene names | CTDP1, FCP1 | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase II subunit A C-terminal domain phosphatase (EC 3.1.3.16) (TFIIF-associating CTD phosphatase). | |||||
|
CO9A1_MOUSE
|
||||||
| NC score | 0.026484 (rank : 8) | θ value | 0.125558 (rank : 5) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 477 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q05722, Q61269, Q61270, Q61433, Q61940 | Gene names | Col9a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(IX) chain precursor. | |||||
|
RPGF2_HUMAN
|
||||||
| NC score | 0.024413 (rank : 9) | θ value | 1.38821 (rank : 10) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y4G8 | Gene names | RAPGEF2, KIAA0313, NRAPGEP, PDZGEF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rap guanine nucleotide exchange factor 2 (Neural RAP guanine nucleotide exchange protein) (nRap GEP) (PDZ domain-containing guanine nucleotide exchange factor 1) (PDZ-GEF1) (RA-GEF). | |||||
|
CO5A2_HUMAN
|
||||||
| NC score | 0.019863 (rank : 10) | θ value | 0.21417 (rank : 7) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 760 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P05997 | Gene names | COL5A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(V) chain precursor. | |||||
|
PIAS3_HUMAN
|
||||||
| NC score | 0.017613 (rank : 11) | θ value | 4.03905 (rank : 17) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y6X2, Q9UFI3 | Gene names | PIAS3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein inhibitor of activated STAT protein 3. | |||||
|
PER3_HUMAN
|
||||||
| NC score | 0.017260 (rank : 12) | θ value | 3.0926 (rank : 14) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P56645, Q5H8X4, Q969K6, Q96S77, Q96S78, Q9C0J3, Q9NSP9, Q9UGU8 | Gene names | PER3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (hPER3). | |||||
|
FHOD1_MOUSE
|
||||||
| NC score | 0.017141 (rank : 13) | θ value | 2.36792 (rank : 12) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6P9Q4, Q8BMK2 | Gene names | Fhod1, Fhos1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FH1/FH2 domain-containing protein (Formin homolog overexpressed in spleen) (FHOS) (Formin homology 2 domain-containing protein 1). | |||||
|
CO4A2_MOUSE
|
||||||
| NC score | 0.016131 (rank : 14) | θ value | 5.27518 (rank : 20) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 492 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P08122, Q61375 | Gene names | Col4a2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(IV) chain precursor. | |||||
|
B4GT1_HUMAN
|
||||||
| NC score | 0.013687 (rank : 15) | θ value | 5.27518 (rank : 19) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P15291, Q12909, Q12910, Q12911, Q14456, Q14509, Q14523 | Gene names | B4GALT1, GGTB2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-1,4-galactosyltransferase 1 (EC 2.4.1.-) (Beta-1,4-GalTase 1) (Beta4Gal-T1) (b4Gal-T1) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 1) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 1) [Includes: Lactose synthase A protein (EC 2.4.1.22); N-acetyllactosamine synthase (EC 2.4.1.90) (Nal synthetase); Beta-N-acetylglucosaminylglycopeptide beta-1,4- galactosyltransferase (EC 2.4.1.38); Beta-N-acetylglucosaminyl- glycolipid beta-1,4-galactosyltransferase (EC 2.4.1.-)]. | |||||
|
PKHG4_HUMAN
|
||||||
| NC score | 0.013446 (rank : 16) | θ value | 4.03905 (rank : 18) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q58EX7, Q4G0J8, Q4H485, Q56A69, Q9H7K4, Q9UFW0 | Gene names | PLEKHG4, PRTPHN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Puratrophin-1 (Pleckstrin homology domain-containing family G member 4) (Purkinje cell atrophy-associated protein 1). | |||||
|
TS1R1_MOUSE
|
||||||
| NC score | 0.011720 (rank : 17) | θ value | 1.38821 (rank : 11) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99PG6, Q6NS58, Q923J9, Q925I5, Q99PG5 | Gene names | Tas1r1, Gpr70, T1r1, Tr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Taste receptor type 1 member 1 precursor (G-protein coupled receptor 70). | |||||
|
RM37_MOUSE
|
||||||
| NC score | 0.011552 (rank : 18) | θ value | 8.99809 (rank : 31) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q921S7 | Gene names | Mrpl37 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 39S ribosomal protein L37, mitochondrial precursor (L37mt) (MRP-L37). | |||||
|
SEPT3_MOUSE
|
||||||
| NC score | 0.011248 (rank : 19) | θ value | 6.88961 (rank : 27) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z1S5 | Gene names | Sept3, Sep3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal-specific septin-3. | |||||
|
SEPT3_HUMAN
|
||||||
| NC score | 0.010887 (rank : 20) | θ value | 6.88961 (rank : 26) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UH03, Q6IBZ6, Q8N3P3 | Gene names | SEPT3, SEP3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal-specific septin-3. | |||||
|
PIAS3_MOUSE
|
||||||
| NC score | 0.010575 (rank : 21) | θ value | 8.99809 (rank : 30) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O54714, Q80WF8, Q8R598 | Gene names | Pias3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein inhibitor of activated STAT protein 3. | |||||
|
PDZD2_HUMAN
|
||||||
| NC score | 0.010416 (rank : 22) | θ value | 3.0926 (rank : 13) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O15018, Q9BXD4 | Gene names | PDZD2, AIPC, KIAA0300, PDZK3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 2 (PDZ domain-containing protein 3) (Activated in prostate cancer protein). | |||||
|
PO121_MOUSE
|
||||||
| NC score | 0.009615 (rank : 23) | θ value | 6.88961 (rank : 25) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8K3Z9, Q7TSH5 | Gene names | Pom121, Nup121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear envelope pore membrane protein POM 121 (Pore membrane protein of 121 kDa). | |||||
|
GEMI5_HUMAN
|
||||||
| NC score | 0.009592 (rank : 24) | θ value | 4.03905 (rank : 16) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8TEQ6, Q8WWV4, Q969W4, Q9H9K3, Q9UFI5 | Gene names | GEMIN5 | |||
|
Domain Architecture |

|
|||||
| Description | Gem-associated protein 5 (Gemin5). | |||||
|
LSD1_HUMAN
|
||||||
| NC score | 0.008195 (rank : 25) | θ value | 8.99809 (rank : 29) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O60341, Q5TH94, Q5TH95, Q86VT7, Q8IXK4, Q8NDP6, Q8TAZ3, Q96AW4 | Gene names | AOF2, KIAA0601, LSD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lysine-specific histone demethylase 1 (EC 1.-.-.-) (Flavin-containing amine oxidase domain-containing protein 2) (BRAF35-HDAC complex protein BHC110). | |||||
|
RIMB1_HUMAN
|
||||||
| NC score | 0.005749 (rank : 26) | θ value | 5.27518 (rank : 21) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1479 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O95153, O75111, Q8N5W3 | Gene names | BZRAP1, KIAA0612, RBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripheral-type benzodiazepine receptor-associated protein 1 (PRAX-1) (Peripheral benzodiazepine receptor-interacting protein) (PBR-IP) (RIM-binding protein 1) (RIM-BP1). | |||||
|
SREC_HUMAN
|
||||||
| NC score | 0.005146 (rank : 27) | θ value | 3.0926 (rank : 15) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14162, O43701 | Gene names | SCARF1, KIAA0149, SREC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endothelial cells scavenger receptor precursor (Acetyl LDL receptor) (Scavenger receptor class F member 1). | |||||
|
ANR47_MOUSE
|
||||||
| NC score | 0.004858 (rank : 28) | θ value | 6.88961 (rank : 22) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Z1P7 | Gene names | Ankrd47, D17Ertd288e, Ng28 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 47. | |||||
|
NKX25_MOUSE
|
||||||
| NC score | 0.003428 (rank : 29) | θ value | 6.88961 (rank : 24) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P42582, P97335 | Gene names | Nkx2-5, Csx, Nkx-2.5, Nkx2e | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Nkx-2.5 (Homeobox protein NK-2 homolog E) (Cardiac- specific homeobox) (Homeobox protein CSX). | |||||
|
KPCG_HUMAN
|
||||||
| NC score | 0.001231 (rank : 30) | θ value | 6.88961 (rank : 23) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1126 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P05129 | Gene names | PRKCG, PKCG | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C gamma type (EC 2.7.11.13) (PKC-gamma). | |||||
|
KPCG_MOUSE
|
||||||
| NC score | 0.000830 (rank : 31) | θ value | 8.99809 (rank : 28) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1123 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P63318, P05697 | Gene names | Prkcg, Pkcc, Pkcg, Prkcc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein kinase C gamma type (EC 2.7.11.13) (PKC-gamma). | |||||