Please be patient as the page loads
|
CND1_MOUSE
|
||||||
| SwissProt Accessions | Q8K2Z4, Q8R0F2, Q8R3H0, Q922B7, Q923A3, Q9CS25, Q9CT32 | Gene names | Ncapd2, Capd2, Cnap1 | |||
|
Domain Architecture |

|
|||||
| Description | Condensin complex subunit 1 (Non-SMC condensin I complex subunit D2) (Chromosome condensation-related SMC-associated protein 1) (Chromosome-associated protein D2) (mCAP-D2) (XCAP-D2 homolog). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
CND1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.990366 (rank : 2) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q15021, Q8N6U3 | Gene names | NCAPD2, CAPD2, CNAP1, KIAA0159 | |||
|
Domain Architecture |

|
|||||
| Description | Condensin complex subunit 1 (Non-SMC condensin I complex subunit D2) (Chromosome condensation-related SMC-associated protein 1) (Chromosome-associated protein D2) (hCAP-D2) (XCAP-D2 homolog). | |||||
|
CND1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8K2Z4, Q8R0F2, Q8R3H0, Q922B7, Q923A3, Q9CS25, Q9CT32 | Gene names | Ncapd2, Capd2, Cnap1 | |||
|
Domain Architecture |

|
|||||
| Description | Condensin complex subunit 1 (Non-SMC condensin I complex subunit D2) (Chromosome condensation-related SMC-associated protein 1) (Chromosome-associated protein D2) (mCAP-D2) (XCAP-D2 homolog). | |||||
|
K0056_HUMAN
|
||||||
| θ value | 3.64472e-09 (rank : 3) | NC score | 0.320501 (rank : 4) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P42695 | Gene names | KIAA0056 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0056. | |||||
|
K0056_MOUSE
|
||||||
| θ value | 1.38499e-08 (rank : 4) | NC score | 0.322646 (rank : 3) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6ZQK0, Q9CS21 | Gene names | Kiaa0056 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0056. | |||||
|
EI2BE_MOUSE
|
||||||
| θ value | 0.21417 (rank : 5) | NC score | 0.051687 (rank : 5) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CHW4, Q3TU39 | Gene names | Eif2b5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Translation initiation factor eIF-2B subunit epsilon (eIF-2B GDP-GTP exchange factor subunit epsilon). | |||||
|
LAMB1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 6) | NC score | 0.019920 (rank : 16) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P02469 | Gene names | Lamb1-1, Lamb-1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-1 chain precursor (Laminin B1 chain). | |||||
|
KCC2B_MOUSE
|
||||||
| θ value | 0.62314 (rank : 7) | NC score | 0.006083 (rank : 32) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 841 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P28652, Q5SVH9 | Gene names | Camk2b, Camk2d | |||
|
Domain Architecture |

|
|||||
| Description | Calcium/calmodulin-dependent protein kinase type II beta chain (EC 2.7.11.17) (CaM-kinase II beta chain) (CaM kinase II subunit beta) (CaMK-II subunit beta). | |||||
|
LAMB1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 8) | NC score | 0.018929 (rank : 17) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 1018 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P07942 | Gene names | LAMB1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-1 chain precursor (Laminin B1 chain). | |||||
|
B4GN1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 9) | NC score | 0.041479 (rank : 6) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q00973 | Gene names | B4GALNT1, GALGT, SIAT2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-1,4 N-acetylgalactosaminyltransferase 1 (EC 2.4.1.92) ((N- acetylneuraminyl)-galactosylglucosylceramide) (GM2/GD2 synthase) (GalNAc-T). | |||||
|
FA76A_MOUSE
|
||||||
| θ value | 2.36792 (rank : 10) | NC score | 0.028504 (rank : 9) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q922G2, Q3U3U8, Q3V3N2 | Gene names | Fam76a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM76A. | |||||
|
MIS12_HUMAN
|
||||||
| θ value | 2.36792 (rank : 11) | NC score | 0.035063 (rank : 7) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9H081, Q96N24 | Gene names | MIS12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein MIS12 homolog. | |||||
|
SGIP1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 12) | NC score | 0.027221 (rank : 10) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8VD37, Q3UFU3, Q3UGA0, Q8BXX4, Q8C034, Q9CXT2 | Gene names | Sgip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3-containing GRB2-like protein 3-interacting protein 1 (Endophilin- 3-interacting protein). | |||||
|
XRCC4_HUMAN
|
||||||
| θ value | 2.36792 (rank : 13) | NC score | 0.032701 (rank : 8) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q13426, Q9BS72, Q9UP94 | Gene names | XRCC4 | |||
|
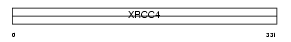
Domain Architecture |
|
|||||
| Description | DNA-repair protein XRCC4 (X-ray repair cross-complementing protein 4). | |||||
|
ESPL1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 14) | NC score | 0.023876 (rank : 13) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P60330 | Gene names | Espl1, Esp1, Kiaa0165 | |||
|
Domain Architecture |

|
|||||
| Description | Separin (EC 3.4.22.49) (Separase) (Caspase-like protein ESPL1) (Extra spindle poles-like 1 protein). | |||||
|
KCC2B_HUMAN
|
||||||
| θ value | 3.0926 (rank : 15) | NC score | 0.004562 (rank : 35) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 866 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13554, O95437, O95438, O95599, Q9UGH7, Q9UGH8, Q9UGH9, Q9UNX0, Q9UNX7, Q9UP00, Q9Y5N4, Q9Y6F4 | Gene names | CAMK2B, CAM2, CAMKB | |||
|
Domain Architecture |

|
|||||
| Description | Calcium/calmodulin-dependent protein kinase type II beta chain (EC 2.7.11.17) (CaM-kinase II beta chain) (CaM kinase II subunit beta) (CaMK-II subunit beta). | |||||
|
KTN1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 16) | NC score | 0.010931 (rank : 26) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 1324 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q61595, Q8BG49, Q8BHF4, Q8BHM8, Q8C9Y5, Q8CG51, Q8CG52, Q8CG53, Q8CG54, Q8CG55, Q8CG56, Q8CG57, Q8CG58, Q8CG59, Q8CG60, Q8CG61, Q8CG62, Q8CG63 | Gene names | Ktn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinectin. | |||||
|
RFC1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 17) | NC score | 0.020252 (rank : 15) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P35251, Q5XKF5, Q6PKU0, Q86V41, Q86V46 | Gene names | RFC1, RFC140 | |||
|
Domain Architecture |

|
|||||
| Description | Replication factor C subunit 1 (Replication factor C large subunit) (RF-C 140 kDa subunit) (Activator 1 140 kDa subunit) (Activator 1 large subunit) (A1 140 kDa subunit) (DNA-binding protein PO-GA). | |||||
|
RILP_HUMAN
|
||||||
| θ value | 3.0926 (rank : 18) | NC score | 0.025791 (rank : 12) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96NA2, Q71RE6, Q9BSL3, Q9BYS3 | Gene names | RILP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab-interacting lysosomal protein. | |||||
|
RP1L1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 19) | NC score | 0.014958 (rank : 19) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 1072 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IWN7, Q86SQ1, Q8IWN8, Q8IWN9, Q8IWP0, Q8IWP1, Q8IWP2 | Gene names | RP1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinitis pigmentosa 1-like 1 protein. | |||||
|
RRBP1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 20) | NC score | 0.010122 (rank : 30) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
DHRS3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 21) | NC score | 0.026038 (rank : 11) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75911, Q6UY38, Q9BUC8 | Gene names | DHRS3 | |||
|
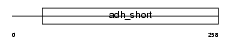
Domain Architecture |
|
|||||
| Description | Short-chain dehydrogenase/reductase 3 (EC 1.1.-.-) (Retinal short- chain dehydrogenase/reductase 1) (retSDR1) (DD83.1). | |||||
|
ABCAD_HUMAN
|
||||||
| θ value | 5.27518 (rank : 22) | NC score | 0.007529 (rank : 31) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q86UQ4, Q6ZTT7, Q86WI2, Q8N248 | Gene names | ABCA13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-binding cassette sub-family A member 13. | |||||
|
DHRS3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 23) | NC score | 0.018357 (rank : 18) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O88876, Q3UAD1, Q91WR0, Q91XC3, Q922A6 | Gene names | Dhrs3, Rsdr1 | |||
|
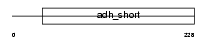
Domain Architecture |
|
|||||
| Description | Short-chain dehydrogenase/reductase 3 (EC 1.1.-.-) (Retinal short- chain dehydrogenase/reductase 1) (retSDR1). | |||||
|
GNRP_HUMAN
|
||||||
| θ value | 5.27518 (rank : 24) | NC score | 0.010413 (rank : 29) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13972, Q16027 | Gene names | RASGRF1, CDC25 | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide-releasing protein (GNRP) (Ras-specific nucleotide exchange factor CDC25) (Ras-specific guanine nucleotide-releasing factor). | |||||
|
GLE1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 25) | NC score | 0.012673 (rank : 21) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q53GS7, O75458, Q53GT9, Q5VVU1, Q8NCP6, Q9UFL6 | Gene names | GLE1L, GLE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoporin GLE1 (GLE1-like protein) (hGLE1). | |||||
|
PNMA2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 26) | NC score | 0.012091 (rank : 24) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BHK0 | Gene names | Pnma2 | |||
|
Domain Architecture |

|
|||||
| Description | Paraneoplastic antigen Ma2 homolog. | |||||
|
RERE_HUMAN
|
||||||
| θ value | 6.88961 (rank : 27) | NC score | 0.012988 (rank : 20) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9P2R6, O43393, O75046, O75359, Q5VXL9, Q6P6B9, Q9Y2W4 | Gene names | RERE, ARG, ARP, ATN1L, KIAA0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-1-like protein) (Atrophin-1-related protein). | |||||
|
RERE_MOUSE
|
||||||
| θ value | 6.88961 (rank : 28) | NC score | 0.012585 (rank : 22) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80TZ9 | Gene names | Rere, Atr2, Kiaa0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-2). | |||||
|
5NT1B_MOUSE
|
||||||
| θ value | 8.99809 (rank : 29) | NC score | 0.012455 (rank : 23) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q91YE9, Q91Y48 | Gene names | Nt5c1b, Airp | |||
|
Domain Architecture |

|
|||||
| Description | Cytosolic 5'-nucleotidase 1B (EC 3.1.3.5) (Cytosolic 5'-nucleotidase IB) (cN1B) (cN-IB) (Autoimmune infertility-related protein). | |||||
|
FA76A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 30) | NC score | 0.020829 (rank : 14) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8TAV0, O95565, O95566, Q8N7J5 | Gene names | FAM76A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM76A. | |||||
|
MYH2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 31) | NC score | 0.003757 (rank : 36) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 1491 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9UKX2, Q14322, Q16229 | Gene names | MYH2, MYHSA2 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, skeletal muscle, adult 2 (Myosin heavy chain IIa) (MyHC-IIa). | |||||
|
MYPC1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 32) | NC score | 0.004776 (rank : 34) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q00872, Q15497 | Gene names | MYBPC1, MYBPCS | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-binding protein C, slow-type (Slow MyBP-C) (C-protein, skeletal muscle slow isoform). | |||||
|
NUP88_MOUSE
|
||||||
| θ value | 8.99809 (rank : 33) | NC score | 0.011724 (rank : 25) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CEC0, Q80Z13, Q8K090 | Gene names | Nup88 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear pore complex protein Nup88 (Nucleoporin Nup88) (88 kDa nuclear pore complex protein). | |||||
|
TWST2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 34) | NC score | 0.010713 (rank : 28) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8WVJ9 | Gene names | TWIST2, DERMO1 | |||
|
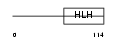
Domain Architecture |
|
|||||
| Description | Twist-related protein 2 (Dermis expressed protein 1) (Dermo-1). | |||||
|
TWST2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 35) | NC score | 0.010731 (rank : 27) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9D030 | Gene names | Twist2, Dermo1 | |||
|
Domain Architecture |
|
|||||
| Description | Twist-related protein 2 (Dermis expressed protein 1) (Dermo-1). | |||||
|
UTRO_HUMAN
|
||||||
| θ value | 8.99809 (rank : 36) | NC score | 0.006049 (rank : 33) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P46939 | Gene names | UTRN, DMDL | |||
|
Domain Architecture |

|
|||||
| Description | Utrophin (Dystrophin-related protein 1) (DRP1) (DRP). | |||||
|
CND1_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8K2Z4, Q8R0F2, Q8R3H0, Q922B7, Q923A3, Q9CS25, Q9CT32 | Gene names | Ncapd2, Capd2, Cnap1 | |||
|
Domain Architecture |

|
|||||
| Description | Condensin complex subunit 1 (Non-SMC condensin I complex subunit D2) (Chromosome condensation-related SMC-associated protein 1) (Chromosome-associated protein D2) (mCAP-D2) (XCAP-D2 homolog). | |||||
|
CND1_HUMAN
|
||||||
| NC score | 0.990366 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q15021, Q8N6U3 | Gene names | NCAPD2, CAPD2, CNAP1, KIAA0159 | |||
|
Domain Architecture |

|
|||||
| Description | Condensin complex subunit 1 (Non-SMC condensin I complex subunit D2) (Chromosome condensation-related SMC-associated protein 1) (Chromosome-associated protein D2) (hCAP-D2) (XCAP-D2 homolog). | |||||
|
K0056_MOUSE
|
||||||
| NC score | 0.322646 (rank : 3) | θ value | 1.38499e-08 (rank : 4) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6ZQK0, Q9CS21 | Gene names | Kiaa0056 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0056. | |||||
|
K0056_HUMAN
|
||||||
| NC score | 0.320501 (rank : 4) | θ value | 3.64472e-09 (rank : 3) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P42695 | Gene names | KIAA0056 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0056. | |||||
|
EI2BE_MOUSE
|
||||||
| NC score | 0.051687 (rank : 5) | θ value | 0.21417 (rank : 5) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CHW4, Q3TU39 | Gene names | Eif2b5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Translation initiation factor eIF-2B subunit epsilon (eIF-2B GDP-GTP exchange factor subunit epsilon). | |||||
|
B4GN1_HUMAN
|
||||||
| NC score | 0.041479 (rank : 6) | θ value | 1.81305 (rank : 9) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q00973 | Gene names | B4GALNT1, GALGT, SIAT2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-1,4 N-acetylgalactosaminyltransferase 1 (EC 2.4.1.92) ((N- acetylneuraminyl)-galactosylglucosylceramide) (GM2/GD2 synthase) (GalNAc-T). | |||||
|
MIS12_HUMAN
|
||||||
| NC score | 0.035063 (rank : 7) | θ value | 2.36792 (rank : 11) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9H081, Q96N24 | Gene names | MIS12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein MIS12 homolog. | |||||
|
XRCC4_HUMAN
|
||||||
| NC score | 0.032701 (rank : 8) | θ value | 2.36792 (rank : 13) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q13426, Q9BS72, Q9UP94 | Gene names | XRCC4 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-repair protein XRCC4 (X-ray repair cross-complementing protein 4). | |||||
|
FA76A_MOUSE
|
||||||
| NC score | 0.028504 (rank : 9) | θ value | 2.36792 (rank : 10) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q922G2, Q3U3U8, Q3V3N2 | Gene names | Fam76a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM76A. | |||||
|
SGIP1_MOUSE
|
||||||
| NC score | 0.027221 (rank : 10) | θ value | 2.36792 (rank : 12) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8VD37, Q3UFU3, Q3UGA0, Q8BXX4, Q8C034, Q9CXT2 | Gene names | Sgip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3-containing GRB2-like protein 3-interacting protein 1 (Endophilin- 3-interacting protein). | |||||
|
DHRS3_HUMAN
|
||||||
| NC score | 0.026038 (rank : 11) | θ value | 4.03905 (rank : 21) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75911, Q6UY38, Q9BUC8 | Gene names | DHRS3 | |||
|
Domain Architecture |

|
|||||
| Description | Short-chain dehydrogenase/reductase 3 (EC 1.1.-.-) (Retinal short- chain dehydrogenase/reductase 1) (retSDR1) (DD83.1). | |||||
|
RILP_HUMAN
|
||||||
| NC score | 0.025791 (rank : 12) | θ value | 3.0926 (rank : 18) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96NA2, Q71RE6, Q9BSL3, Q9BYS3 | Gene names | RILP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab-interacting lysosomal protein. | |||||
|
ESPL1_MOUSE
|
||||||
| NC score | 0.023876 (rank : 13) | θ value | 3.0926 (rank : 14) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P60330 | Gene names | Espl1, Esp1, Kiaa0165 | |||
|
Domain Architecture |

|
|||||
| Description | Separin (EC 3.4.22.49) (Separase) (Caspase-like protein ESPL1) (Extra spindle poles-like 1 protein). | |||||
|
FA76A_HUMAN
|
||||||
| NC score | 0.020829 (rank : 14) | θ value | 8.99809 (rank : 30) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8TAV0, O95565, O95566, Q8N7J5 | Gene names | FAM76A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM76A. | |||||
|
RFC1_HUMAN
|
||||||
| NC score | 0.020252 (rank : 15) | θ value | 3.0926 (rank : 17) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P35251, Q5XKF5, Q6PKU0, Q86V41, Q86V46 | Gene names | RFC1, RFC140 | |||
|
Domain Architecture |

|
|||||
| Description | Replication factor C subunit 1 (Replication factor C large subunit) (RF-C 140 kDa subunit) (Activator 1 140 kDa subunit) (Activator 1 large subunit) (A1 140 kDa subunit) (DNA-binding protein PO-GA). | |||||
|
LAMB1_MOUSE
|
||||||
| NC score | 0.019920 (rank : 16) | θ value | 0.47712 (rank : 6) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P02469 | Gene names | Lamb1-1, Lamb-1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-1 chain precursor (Laminin B1 chain). | |||||
|
LAMB1_HUMAN
|
||||||
| NC score | 0.018929 (rank : 17) | θ value | 0.813845 (rank : 8) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 1018 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P07942 | Gene names | LAMB1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-1 chain precursor (Laminin B1 chain). | |||||
|
DHRS3_MOUSE
|
||||||
| NC score | 0.018357 (rank : 18) | θ value | 5.27518 (rank : 23) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O88876, Q3UAD1, Q91WR0, Q91XC3, Q922A6 | Gene names | Dhrs3, Rsdr1 | |||
|
Domain Architecture |

|
|||||
| Description | Short-chain dehydrogenase/reductase 3 (EC 1.1.-.-) (Retinal short- chain dehydrogenase/reductase 1) (retSDR1). | |||||
|
RP1L1_HUMAN
|
||||||
| NC score | 0.014958 (rank : 19) | θ value | 3.0926 (rank : 19) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 1072 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IWN7, Q86SQ1, Q8IWN8, Q8IWN9, Q8IWP0, Q8IWP1, Q8IWP2 | Gene names | RP1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinitis pigmentosa 1-like 1 protein. | |||||
|
RERE_HUMAN
|
||||||
| NC score | 0.012988 (rank : 20) | θ value | 6.88961 (rank : 27) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9P2R6, O43393, O75046, O75359, Q5VXL9, Q6P6B9, Q9Y2W4 | Gene names | RERE, ARG, ARP, ATN1L, KIAA0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-1-like protein) (Atrophin-1-related protein). | |||||
|
GLE1_HUMAN
|
||||||
| NC score | 0.012673 (rank : 21) | θ value | 6.88961 (rank : 25) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q53GS7, O75458, Q53GT9, Q5VVU1, Q8NCP6, Q9UFL6 | Gene names | GLE1L, GLE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoporin GLE1 (GLE1-like protein) (hGLE1). | |||||
|
RERE_MOUSE
|
||||||
| NC score | 0.012585 (rank : 22) | θ value | 6.88961 (rank : 28) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80TZ9 | Gene names | Rere, Atr2, Kiaa0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-2). | |||||
|
5NT1B_MOUSE
|
||||||
| NC score | 0.012455 (rank : 23) | θ value | 8.99809 (rank : 29) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q91YE9, Q91Y48 | Gene names | Nt5c1b, Airp | |||
|
Domain Architecture |

|
|||||
| Description | Cytosolic 5'-nucleotidase 1B (EC 3.1.3.5) (Cytosolic 5'-nucleotidase IB) (cN1B) (cN-IB) (Autoimmune infertility-related protein). | |||||
|
PNMA2_MOUSE
|
||||||
| NC score | 0.012091 (rank : 24) | θ value | 6.88961 (rank : 26) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BHK0 | Gene names | Pnma2 | |||
|
Domain Architecture |

|
|||||
| Description | Paraneoplastic antigen Ma2 homolog. | |||||
|
NUP88_MOUSE
|
||||||
| NC score | 0.011724 (rank : 25) | θ value | 8.99809 (rank : 33) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CEC0, Q80Z13, Q8K090 | Gene names | Nup88 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear pore complex protein Nup88 (Nucleoporin Nup88) (88 kDa nuclear pore complex protein). | |||||
|
KTN1_MOUSE
|
||||||
| NC score | 0.010931 (rank : 26) | θ value | 3.0926 (rank : 16) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 1324 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q61595, Q8BG49, Q8BHF4, Q8BHM8, Q8C9Y5, Q8CG51, Q8CG52, Q8CG53, Q8CG54, Q8CG55, Q8CG56, Q8CG57, Q8CG58, Q8CG59, Q8CG60, Q8CG61, Q8CG62, Q8CG63 | Gene names | Ktn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinectin. | |||||
|
TWST2_MOUSE
|
||||||
| NC score | 0.010731 (rank : 27) | θ value | 8.99809 (rank : 35) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9D030 | Gene names | Twist2, Dermo1 | |||
|
Domain Architecture |

|
|||||
| Description | Twist-related protein 2 (Dermis expressed protein 1) (Dermo-1). | |||||
|
TWST2_HUMAN
|
||||||
| NC score | 0.010713 (rank : 28) | θ value | 8.99809 (rank : 34) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8WVJ9 | Gene names | TWIST2, DERMO1 | |||
|
Domain Architecture |

|
|||||
| Description | Twist-related protein 2 (Dermis expressed protein 1) (Dermo-1). | |||||
|
GNRP_HUMAN
|
||||||
| NC score | 0.010413 (rank : 29) | θ value | 5.27518 (rank : 24) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13972, Q16027 | Gene names | RASGRF1, CDC25 | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide-releasing protein (GNRP) (Ras-specific nucleotide exchange factor CDC25) (Ras-specific guanine nucleotide-releasing factor). | |||||
|
RRBP1_MOUSE
|
||||||
| NC score | 0.010122 (rank : 30) | θ value | 3.0926 (rank : 20) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
ABCAD_HUMAN
|
||||||
| NC score | 0.007529 (rank : 31) | θ value | 5.27518 (rank : 22) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q86UQ4, Q6ZTT7, Q86WI2, Q8N248 | Gene names | ABCA13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-binding cassette sub-family A member 13. | |||||
|
KCC2B_MOUSE
|
||||||
| NC score | 0.006083 (rank : 32) | θ value | 0.62314 (rank : 7) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 841 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P28652, Q5SVH9 | Gene names | Camk2b, Camk2d | |||
|
Domain Architecture |

|
|||||
| Description | Calcium/calmodulin-dependent protein kinase type II beta chain (EC 2.7.11.17) (CaM-kinase II beta chain) (CaM kinase II subunit beta) (CaMK-II subunit beta). | |||||
|
UTRO_HUMAN
|
||||||
| NC score | 0.006049 (rank : 33) | θ value | 8.99809 (rank : 36) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P46939 | Gene names | UTRN, DMDL | |||
|
Domain Architecture |

|
|||||
| Description | Utrophin (Dystrophin-related protein 1) (DRP1) (DRP). | |||||
|
MYPC1_HUMAN
|
||||||
| NC score | 0.004776 (rank : 34) | θ value | 8.99809 (rank : 32) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q00872, Q15497 | Gene names | MYBPC1, MYBPCS | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-binding protein C, slow-type (Slow MyBP-C) (C-protein, skeletal muscle slow isoform). | |||||
|
KCC2B_HUMAN
|
||||||
| NC score | 0.004562 (rank : 35) | θ value | 3.0926 (rank : 15) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 866 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13554, O95437, O95438, O95599, Q9UGH7, Q9UGH8, Q9UGH9, Q9UNX0, Q9UNX7, Q9UP00, Q9Y5N4, Q9Y6F4 | Gene names | CAMK2B, CAM2, CAMKB | |||
|
Domain Architecture |

|
|||||
| Description | Calcium/calmodulin-dependent protein kinase type II beta chain (EC 2.7.11.17) (CaM-kinase II beta chain) (CaM kinase II subunit beta) (CaMK-II subunit beta). | |||||
|
MYH2_HUMAN
|
||||||
| NC score | 0.003757 (rank : 36) | θ value | 8.99809 (rank : 31) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 1491 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9UKX2, Q14322, Q16229 | Gene names | MYH2, MYHSA2 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, skeletal muscle, adult 2 (Myosin heavy chain IIa) (MyHC-IIa). | |||||