Please be patient as the page loads
|
GRIN2_HUMAN
|
||||||
| SwissProt Accessions | O60269, Q5SVF0 | Gene names | GPRIN2, KIAA0514 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 2 (GRIN2). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
GRIN2_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O60269, Q5SVF0 | Gene names | GPRIN2, KIAA0514 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 2 (GRIN2). | |||||
|
GRIN1_HUMAN
|
||||||
| θ value | 8.65492e-19 (rank : 2) | NC score | 0.433329 (rank : 5) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q7Z2K8, Q8ND74, Q96PZ4 | Gene names | GPRIN1, KIAA1893 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
GRIN1_MOUSE
|
||||||
| θ value | 1.92812e-18 (rank : 3) | NC score | 0.550518 (rank : 3) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q3UNH4, Q6PGF9, Q9QZY2 | Gene names | Gprin1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
K2027_MOUSE
|
||||||
| θ value | 2.43908e-13 (rank : 4) | NC score | 0.551388 (rank : 2) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BWS5, Q69Z30 | Gene names | Kiaa2027 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA2027. | |||||
|
K2027_HUMAN
|
||||||
| θ value | 2.0648e-12 (rank : 5) | NC score | 0.545948 (rank : 4) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q6ZVF9, Q8IVE4 | Gene names | KIAA2027 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA2027. | |||||
|
CAC1H_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 6) | NC score | 0.041016 (rank : 40) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O95180, O95802, Q8WWI6, Q96QI6, Q96RZ9, Q9NYY4, Q9NYY5 | Gene names | CACNA1H | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1H (Voltage- gated calcium channel subunit alpha Cav3.2) (Low-voltage-activated calcium channel alpha1 3.2 subunit). | |||||
|
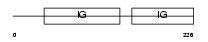
ROBO4_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 7) | NC score | 0.038900 (rank : 41) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 358 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8C310, Q9DBW1 | Gene names | Robo4 | |||
|
Domain Architecture |
|
|||||
| Description | Roundabout homolog 4 precursor. | |||||
|
MUC1_HUMAN
|
||||||
| θ value | 0.163984 (rank : 8) | NC score | 0.052739 (rank : 32) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
HORN_MOUSE
|
||||||
| θ value | 0.21417 (rank : 9) | NC score | 0.058526 (rank : 25) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8VHD8 | Gene names | Hrnr | |||
|
Domain Architecture |

|
|||||
| Description | Hornerin. | |||||
|
GON4L_MOUSE
|
||||||
| θ value | 0.365318 (rank : 10) | NC score | 0.050073 (rank : 36) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9DB00, Q80TB4, Q91YI9 | Gene names | Gon4l, Gon4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
SYN2_MOUSE
|
||||||
| θ value | 0.365318 (rank : 11) | NC score | 0.066575 (rank : 17) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q64332, Q6NZR0, Q9QWV7 | Gene names | Syn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synapsin-2 (Synapsin II). | |||||
|
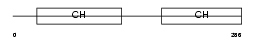
CLMN_MOUSE
|
||||||
| θ value | 0.47712 (rank : 12) | NC score | 0.032460 (rank : 45) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8C5W0, Q91V71, Q91XT7, Q91XT8, Q91XU9 | Gene names | Clmn | |||
|
Domain Architecture |
|
|||||
| Description | Calmin. | |||||
|
MN1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 13) | NC score | 0.094686 (rank : 8) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q10571 | Gene names | MN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable tumor suppressor protein MN1. | |||||
|
K1543_HUMAN
|
||||||
| θ value | 0.813845 (rank : 14) | NC score | 0.058039 (rank : 26) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9P1Y5, Q8NDF1 | Gene names | KIAA1543 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1543. | |||||
|
MY18B_HUMAN
|
||||||
| θ value | 1.06291 (rank : 15) | NC score | 0.022902 (rank : 52) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1299 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8IUG5, Q8NDI8, Q8TE65, Q8WWS0, Q96KH2, Q96KR8, Q96KR9 | Gene names | MYO18B | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18B (Myosin XVIIIb). | |||||
|
SCAP_HUMAN
|
||||||
| θ value | 1.38821 (rank : 16) | NC score | 0.036753 (rank : 42) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q12770, Q8N2E0, Q8WUA1 | Gene names | SCAP, KIAA0199 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein cleavage-activating protein (SREBP cleavage-activating protein) (SCAP). | |||||
|
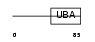
UBP2L_HUMAN
|
||||||
| θ value | 1.38821 (rank : 17) | NC score | 0.065045 (rank : 19) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q14157, Q5VU75, Q5VU76, Q9BTU3, Q9UGL5 | Gene names | UBAP2L, KIAA0144, NICE4 | |||
|
Domain Architecture |
|
|||||
| Description | Ubiquitin-associated protein 2-like (Protein NICE-4). | |||||
|
K0999_HUMAN
|
||||||
| θ value | 1.81305 (rank : 18) | NC score | 0.004608 (rank : 66) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 899 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y2K2, Q59FY2, Q5M9N1, Q6P3R6, Q8IYM8, Q9HA50 | Gene names | KIAA0999 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized serine/threonine-protein kinase KIAA0999 (EC 2.7.11.1). | |||||
|
TARA_HUMAN
|
||||||
| θ value | 1.81305 (rank : 19) | NC score | 0.053498 (rank : 31) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
TCGAP_HUMAN
|
||||||
| θ value | 1.81305 (rank : 20) | NC score | 0.028546 (rank : 48) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 872 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O14559, O14552, O14560, Q6ZSP6, Q96CP3, Q9NT23 | Gene names | SNX26, TCGAP | |||
|
Domain Architecture |

|
|||||
| Description | TC10/CDC42 GTPase-activating protein (Sorting nexin-26). | |||||
|
TCOF_MOUSE
|
||||||
| θ value | 1.81305 (rank : 21) | NC score | 0.101421 (rank : 7) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O08784, O08857 | Gene names | Tcof1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein homolog). | |||||
|
TMPSD_MOUSE
|
||||||
| θ value | 1.81305 (rank : 22) | NC score | 0.015828 (rank : 63) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 334 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5U405, Q8CFE0, Q91VQ8 | Gene names | Tmprss13, Msp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protease, serine 13 (EC 3.4.21.-) (Mosaic serine protease) (Membrane-type mosaic serine protease). | |||||
|
TULP4_HUMAN
|
||||||
| θ value | 1.81305 (rank : 23) | NC score | 0.030940 (rank : 46) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 426 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NRJ4, Q9HD22, Q9P2F0 | Gene names | TULP4, KIAA1397, TUSP | |||
|
Domain Architecture |

|
|||||
| Description | Tubby-like protein 4 (Tubby superfamily protein). | |||||
|
KCNH6_HUMAN
|
||||||
| θ value | 2.36792 (rank : 24) | NC score | 0.016566 (rank : 61) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H252, Q9BRD7 | Gene names | KCNH6, ERG2, HERG2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 6 (Voltage-gated potassium channel subunit Kv11.2) (Ether-a-go-go-related gene potassium channel 2) (Ether-a-go-go-related protein 2) (Eag-related protein 2). | |||||
|
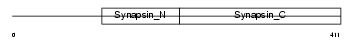
SYN1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 25) | NC score | 0.072659 (rank : 13) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 363 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O88935, Q62279, Q8QZT8 | Gene names | Syn1, Syn-1 | |||
|
Domain Architecture |
|
|||||
| Description | Synapsin-1 (Synapsin I). | |||||
|
TAF4_HUMAN
|
||||||
| θ value | 2.36792 (rank : 26) | NC score | 0.073573 (rank : 12) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O00268, Q5TBP6, Q99721, Q9BR40, Q9BX42 | Gene names | TAF4, TAF2C, TAF2C1, TAF4A, TAFII130, TAFII135 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 4 (TBP-associated factor 4) (Transcription initiation factor TFIID 135 kDa subunit) (TAF(II)135) (TAFII-135) (TAFII135) (TAFII-130) (TAFII130). | |||||
|
BCAS1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 27) | NC score | 0.053994 (rank : 30) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q80YN3, Q9CVA1 | Gene names | Bcas1, Nabc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Breast carcinoma amplified sequence 1 homolog (Novel amplified in breast cancer 1 homolog). | |||||
|
BRWD1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 28) | NC score | 0.020670 (rank : 57) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 448 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NSI6, O43721, Q5R2V0, Q5R2V1, Q8TCV3, Q96QG9, Q96QH0, Q9NUK1 | Gene names | BRWD1, C21orf107, WDR9 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain and WD repeat domain-containing protein 1 (WD repeat protein 9). | |||||
|
DLGP1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 29) | NC score | 0.022918 (rank : 51) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O14490, O14489, P78335 | Gene names | DLGAP1, DAP1, GKAP | |||
|
Domain Architecture |

|
|||||
| Description | Disks large-associated protein 1 (DAP-1) (Guanylate kinase-associated protein) (hGKAP) (SAP90/PSD-95-associated protein 1) (SAPAP1) (PSD- 95/SAP90-binding protein 1). | |||||
|
DOK7_MOUSE
|
||||||
| θ value | 3.0926 (rank : 30) | NC score | 0.044534 (rank : 38) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q18PE0, Q3TCZ6, Q5FW70, Q8C8U7 | Gene names | Dok7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Dok-7 (Downstream of tyrosine kinase 7). | |||||
|
NTKL_MOUSE
|
||||||
| θ value | 3.0926 (rank : 31) | NC score | 0.022755 (rank : 53) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9EQC5, Q3TST7, Q3UKU0, Q8K222 | Gene names | Scyl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-terminal kinase-like protein (SCY1-like protein 1) (105 kDa kinase- like protein) (Mitosis-associated kinase-like protein NTKL). | |||||
|
PHC1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 32) | NC score | 0.051432 (rank : 34) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q64028, P70359, Q64307, Q8BZ80 | Gene names | Phc1, Edr, Edr1, Rae28 | |||
|
Domain Architecture |

|
|||||
| Description | Polyhomeotic-like protein 1 (mPH1) (Early development regulatory protein 1) (RAE-28). | |||||
|
SYN3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 33) | NC score | 0.064919 (rank : 20) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O14994 | Gene names | SYN3 | |||
|
Domain Architecture |
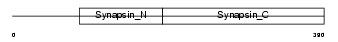
|
|||||
| Description | Synapsin-3 (Synapsin III). | |||||
|
ASTN_MOUSE
|
||||||
| θ value | 4.03905 (rank : 34) | NC score | 0.025805 (rank : 49) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q61137, Q505A0, Q7TQG3, Q8CHH2 | Gene names | Astn, Astn1, Kiaa0289 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Astrotactin-1 precursor (Neuronal migration protein GC14). | |||||
|
BSN_MOUSE
|
||||||
| θ value | 4.03905 (rank : 35) | NC score | 0.043923 (rank : 39) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
CBX2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 36) | NC score | 0.030318 (rank : 47) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P30658, O35731, Q8CIA0 | Gene names | Cbx2, M33 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 2 (Modifier 3 protein) (M33). | |||||
|
PHF12_MOUSE
|
||||||
| θ value | 4.03905 (rank : 37) | NC score | 0.023234 (rank : 50) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5SPL2, Q5SPL3, Q6ZPN8, Q80W50 | Gene names | Phf12, Kiaa1523 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 12 (PHD factor 1) (Pf1). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 38) | NC score | 0.067530 (rank : 15) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
CO1A1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 39) | NC score | 0.033424 (rank : 44) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P11087, Q53WT0, Q60635, Q61367, Q61427, Q63919, Q6PCL3, Q810J9 | Gene names | Col1a1, Cola1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(I) chain precursor. | |||||
|
CS007_MOUSE
|
||||||
| θ value | 5.27518 (rank : 40) | NC score | 0.045466 (rank : 37) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 464 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q6ZPZ3 | Gene names | Kiaa1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7 homolog. | |||||
|
HXB5_HUMAN
|
||||||
| θ value | 5.27518 (rank : 41) | NC score | 0.006179 (rank : 65) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P09067, P09069 | Gene names | HOXB5, HOX2A | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-B5 (Hox-2A) (HHO.C10) (HU-1). | |||||
|
NFAC2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 42) | NC score | 0.021641 (rank : 54) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q60591, Q60984, Q60985 | Gene names | Nfatc2, Nfat1, Nfatp | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor of activated T-cells, cytoplasmic 2 (T cell transcription factor NFAT1) (NFAT pre-existing subunit) (NF-ATp). | |||||
|
NOLC1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 43) | NC score | 0.080777 (rank : 9) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
PHC2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 44) | NC score | 0.034851 (rank : 43) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9QWH1, O88463, Q8K5D9 | Gene names | Phc2, Edr2, Ph2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyhomeotic-like protein 2 (mPH2) (Early development regulatory protein 2) (p36). | |||||
|
TMPSD_HUMAN
|
||||||
| θ value | 5.27518 (rank : 45) | NC score | 0.012112 (rank : 64) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9BYE2, Q86YM4, Q96JY8, Q9BYE1 | Gene names | TMPRSS13, MSP, TMPRSS11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protease, serine 13 (EC 3.4.21.-) (Mosaic serine protease) (Membrane-type mosaic serine protease). | |||||
|
ZSWM5_HUMAN
|
||||||
| θ value | 5.27518 (rank : 46) | NC score | 0.019124 (rank : 59) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9P217, Q5SXQ9 | Gene names | ZSWIM5, KIAA1511 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger SWIM domain-containing protein 5. | |||||
|
BRWD1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 47) | NC score | 0.019539 (rank : 58) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 478 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q921C3, Q921C2 | Gene names | Brwd1, Wdr9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain and WD repeat domain-containing protein 1 (WD repeat protein 9). | |||||
|
GNAS1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 48) | NC score | 0.020899 (rank : 55) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 651 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6R0H7, Q6R0H4, Q6R0H5, Q6R2J5, Q9JJX0, Q9Z1N8 | Gene names | Gnas, Gnas1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Guanine nucleotide-binding protein G(s) subunit alpha isoforms XLas (Adenylate cyclase-stimulating G alpha protein) (Extra large alphas protein) (XLalphas). | |||||
|
HRBL_MOUSE
|
||||||
| θ value | 6.88961 (rank : 49) | NC score | 0.020674 (rank : 56) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q80WC7, Q8BKS5, Q99J67 | Gene names | Hrbl, Rabr | |||
|
Domain Architecture |
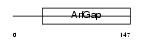
|
|||||
| Description | HIV-1 Rev-binding protein-like protein (Rev/Rex activation domain- binding protein related) (RAB-R). | |||||
|
MAST2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 50) | NC score | 0.002780 (rank : 67) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1110 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q60592, Q6P9M1, Q8CHD1 | Gene names | Mast2, Mast205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 2 (EC 2.7.11.1). | |||||
|
UBP2L_MOUSE
|
||||||
| θ value | 6.88961 (rank : 51) | NC score | 0.055184 (rank : 29) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q80X50, Q8BIT6, Q8BIW4, Q8BJ01, Q8CIG7, Q8K102, Q9CRT6 | Gene names | Ubap2l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-associated protein 2-like. | |||||
|
MEGF9_MOUSE
|
||||||
| θ value | 8.99809 (rank : 52) | NC score | 0.017621 (rank : 60) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BH27, Q8BWI4 | Gene names | Megf9, Egfl5, Kiaa0818 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 9 precursor (EGF-like domain-containing protein 5) (Multiple EGF-like domain protein 5). | |||||
|
ZFY28_HUMAN
|
||||||
| θ value | 8.99809 (rank : 53) | NC score | 0.015936 (rank : 62) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9HCC9 | Gene names | ZFYVE28, KIAA1643 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger FYVE domain-containing protein 28. | |||||
|
BASP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 54) | NC score | 0.051101 (rank : 35) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
BASP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 55) | NC score | 0.051648 (rank : 33) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
GGN_MOUSE
|
||||||
| θ value | θ > 10 (rank : 56) | NC score | 0.055356 (rank : 28) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q80WJ1, Q5EBP4, Q80WI9, Q80WJ0 | Gene names | Ggn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin. | |||||
|
MARCS_HUMAN
|
||||||
| θ value | θ > 10 (rank : 57) | NC score | 0.067396 (rank : 16) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P29966, Q2LA83, Q5TDB7 | Gene names | MARCKS, MACS, PRKCSL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS) (Protein kinase C substrate, 80 kDa protein, light chain) (PKCSL) (80K-L protein). | |||||
|
MARCS_MOUSE
|
||||||
| θ value | θ > 10 (rank : 58) | NC score | 0.060334 (rank : 23) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P26645 | Gene names | Marcks, Macs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS). | |||||
|
MUC13_MOUSE
|
||||||
| θ value | θ > 10 (rank : 59) | NC score | 0.061280 (rank : 22) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P19467 | Gene names | Muc13, Ly64 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-13 precursor (Cell surface antigen 114/A10) (Lymphocyte antigen 64). | |||||
|
MUC1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 60) | NC score | 0.079901 (rank : 10) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q02496 | Gene names | Muc1, Muc-1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (Polymorphic epithelial mucin) (PEMT) (Episialin). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | θ > 10 (rank : 61) | NC score | 0.059810 (rank : 24) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
PRG4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 62) | NC score | 0.075455 (rank : 11) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
PRG4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 63) | NC score | 0.066181 (rank : 18) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9JM99, Q3UEL1, Q3V198 | Gene names | Prg4, Msf, Szp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 64) | NC score | 0.062200 (rank : 21) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
SYN1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 65) | NC score | 0.067554 (rank : 14) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P17600, O75825, Q5H9A9 | Gene names | SYN1 | |||
|
Domain Architecture |

|
|||||
| Description | Synapsin-1 (Synapsin I) (Brain protein 4.1). | |||||
|
TCOF_HUMAN
|
||||||
| θ value | θ > 10 (rank : 66) | NC score | 0.104759 (rank : 6) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q13428, Q99408, Q99860 | Gene names | TCOF1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein). | |||||
|
ZN207_HUMAN
|
||||||
| θ value | θ > 10 (rank : 67) | NC score | 0.056611 (rank : 27) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O43670, Q96HW5, Q9BUQ7 | Gene names | ZNF207 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 207. | |||||
|
GRIN2_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | O60269, Q5SVF0 | Gene names | GPRIN2, KIAA0514 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 2 (GRIN2). | |||||
|
K2027_MOUSE
|
||||||
| NC score | 0.551388 (rank : 2) | θ value | 2.43908e-13 (rank : 4) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BWS5, Q69Z30 | Gene names | Kiaa2027 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA2027. | |||||
|
GRIN1_MOUSE
|
||||||
| NC score | 0.550518 (rank : 3) | θ value | 1.92812e-18 (rank : 3) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q3UNH4, Q6PGF9, Q9QZY2 | Gene names | Gprin1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
K2027_HUMAN
|
||||||
| NC score | 0.545948 (rank : 4) | θ value | 2.0648e-12 (rank : 5) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q6ZVF9, Q8IVE4 | Gene names | KIAA2027 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA2027. | |||||
|
GRIN1_HUMAN
|
||||||
| NC score | 0.433329 (rank : 5) | θ value | 8.65492e-19 (rank : 2) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q7Z2K8, Q8ND74, Q96PZ4 | Gene names | GPRIN1, KIAA1893 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
TCOF_HUMAN
|
||||||
| NC score | 0.104759 (rank : 6) | θ value | θ > 10 (rank : 66) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q13428, Q99408, Q99860 | Gene names | TCOF1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein). | |||||
|
TCOF_MOUSE
|
||||||
| NC score | 0.101421 (rank : 7) | θ value | 1.81305 (rank : 21) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O08784, O08857 | Gene names | Tcof1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein homolog). | |||||
|
MN1_HUMAN
|
||||||
| NC score | 0.094686 (rank : 8) | θ value | 0.62314 (rank : 13) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q10571 | Gene names | MN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable tumor suppressor protein MN1. | |||||
|
NOLC1_HUMAN
|
||||||
| NC score | 0.080777 (rank : 9) | θ value | 5.27518 (rank : 43) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
MUC1_MOUSE
|
||||||
| NC score | 0.079901 (rank : 10) | θ value | θ > 10 (rank : 60) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q02496 | Gene names | Muc1, Muc-1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (Polymorphic epithelial mucin) (PEMT) (Episialin). | |||||
|
PRG4_HUMAN
|
||||||
| NC score | 0.075455 (rank : 11) | θ value | θ > 10 (rank : 62) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
TAF4_HUMAN
|
||||||
| NC score | 0.073573 (rank : 12) | θ value | 2.36792 (rank : 26) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O00268, Q5TBP6, Q99721, Q9BR40, Q9BX42 | Gene names | TAF4, TAF2C, TAF2C1, TAF4A, TAFII130, TAFII135 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 4 (TBP-associated factor 4) (Transcription initiation factor TFIID 135 kDa subunit) (TAF(II)135) (TAFII-135) (TAFII135) (TAFII-130) (TAFII130). | |||||
|
SYN1_MOUSE
|
||||||
| NC score | 0.072659 (rank : 13) | θ value | 2.36792 (rank : 25) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 363 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O88935, Q62279, Q8QZT8 | Gene names | Syn1, Syn-1 | |||
|
Domain Architecture |

|
|||||
| Description | Synapsin-1 (Synapsin I). | |||||
|
SYN1_HUMAN
|
||||||
| NC score | 0.067554 (rank : 14) | θ value | θ > 10 (rank : 65) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P17600, O75825, Q5H9A9 | Gene names | SYN1 | |||
|
Domain Architecture |

|
|||||
| Description | Synapsin-1 (Synapsin I) (Brain protein 4.1). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.067530 (rank : 15) | θ value | 4.03905 (rank : 38) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
MARCS_HUMAN
|
||||||
| NC score | 0.067396 (rank : 16) | θ value | θ > 10 (rank : 57) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P29966, Q2LA83, Q5TDB7 | Gene names | MARCKS, MACS, PRKCSL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS) (Protein kinase C substrate, 80 kDa protein, light chain) (PKCSL) (80K-L protein). | |||||
|
SYN2_MOUSE
|
||||||
| NC score | 0.066575 (rank : 17) | θ value | 0.365318 (rank : 11) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q64332, Q6NZR0, Q9QWV7 | Gene names | Syn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synapsin-2 (Synapsin II). | |||||
|
PRG4_MOUSE
|
||||||
| NC score | 0.066181 (rank : 18) | θ value | θ > 10 (rank : 63) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9JM99, Q3UEL1, Q3V198 | Gene names | Prg4, Msf, Szp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
UBP2L_HUMAN
|
||||||
| NC score | 0.065045 (rank : 19) | θ value | 1.38821 (rank : 17) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q14157, Q5VU75, Q5VU76, Q9BTU3, Q9UGL5 | Gene names | UBAP2L, KIAA0144, NICE4 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-associated protein 2-like (Protein NICE-4). | |||||
|
SYN3_HUMAN
|
||||||
| NC score | 0.064919 (rank : 20) | θ value | 3.0926 (rank : 33) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O14994 | Gene names | SYN3 | |||
|
Domain Architecture |

|
|||||
| Description | Synapsin-3 (Synapsin III). | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.062200 (rank : 21) | θ value | θ > 10 (rank : 64) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
MUC13_MOUSE
|
||||||
| NC score | 0.061280 (rank : 22) | θ value | θ > 10 (rank : 59) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P19467 | Gene names | Muc13, Ly64 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-13 precursor (Cell surface antigen 114/A10) (Lymphocyte antigen 64). | |||||
|
MARCS_MOUSE
|
||||||
| NC score | 0.060334 (rank : 23) | θ value | θ > 10 (rank : 58) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P26645 | Gene names | Marcks, Macs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS). | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.059810 (rank : 24) | θ value | θ > 10 (rank : 61) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
HORN_MOUSE
|
||||||
| NC score | 0.058526 (rank : 25) | θ value | 0.21417 (rank : 9) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8VHD8 | Gene names | Hrnr | |||
|
Domain Architecture |

|
|||||
| Description | Hornerin. | |||||
|
K1543_HUMAN
|
||||||
| NC score | 0.058039 (rank : 26) | θ value | 0.813845 (rank : 14) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9P1Y5, Q8NDF1 | Gene names | KIAA1543 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1543. | |||||
|
ZN207_HUMAN
|
||||||
| NC score | 0.056611 (rank : 27) | θ value | θ > 10 (rank : 67) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O43670, Q96HW5, Q9BUQ7 | Gene names | ZNF207 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 207. | |||||
|
GGN_MOUSE
|
||||||
| NC score | 0.055356 (rank : 28) | θ value | θ > 10 (rank : 56) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q80WJ1, Q5EBP4, Q80WI9, Q80WJ0 | Gene names | Ggn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin. | |||||
|
UBP2L_MOUSE
|
||||||
| NC score | 0.055184 (rank : 29) | θ value | 6.88961 (rank : 51) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q80X50, Q8BIT6, Q8BIW4, Q8BJ01, Q8CIG7, Q8K102, Q9CRT6 | Gene names | Ubap2l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-associated protein 2-like. | |||||
|
BCAS1_MOUSE
|
||||||
| NC score | 0.053994 (rank : 30) | θ value | 3.0926 (rank : 27) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q80YN3, Q9CVA1 | Gene names | Bcas1, Nabc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Breast carcinoma amplified sequence 1 homolog (Novel amplified in breast cancer 1 homolog). | |||||
|
TARA_HUMAN
|
||||||
| NC score | 0.053498 (rank : 31) | θ value | 1.81305 (rank : 19) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
MUC1_HUMAN
|
||||||
| NC score | 0.052739 (rank : 32) | θ value | 0.163984 (rank : 8) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
BASP_MOUSE
|
||||||
| NC score | 0.051648 (rank : 33) | θ value | θ > 10 (rank : 55) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
PHC1_MOUSE
|
||||||
| NC score | 0.051432 (rank : 34) | θ value | 3.0926 (rank : 32) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q64028, P70359, Q64307, Q8BZ80 | Gene names | Phc1, Edr, Edr1, Rae28 | |||
|
Domain Architecture |

|
|||||
| Description | Polyhomeotic-like protein 1 (mPH1) (Early development regulatory protein 1) (RAE-28). | |||||
|
BASP_HUMAN
|
||||||
| NC score | 0.051101 (rank : 35) | θ value | θ > 10 (rank : 54) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
GON4L_MOUSE
|
||||||
| NC score | 0.050073 (rank : 36) | θ value | 0.365318 (rank : 10) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9DB00, Q80TB4, Q91YI9 | Gene names | Gon4l, Gon4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
CS007_MOUSE
|
||||||
| NC score | 0.045466 (rank : 37) | θ value | 5.27518 (rank : 40) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 464 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q6ZPZ3 | Gene names | Kiaa1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7 homolog. | |||||
|
DOK7_MOUSE
|
||||||
| NC score | 0.044534 (rank : 38) | θ value | 3.0926 (rank : 30) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q18PE0, Q3TCZ6, Q5FW70, Q8C8U7 | Gene names | Dok7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Dok-7 (Downstream of tyrosine kinase 7). | |||||
|
BSN_MOUSE
|
||||||
| NC score | 0.043923 (rank : 39) | θ value | 4.03905 (rank : 35) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
CAC1H_HUMAN
|
||||||
| NC score | 0.041016 (rank : 40) | θ value | 0.0431538 (rank : 6) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O95180, O95802, Q8WWI6, Q96QI6, Q96RZ9, Q9NYY4, Q9NYY5 | Gene names | CACNA1H | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1H (Voltage- gated calcium channel subunit alpha Cav3.2) (Low-voltage-activated calcium channel alpha1 3.2 subunit). | |||||
|
ROBO4_MOUSE
|
||||||
| NC score | 0.038900 (rank : 41) | θ value | 0.0736092 (rank : 7) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 358 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8C310, Q9DBW1 | Gene names | Robo4 | |||
|
Domain Architecture |

|
|||||
| Description | Roundabout homolog 4 precursor. | |||||
|
SCAP_HUMAN
|
||||||
| NC score | 0.036753 (rank : 42) | θ value | 1.38821 (rank : 16) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q12770, Q8N2E0, Q8WUA1 | Gene names | SCAP, KIAA0199 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein cleavage-activating protein (SREBP cleavage-activating protein) (SCAP). | |||||
|
PHC2_MOUSE
|
||||||
| NC score | 0.034851 (rank : 43) | θ value | 5.27518 (rank : 44) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9QWH1, O88463, Q8K5D9 | Gene names | Phc2, Edr2, Ph2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyhomeotic-like protein 2 (mPH2) (Early development regulatory protein 2) (p36). | |||||
|
CO1A1_MOUSE
|
||||||
| NC score | 0.033424 (rank : 44) | θ value | 5.27518 (rank : 39) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P11087, Q53WT0, Q60635, Q61367, Q61427, Q63919, Q6PCL3, Q810J9 | Gene names | Col1a1, Cola1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(I) chain precursor. | |||||
|
CLMN_MOUSE
|
||||||
| NC score | 0.032460 (rank : 45) | θ value | 0.47712 (rank : 12) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8C5W0, Q91V71, Q91XT7, Q91XT8, Q91XU9 | Gene names | Clmn | |||
|
Domain Architecture |

|
|||||
| Description | Calmin. | |||||
|
TULP4_HUMAN
|
||||||
| NC score | 0.030940 (rank : 46) | θ value | 1.81305 (rank : 23) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 426 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NRJ4, Q9HD22, Q9P2F0 | Gene names | TULP4, KIAA1397, TUSP | |||
|
Domain Architecture |

|
|||||
| Description | Tubby-like protein 4 (Tubby superfamily protein). | |||||
|
CBX2_MOUSE
|
||||||
| NC score | 0.030318 (rank : 47) | θ value | 4.03905 (rank : 36) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P30658, O35731, Q8CIA0 | Gene names | Cbx2, M33 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 2 (Modifier 3 protein) (M33). | |||||
|
TCGAP_HUMAN
|
||||||
| NC score | 0.028546 (rank : 48) | θ value | 1.81305 (rank : 20) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 872 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O14559, O14552, O14560, Q6ZSP6, Q96CP3, Q9NT23 | Gene names | SNX26, TCGAP | |||
|
Domain Architecture |

|
|||||
| Description | TC10/CDC42 GTPase-activating protein (Sorting nexin-26). | |||||
|
ASTN_MOUSE
|
||||||
| NC score | 0.025805 (rank : 49) | θ value | 4.03905 (rank : 34) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q61137, Q505A0, Q7TQG3, Q8CHH2 | Gene names | Astn, Astn1, Kiaa0289 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Astrotactin-1 precursor (Neuronal migration protein GC14). | |||||
|
PHF12_MOUSE
|
||||||
| NC score | 0.023234 (rank : 50) | θ value | 4.03905 (rank : 37) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5SPL2, Q5SPL3, Q6ZPN8, Q80W50 | Gene names | Phf12, Kiaa1523 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 12 (PHD factor 1) (Pf1). | |||||
|
DLGP1_HUMAN
|
||||||
| NC score | 0.022918 (rank : 51) | θ value | 3.0926 (rank : 29) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O14490, O14489, P78335 | Gene names | DLGAP1, DAP1, GKAP | |||
|
Domain Architecture |

|
|||||
| Description | Disks large-associated protein 1 (DAP-1) (Guanylate kinase-associated protein) (hGKAP) (SAP90/PSD-95-associated protein 1) (SAPAP1) (PSD- 95/SAP90-binding protein 1). | |||||
|
MY18B_HUMAN
|
||||||
| NC score | 0.022902 (rank : 52) | θ value | 1.06291 (rank : 15) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1299 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8IUG5, Q8NDI8, Q8TE65, Q8WWS0, Q96KH2, Q96KR8, Q96KR9 | Gene names | MYO18B | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18B (Myosin XVIIIb). | |||||
|
NTKL_MOUSE
|
||||||
| NC score | 0.022755 (rank : 53) | θ value | 3.0926 (rank : 31) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9EQC5, Q3TST7, Q3UKU0, Q8K222 | Gene names | Scyl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-terminal kinase-like protein (SCY1-like protein 1) (105 kDa kinase- like protein) (Mitosis-associated kinase-like protein NTKL). | |||||
|
NFAC2_MOUSE
|
||||||
| NC score | 0.021641 (rank : 54) | θ value | 5.27518 (rank : 42) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q60591, Q60984, Q60985 | Gene names | Nfatc2, Nfat1, Nfatp | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor of activated T-cells, cytoplasmic 2 (T cell transcription factor NFAT1) (NFAT pre-existing subunit) (NF-ATp). | |||||
|
GNAS1_MOUSE
|
||||||
| NC score | 0.020899 (rank : 55) | θ value | 6.88961 (rank : 48) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 651 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6R0H7, Q6R0H4, Q6R0H5, Q6R2J5, Q9JJX0, Q9Z1N8 | Gene names | Gnas, Gnas1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Guanine nucleotide-binding protein G(s) subunit alpha isoforms XLas (Adenylate cyclase-stimulating G alpha protein) (Extra large alphas protein) (XLalphas). | |||||
|
HRBL_MOUSE
|
||||||
| NC score | 0.020674 (rank : 56) | θ value | 6.88961 (rank : 49) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q80WC7, Q8BKS5, Q99J67 | Gene names | Hrbl, Rabr | |||
|
Domain Architecture |

|
|||||
| Description | HIV-1 Rev-binding protein-like protein (Rev/Rex activation domain- binding protein related) (RAB-R). | |||||
|
BRWD1_HUMAN
|
||||||
| NC score | 0.020670 (rank : 57) | θ value | 3.0926 (rank : 28) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 448 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NSI6, O43721, Q5R2V0, Q5R2V1, Q8TCV3, Q96QG9, Q96QH0, Q9NUK1 | Gene names | BRWD1, C21orf107, WDR9 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain and WD repeat domain-containing protein 1 (WD repeat protein 9). | |||||
|
BRWD1_MOUSE
|
||||||
| NC score | 0.019539 (rank : 58) | θ value | 6.88961 (rank : 47) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 478 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q921C3, Q921C2 | Gene names | Brwd1, Wdr9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain and WD repeat domain-containing protein 1 (WD repeat protein 9). | |||||
|
ZSWM5_HUMAN
|
||||||
| NC score | 0.019124 (rank : 59) | θ value | 5.27518 (rank : 46) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9P217, Q5SXQ9 | Gene names | ZSWIM5, KIAA1511 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger SWIM domain-containing protein 5. | |||||
|
MEGF9_MOUSE
|
||||||
| NC score | 0.017621 (rank : 60) | θ value | 8.99809 (rank : 52) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BH27, Q8BWI4 | Gene names | Megf9, Egfl5, Kiaa0818 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 9 precursor (EGF-like domain-containing protein 5) (Multiple EGF-like domain protein 5). | |||||
|
KCNH6_HUMAN
|
||||||
| NC score | 0.016566 (rank : 61) | θ value | 2.36792 (rank : 24) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H252, Q9BRD7 | Gene names | KCNH6, ERG2, HERG2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 6 (Voltage-gated potassium channel subunit Kv11.2) (Ether-a-go-go-related gene potassium channel 2) (Ether-a-go-go-related protein 2) (Eag-related protein 2). | |||||
|
ZFY28_HUMAN
|
||||||
| NC score | 0.015936 (rank : 62) | θ value | 8.99809 (rank : 53) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9HCC9 | Gene names | ZFYVE28, KIAA1643 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger FYVE domain-containing protein 28. | |||||
|
TMPSD_MOUSE
|
||||||
| NC score | 0.015828 (rank : 63) | θ value | 1.81305 (rank : 22) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 334 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5U405, Q8CFE0, Q91VQ8 | Gene names | Tmprss13, Msp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protease, serine 13 (EC 3.4.21.-) (Mosaic serine protease) (Membrane-type mosaic serine protease). | |||||
|
TMPSD_HUMAN
|
||||||
| NC score | 0.012112 (rank : 64) | θ value | 5.27518 (rank : 45) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9BYE2, Q86YM4, Q96JY8, Q9BYE1 | Gene names | TMPRSS13, MSP, TMPRSS11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protease, serine 13 (EC 3.4.21.-) (Mosaic serine protease) (Membrane-type mosaic serine protease). | |||||
|
HXB5_HUMAN
|
||||||
| NC score | 0.006179 (rank : 65) | θ value | 5.27518 (rank : 41) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P09067, P09069 | Gene names | HOXB5, HOX2A | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-B5 (Hox-2A) (HHO.C10) (HU-1). | |||||
|
K0999_HUMAN
|
||||||
| NC score | 0.004608 (rank : 66) | θ value | 1.81305 (rank : 18) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 899 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y2K2, Q59FY2, Q5M9N1, Q6P3R6, Q8IYM8, Q9HA50 | Gene names | KIAA0999 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized serine/threonine-protein kinase KIAA0999 (EC 2.7.11.1). | |||||
|
MAST2_MOUSE
|
||||||
| NC score | 0.002780 (rank : 67) | θ value | 6.88961 (rank : 50) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1110 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q60592, Q6P9M1, Q8CHD1 | Gene names | Mast2, Mast205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 2 (EC 2.7.11.1). | |||||