Please be patient as the page loads
|
VEGFD_HUMAN
|
||||||
| SwissProt Accessions | O43915 | Gene names | FIGF, VEGFD | |||
|
Domain Architecture |
|
|||||
| Description | Vascular endothelial growth factor D precursor (VEGF-D) (c-fos-induced growth factor) (FIGF). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
VEGFD_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O43915 | Gene names | FIGF, VEGFD | |||
|
Domain Architecture |

|
|||||
| Description | Vascular endothelial growth factor D precursor (VEGF-D) (c-fos-induced growth factor) (FIGF). | |||||
|
VEGFD_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.993117 (rank : 2) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P97946 | Gene names | Figf, Vegfd | |||
|
Domain Architecture |
|
|||||
| Description | Vascular endothelial growth factor D precursor (VEGF-D) (c-fos-induced growth factor) (FIGF). | |||||
|
VEGFC_HUMAN
|
||||||
| θ value | 9.11921e-69 (rank : 3) | NC score | 0.884205 (rank : 4) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P49767 | Gene names | VEGFC | |||
|
Domain Architecture |
|
|||||
| Description | Vascular endothelial growth factor C precursor (VEGF-C) (Vascular endothelial growth factor-related protein) (VRP) (Flt4 ligand) (Flt4- L). | |||||
|
VEGFC_MOUSE
|
||||||
| θ value | 4.52582e-68 (rank : 4) | NC score | 0.895870 (rank : 3) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P97953, Q543R6 | Gene names | Vegfc | |||
|
Domain Architecture |
|
|||||
| Description | Vascular endothelial growth factor C precursor (VEGF-C) (Vascular endothelial growth factor-related protein) (VRP) (Flt4 ligand) (Flt4- L). | |||||
|
VEGFA_HUMAN
|
||||||
| θ value | 4.59992e-12 (rank : 5) | NC score | 0.668635 (rank : 5) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P15692, O60720, O75875, Q16889, Q5UB46, Q6P0P5, Q96KJ0, Q96L82, Q96NW5, Q9H1W8, Q9H1W9, Q9UH58, Q9UL23 | Gene names | VEGF, VEGFA | |||
|
Domain Architecture |
|
|||||
| Description | Vascular endothelial growth factor A precursor (VEGF-A) (Vascular permeability factor) (VPF). | |||||
|
VEGFA_MOUSE
|
||||||
| θ value | 6.00763e-12 (rank : 6) | NC score | 0.653903 (rank : 6) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q00731, O70123, Q66S31, Q6GT23, Q6WZL9 | Gene names | Vegf, Vegfa | |||
|
Domain Architecture |
|
|||||
| Description | Vascular endothelial growth factor A precursor (VEGF-A) (Vascular permeability factor) (VPF). | |||||
|
PLGF_HUMAN
|
||||||
| θ value | 5.62301e-10 (rank : 7) | NC score | 0.638262 (rank : 8) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P49763, Q07101, Q9BV78, Q9Y6S8 | Gene names | PGF, PLGF | |||
|
Domain Architecture |
|
|||||
| Description | Placenta growth factor precursor (PlGF). | |||||
|
VEGFB_MOUSE
|
||||||
| θ value | 4.76016e-09 (rank : 8) | NC score | 0.630278 (rank : 9) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P49766, Q3UG04, Q5D0B1, Q64290 | Gene names | Vegfb, Vrf | |||
|
Domain Architecture |
|
|||||
| Description | Vascular endothelial growth factor B precursor (VEGF-B) (VEGF-related factor) (VRF). | |||||
|
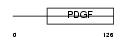
PLGF_MOUSE
|
||||||
| θ value | 8.11959e-09 (rank : 9) | NC score | 0.638975 (rank : 7) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P49764 | Gene names | Pgf, Plgf | |||
|
Domain Architecture |
|
|||||
| Description | Placenta growth factor precursor (PlGF). | |||||
|
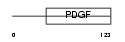
VEGFB_HUMAN
|
||||||
| θ value | 8.11959e-09 (rank : 10) | NC score | 0.627102 (rank : 10) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P49765, Q16528 | Gene names | VEGFB, VRF | |||
|
Domain Architecture |
|
|||||
| Description | Vascular endothelial growth factor B precursor (VEGF-B) (VEGF-related factor) (VRF). | |||||
|
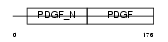
PDGFB_HUMAN
|
||||||
| θ value | 2.88788e-06 (rank : 11) | NC score | 0.465989 (rank : 11) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P01127, P78431, Q9UF23 | Gene names | PDGFB, PDGF2, SIS | |||
|
Domain Architecture |
|
|||||
| Description | Platelet-derived growth factor B chain precursor (PDGF B-chain) (Platelet-derived growth factor beta polypeptide) (PDGF-2) (c-sis) (Becaplermin). | |||||
|
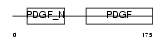
PDGFB_MOUSE
|
||||||
| θ value | 2.44474e-05 (rank : 12) | NC score | 0.444816 (rank : 12) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P31240, Q6P3C4 | Gene names | Pdgfb, Sis | |||
|
Domain Architecture |
|
|||||
| Description | Platelet-derived growth factor B chain precursor (PDGF B-chain) (Platelet-derived growth factor beta polypeptide) (PDGF-2) (c-sis). | |||||
|
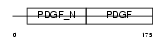
PDGFA_HUMAN
|
||||||
| θ value | 0.000121331 (rank : 13) | NC score | 0.438683 (rank : 13) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P04085 | Gene names | PDGFA, PDGF1 | |||
|
Domain Architecture |
|
|||||
| Description | Platelet-derived growth factor A chain precursor (PDGF A-chain) (Platelet-derived growth factor alpha polypeptide) (PDGF-1). | |||||
|
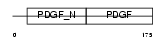
PDGFA_MOUSE
|
||||||
| θ value | 0.000461057 (rank : 14) | NC score | 0.422199 (rank : 14) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P20033 | Gene names | Pdgfa | |||
|
Domain Architecture |
|
|||||
| Description | Platelet-derived growth factor A chain precursor (PDGF A-chain) (Platelet-derived growth factor alpha polypeptide) (PDGF-1). | |||||
|
PDGFD_HUMAN
|
||||||
| θ value | 0.00228821 (rank : 15) | NC score | 0.184105 (rank : 15) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9GZP0, Q9BWV5 | Gene names | PDGFD, IEGF, SCDGFB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Platelet-derived growth factor D precursor (PDGF D) (Iris-expressed growth factor) (Spinal cord-derived growth factor B) (SCDGF-B). | |||||
|
PDGFD_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 16) | NC score | 0.164863 (rank : 16) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q925I7, Q3URF6, Q8K2L3, Q9D1L8 | Gene names | Pdgfd, Scdgfb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Platelet-derived growth factor chain precursor (PDGF D) (Spinal cord- derived growth factor B) (SCDGF-B). | |||||
|
ADA28_HUMAN
|
||||||
| θ value | 0.365318 (rank : 17) | NC score | 0.029688 (rank : 29) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9UKQ2, Q9Y339, Q9Y3S0 | Gene names | ADAM28, MDCL | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 28 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 28) (Metalloproteinase-like, disintegrin-like, and cysteine- rich protein-L) (MDC-L) (eMDC II) (ADAM23). | |||||
|
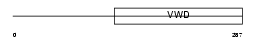
BMPER_MOUSE
|
||||||
| θ value | 0.47712 (rank : 18) | NC score | 0.042957 (rank : 20) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8CJ69, Q7TN57, Q80UZ1, Q9CXM8 | Gene names | Bmper, Cv2 | |||
|
Domain Architecture |
|
|||||
| Description | BMP-binding endothelial regulator protein precursor (Bone morphogenetic protein-binding endothelial cell precursor-derived regulator) (Protein crossveinless-2) (mCV2). | |||||
|
PCSK5_HUMAN
|
||||||
| θ value | 0.62314 (rank : 19) | NC score | 0.022507 (rank : 34) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q92824, Q13527 | Gene names | PCSK5, PC5, PC6 | |||
|
Domain Architecture |

|
|||||
| Description | Proprotein convertase subtilisin/kexin type 5 precursor (EC 3.4.21.-) (Proprotein convertase PC5) (Subtilisin/kexin-like protease PC5) (PC6) (hPC6). | |||||
|
SREC_HUMAN
|
||||||
| θ value | 0.62314 (rank : 20) | NC score | 0.039586 (rank : 24) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q14162, O43701 | Gene names | SCARF1, KIAA0149, SREC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endothelial cells scavenger receptor precursor (Acetyl LDL receptor) (Scavenger receptor class F member 1). | |||||
|
KR101_HUMAN
|
||||||
| θ value | 1.38821 (rank : 21) | NC score | 0.042689 (rank : 21) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P60331 | Gene names | KRTAP10-1, KAP10.1, KAP18-1, KRTAP10.1, KRTAP18-1, KRTAP18.1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 10-1 (Keratin-associated protein 10.1) (High sulfur keratin-associated protein 10.1) (Keratin-associated protein 18-1) (Keratin-associated protein 18.1). | |||||
|
RSPO3_HUMAN
|
||||||
| θ value | 1.38821 (rank : 22) | NC score | 0.040170 (rank : 22) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9BXY4, Q5VTV4, Q96K87 | Gene names | RSPO3, PWTSR, THSD2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | R-spondin-3 precursor (Roof plate-specific spondin-3) (hRspo3) (Thrombospondin type-1 domain-containing protein 2) (Protein with TSP type-1 repeat) (hPWTSR). | |||||
|
TRI42_HUMAN
|
||||||
| θ value | 1.38821 (rank : 23) | NC score | 0.034778 (rank : 25) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IWZ5, Q8N832, Q8NDL3 | Gene names | TRIM42 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 42. | |||||
|
ADA28_MOUSE
|
||||||
| θ value | 1.81305 (rank : 24) | NC score | 0.027272 (rank : 31) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9JLN6, Q8K5D2, Q8K5D3 | Gene names | Adam28 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 28 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 28) (Thymic epithelial cell-ADAM) (TECADAM). | |||||
|
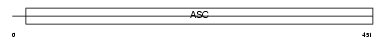
ACCN1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 25) | NC score | 0.020644 (rank : 38) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q16515, Q13553, Q8N3E2 | Gene names | ACCN1, ACCN, BNAC1, MDEG | |||
|
Domain Architecture |
|
|||||
| Description | Amiloride-sensitive cation channel 1, neuronal (Amiloride-sensitive brain sodium channel) (Amiloride-sensitive cation channel neuronal 1) (Acid-sensing ion channel 2) (ASIC2) (Brain sodium channel 1) (BNaC1) (BNC1) (Mammalian degenerin homolog). | |||||
|
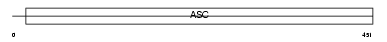
ACCN1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 26) | NC score | 0.020650 (rank : 37) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q925H0, Q5SUU2, Q61203 | Gene names | Accn1, Asic2, Bnac1 | |||
|
Domain Architecture |
|
|||||
| Description | Amiloride-sensitive cation channel 1, neuronal (Amiloride-sensitive brain sodium channel) (Acid-sensing ion channel 2) (Brain sodium channel 1) (BNaC1). | |||||
|
ATRN_MOUSE
|
||||||
| θ value | 2.36792 (rank : 27) | NC score | 0.017356 (rank : 40) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9WU60, Q9R263, Q9WU77 | Gene names | Atrn, mg, Mgca | |||
|
Domain Architecture |

|
|||||
| Description | Attractin precursor (Mahogany protein). | |||||
|
MCSP_MOUSE
|
||||||
| θ value | 2.36792 (rank : 28) | NC score | 0.051694 (rank : 19) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P15265, O70613, Q6P8N3 | Gene names | Smcp, Mcs, Mcsp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm mitochondrial-associated cysteine-rich protein. | |||||
|
ADA32_MOUSE
|
||||||
| θ value | 3.0926 (rank : 29) | NC score | 0.028227 (rank : 30) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8K410, Q6P901, Q8BJ80 | Gene names | Adam32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAM 32 precursor (A disintegrin and metalloproteinase domain 32). | |||||
|
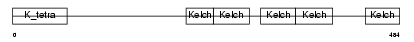
KLHL5_HUMAN
|
||||||
| θ value | 3.0926 (rank : 30) | NC score | 0.005635 (rank : 48) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96PQ7, Q86XW0, Q9NUK3, Q9NV27, Q9NVA9, Q9Y2X2 | Gene names | KLHL5 | |||
|
Domain Architecture |
|
|||||
| Description | Kelch-like protein 5. | |||||
|
RSPO3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 31) | NC score | 0.034396 (rank : 26) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q2TJ95, Q3SYI9, Q5R2V4, Q8BVW2, Q9CSB2 | Gene names | Rspo3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | R-spondin-3 precursor (Roof plate-specific spondin-3) (Cysteine-rich and single thrombospondin domain-containing protein 1) (Cristin-1) (Nucleopondin) (Cabriolet). | |||||
|
CRIM1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 32) | NC score | 0.031281 (rank : 27) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9NZV1, Q59GH0, Q7LCQ5, Q9H318 | Gene names | CRIM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cysteine-rich motor neuron 1 protein precursor (CRIM-1) (Cysteine-rich repeat-containing protein S52). | |||||
|
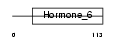
GLHA_HUMAN
|
||||||
| θ value | 4.03905 (rank : 33) | NC score | 0.055990 (rank : 17) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P01215 | Gene names | CGA | |||
|
Domain Architecture |
|
|||||
| Description | Glycoprotein hormones alpha chain precursor (Anterior pituitary glycoprotein hormones common subunit alpha) (Follitropin alpha chain) (Follicle-stimulating hormone alpha chain) (FSH-alpha) (Lutropin alpha chain) (Luteinizing hormone alpha chain) (LSH-alpha) (Thyrotropin alpha chain) (Thyroid-stimulating hormone alpha chain) (TSH-alpha) (Choriogonadotropin alpha chain) (Chorionic gonadotrophin alpha subunit) (CG-alpha). | |||||
|
MEGF6_HUMAN
|
||||||
| θ value | 4.03905 (rank : 34) | NC score | 0.018536 (rank : 39) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O75095, Q5VV39 | Gene names | MEGF6, EGFL3, KIAA0815 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Multiple epidermal growth factor-like domains 6 precursor (EGF-like domain-containing protein 3) (Multiple EGF-like domain protein 3). | |||||
|
SLIT3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 35) | NC score | 0.006571 (rank : 46) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O75094, O95804, Q9UFH5 | Gene names | SLIT3, KIAA0814, MEGF5, SLIL2 | |||
|
Domain Architecture |

|
|||||
| Description | Slit homolog 3 protein precursor (Slit-3) (Multiple epidermal growth factor-like domains 5). | |||||
|
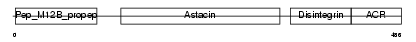
ADA11_HUMAN
|
||||||
| θ value | 6.88961 (rank : 36) | NC score | 0.026995 (rank : 32) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O75078, Q14808, Q14809, Q14810 | Gene names | ADAM11, MDC | |||
|
Domain Architecture |
|
|||||
| Description | ADAM 11 precursor (A disintegrin and metalloproteinase domain 11) (Metalloproteinase-like, disintegrin-like, and cysteine-rich protein) (MDC). | |||||
|
ADA11_MOUSE
|
||||||
| θ value | 6.88961 (rank : 37) | NC score | 0.026822 (rank : 33) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9R1V4 | Gene names | Adam11, Mdc | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 11 precursor (A disintegrin and metalloproteinase domain 11) (Metalloproteinase-like, disintegrin-like, and cysteine-rich protein) (MDC). | |||||
|
ADA22_HUMAN
|
||||||
| θ value | 6.88961 (rank : 38) | NC score | 0.021940 (rank : 35) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9P0K1, O75075, O75076, Q9P0K2, Q9UIA1, Q9UKK2 | Gene names | ADAM22, MDC2 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 22 precursor (A disintegrin and metalloproteinase domain 22) (Metalloproteinase-like, disintegrin-like, and cysteine-rich protein 2) (Metalloproteinase-disintegrin ADAM22-3). | |||||
|
FBLI1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 39) | NC score | 0.005666 (rank : 47) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q71FD7, Q99J35 | Gene names | Fblim1, Cal | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-binding LIM protein 1 (CSX-associated LIM). | |||||
|
LTBP2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 40) | NC score | 0.012700 (rank : 42) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 552 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q14767, Q99907, Q9NS51 | Gene names | LTBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein 2 precursor (LTBP-2). | |||||
|
MT1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 41) | NC score | 0.040016 (rank : 23) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P02802, Q64485 | Gene names | Mt1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-1 (MT-1) (Metallothionein-I) (MT-I). | |||||
|
ZAN_MOUSE
|
||||||
| θ value | 6.88961 (rank : 42) | NC score | 0.030318 (rank : 28) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 808 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O88799, O08647 | Gene names | Zan | |||
|
Domain Architecture |

|
|||||
| Description | Zonadhesin precursor. | |||||
|
ZN180_HUMAN
|
||||||
| θ value | 6.88961 (rank : 43) | NC score | -0.001943 (rank : 49) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 752 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UJW8, Q9P1U2 | Gene names | ZNF180 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger protein 180 (HHZ168). | |||||
|
ADA22_MOUSE
|
||||||
| θ value | 8.99809 (rank : 44) | NC score | 0.021616 (rank : 36) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9R1V6, Q5TLI8, Q5TLI9, Q5TLJ0, Q5TLJ1, Q5TLJ2, Q5TLJ3, Q5TLJ4, Q5TLJ5, Q5TLJ6, Q5TLJ7, Q5TLJ8, Q5TLJ9, Q5TLK0, Q5TLK1, Q5TLK2, Q5TLK3, Q5TLK4, Q5TLK5, Q5TLK6, Q5TLK7, Q5TLK8, Q8BSF2, Q9R1V5 | Gene names | Adam22 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 22 precursor (A disintegrin and metalloproteinase domain 22). | |||||
|
DLL1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 45) | NC score | 0.011676 (rank : 45) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 456 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q61483 | Gene names | Dll1 | |||
|
Domain Architecture |
|
|||||
| Description | Delta-like protein 1 precursor (Drosophila Delta homolog 1) (Delta1). | |||||
|
EGFL8_HUMAN
|
||||||
| θ value | 8.99809 (rank : 46) | NC score | 0.011980 (rank : 44) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 270 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q99944, Q8IV30 | Gene names | EGFL8, C6orf8, NG3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EGF-like domain-containing protein 8 precursor (Epidermal growth factor-like protein 8) (Multiple EGF-like domain protein 8) (Vascular endothelial-statin 2) (VE-statin-2). | |||||
|
LTBP2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 47) | NC score | 0.012002 (rank : 43) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 617 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O08999, Q8C6W9 | Gene names | Ltbp2 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein 2 precursor (LTBP-2). | |||||
|
OLR1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 48) | NC score | 0.016444 (rank : 41) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P78380, Q2PP00, Q7Z484 | Gene names | OLR1, LOX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oxidized low-density lipoprotein receptor 1 (Ox-LDL receptor 1) (Lectin-type oxidized LDL receptor 1) (Lectin-like oxidized LDL receptor 1) (Lectin-like oxLDL receptor 1) (LOX-1) (hLOX-1) [Contains: Oxidized low-density lipoprotein receptor 1, soluble form]. | |||||
|
SEPP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 49) | NC score | 0.053349 (rank : 18) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P70274, Q9Z2T7 | Gene names | Sepp1, Selp | |||
|
Domain Architecture |

|
|||||
| Description | Selenoprotein P precursor (SeP) (Plasma selenoprotein P). | |||||
|
VEGFD_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O43915 | Gene names | FIGF, VEGFD | |||
|
Domain Architecture |

|
|||||
| Description | Vascular endothelial growth factor D precursor (VEGF-D) (c-fos-induced growth factor) (FIGF). | |||||
|
VEGFD_MOUSE
|
||||||
| NC score | 0.993117 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P97946 | Gene names | Figf, Vegfd | |||
|
Domain Architecture |

|
|||||
| Description | Vascular endothelial growth factor D precursor (VEGF-D) (c-fos-induced growth factor) (FIGF). | |||||
|
VEGFC_MOUSE
|
||||||
| NC score | 0.895870 (rank : 3) | θ value | 4.52582e-68 (rank : 4) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P97953, Q543R6 | Gene names | Vegfc | |||
|
Domain Architecture |

|
|||||
| Description | Vascular endothelial growth factor C precursor (VEGF-C) (Vascular endothelial growth factor-related protein) (VRP) (Flt4 ligand) (Flt4- L). | |||||
|
VEGFC_HUMAN
|
||||||
| NC score | 0.884205 (rank : 4) | θ value | 9.11921e-69 (rank : 3) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P49767 | Gene names | VEGFC | |||
|
Domain Architecture |

|
|||||
| Description | Vascular endothelial growth factor C precursor (VEGF-C) (Vascular endothelial growth factor-related protein) (VRP) (Flt4 ligand) (Flt4- L). | |||||
|
VEGFA_HUMAN
|
||||||
| NC score | 0.668635 (rank : 5) | θ value | 4.59992e-12 (rank : 5) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P15692, O60720, O75875, Q16889, Q5UB46, Q6P0P5, Q96KJ0, Q96L82, Q96NW5, Q9H1W8, Q9H1W9, Q9UH58, Q9UL23 | Gene names | VEGF, VEGFA | |||
|
Domain Architecture |

|
|||||
| Description | Vascular endothelial growth factor A precursor (VEGF-A) (Vascular permeability factor) (VPF). | |||||
|
VEGFA_MOUSE
|
||||||
| NC score | 0.653903 (rank : 6) | θ value | 6.00763e-12 (rank : 6) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q00731, O70123, Q66S31, Q6GT23, Q6WZL9 | Gene names | Vegf, Vegfa | |||
|
Domain Architecture |

|
|||||
| Description | Vascular endothelial growth factor A precursor (VEGF-A) (Vascular permeability factor) (VPF). | |||||
|
PLGF_MOUSE
|
||||||
| NC score | 0.638975 (rank : 7) | θ value | 8.11959e-09 (rank : 9) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P49764 | Gene names | Pgf, Plgf | |||
|
Domain Architecture |

|
|||||
| Description | Placenta growth factor precursor (PlGF). | |||||
|
PLGF_HUMAN
|
||||||
| NC score | 0.638262 (rank : 8) | θ value | 5.62301e-10 (rank : 7) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P49763, Q07101, Q9BV78, Q9Y6S8 | Gene names | PGF, PLGF | |||
|
Domain Architecture |

|
|||||
| Description | Placenta growth factor precursor (PlGF). | |||||
|
VEGFB_MOUSE
|
||||||
| NC score | 0.630278 (rank : 9) | θ value | 4.76016e-09 (rank : 8) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P49766, Q3UG04, Q5D0B1, Q64290 | Gene names | Vegfb, Vrf | |||
|
Domain Architecture |

|
|||||
| Description | Vascular endothelial growth factor B precursor (VEGF-B) (VEGF-related factor) (VRF). | |||||
|
VEGFB_HUMAN
|
||||||
| NC score | 0.627102 (rank : 10) | θ value | 8.11959e-09 (rank : 10) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P49765, Q16528 | Gene names | VEGFB, VRF | |||
|
Domain Architecture |

|
|||||
| Description | Vascular endothelial growth factor B precursor (VEGF-B) (VEGF-related factor) (VRF). | |||||
|
PDGFB_HUMAN
|
||||||
| NC score | 0.465989 (rank : 11) | θ value | 2.88788e-06 (rank : 11) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P01127, P78431, Q9UF23 | Gene names | PDGFB, PDGF2, SIS | |||
|
Domain Architecture |

|
|||||
| Description | Platelet-derived growth factor B chain precursor (PDGF B-chain) (Platelet-derived growth factor beta polypeptide) (PDGF-2) (c-sis) (Becaplermin). | |||||
|
PDGFB_MOUSE
|
||||||
| NC score | 0.444816 (rank : 12) | θ value | 2.44474e-05 (rank : 12) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P31240, Q6P3C4 | Gene names | Pdgfb, Sis | |||
|
Domain Architecture |

|
|||||
| Description | Platelet-derived growth factor B chain precursor (PDGF B-chain) (Platelet-derived growth factor beta polypeptide) (PDGF-2) (c-sis). | |||||
|
PDGFA_HUMAN
|
||||||
| NC score | 0.438683 (rank : 13) | θ value | 0.000121331 (rank : 13) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P04085 | Gene names | PDGFA, PDGF1 | |||
|
Domain Architecture |

|
|||||
| Description | Platelet-derived growth factor A chain precursor (PDGF A-chain) (Platelet-derived growth factor alpha polypeptide) (PDGF-1). | |||||
|
PDGFA_MOUSE
|
||||||
| NC score | 0.422199 (rank : 14) | θ value | 0.000461057 (rank : 14) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P20033 | Gene names | Pdgfa | |||
|
Domain Architecture |

|
|||||
| Description | Platelet-derived growth factor A chain precursor (PDGF A-chain) (Platelet-derived growth factor alpha polypeptide) (PDGF-1). | |||||
|
PDGFD_HUMAN
|
||||||
| NC score | 0.184105 (rank : 15) | θ value | 0.00228821 (rank : 15) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9GZP0, Q9BWV5 | Gene names | PDGFD, IEGF, SCDGFB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Platelet-derived growth factor D precursor (PDGF D) (Iris-expressed growth factor) (Spinal cord-derived growth factor B) (SCDGF-B). | |||||
|
PDGFD_MOUSE
|
||||||
| NC score | 0.164863 (rank : 16) | θ value | 0.00390308 (rank : 16) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q925I7, Q3URF6, Q8K2L3, Q9D1L8 | Gene names | Pdgfd, Scdgfb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Platelet-derived growth factor chain precursor (PDGF D) (Spinal cord- derived growth factor B) (SCDGF-B). | |||||
|
GLHA_HUMAN
|
||||||
| NC score | 0.055990 (rank : 17) | θ value | 4.03905 (rank : 33) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P01215 | Gene names | CGA | |||
|
Domain Architecture |

|
|||||
| Description | Glycoprotein hormones alpha chain precursor (Anterior pituitary glycoprotein hormones common subunit alpha) (Follitropin alpha chain) (Follicle-stimulating hormone alpha chain) (FSH-alpha) (Lutropin alpha chain) (Luteinizing hormone alpha chain) (LSH-alpha) (Thyrotropin alpha chain) (Thyroid-stimulating hormone alpha chain) (TSH-alpha) (Choriogonadotropin alpha chain) (Chorionic gonadotrophin alpha subunit) (CG-alpha). | |||||
|
SEPP1_MOUSE
|
||||||
| NC score | 0.053349 (rank : 18) | θ value | θ > 10 (rank : 49) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P70274, Q9Z2T7 | Gene names | Sepp1, Selp | |||
|
Domain Architecture |

|
|||||
| Description | Selenoprotein P precursor (SeP) (Plasma selenoprotein P). | |||||
|
MCSP_MOUSE
|
||||||
| NC score | 0.051694 (rank : 19) | θ value | 2.36792 (rank : 28) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P15265, O70613, Q6P8N3 | Gene names | Smcp, Mcs, Mcsp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm mitochondrial-associated cysteine-rich protein. | |||||
|
BMPER_MOUSE
|
||||||
| NC score | 0.042957 (rank : 20) | θ value | 0.47712 (rank : 18) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8CJ69, Q7TN57, Q80UZ1, Q9CXM8 | Gene names | Bmper, Cv2 | |||
|
Domain Architecture |

|
|||||
| Description | BMP-binding endothelial regulator protein precursor (Bone morphogenetic protein-binding endothelial cell precursor-derived regulator) (Protein crossveinless-2) (mCV2). | |||||
|
KR101_HUMAN
|
||||||
| NC score | 0.042689 (rank : 21) | θ value | 1.38821 (rank : 21) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P60331 | Gene names | KRTAP10-1, KAP10.1, KAP18-1, KRTAP10.1, KRTAP18-1, KRTAP18.1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 10-1 (Keratin-associated protein 10.1) (High sulfur keratin-associated protein 10.1) (Keratin-associated protein 18-1) (Keratin-associated protein 18.1). | |||||
|
RSPO3_HUMAN
|
||||||
| NC score | 0.040170 (rank : 22) | θ value | 1.38821 (rank : 22) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9BXY4, Q5VTV4, Q96K87 | Gene names | RSPO3, PWTSR, THSD2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | R-spondin-3 precursor (Roof plate-specific spondin-3) (hRspo3) (Thrombospondin type-1 domain-containing protein 2) (Protein with TSP type-1 repeat) (hPWTSR). | |||||
|
MT1_MOUSE
|
||||||
| NC score | 0.040016 (rank : 23) | θ value | 6.88961 (rank : 41) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P02802, Q64485 | Gene names | Mt1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metallothionein-1 (MT-1) (Metallothionein-I) (MT-I). | |||||
|
SREC_HUMAN
|
||||||
| NC score | 0.039586 (rank : 24) | θ value | 0.62314 (rank : 20) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q14162, O43701 | Gene names | SCARF1, KIAA0149, SREC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endothelial cells scavenger receptor precursor (Acetyl LDL receptor) (Scavenger receptor class F member 1). | |||||
|
TRI42_HUMAN
|
||||||
| NC score | 0.034778 (rank : 25) | θ value | 1.38821 (rank : 23) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IWZ5, Q8N832, Q8NDL3 | Gene names | TRIM42 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 42. | |||||
|
RSPO3_MOUSE
|
||||||
| NC score | 0.034396 (rank : 26) | θ value | 3.0926 (rank : 31) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q2TJ95, Q3SYI9, Q5R2V4, Q8BVW2, Q9CSB2 | Gene names | Rspo3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | R-spondin-3 precursor (Roof plate-specific spondin-3) (Cysteine-rich and single thrombospondin domain-containing protein 1) (Cristin-1) (Nucleopondin) (Cabriolet). | |||||
|
CRIM1_HUMAN
|
||||||
| NC score | 0.031281 (rank : 27) | θ value | 4.03905 (rank : 32) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9NZV1, Q59GH0, Q7LCQ5, Q9H318 | Gene names | CRIM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cysteine-rich motor neuron 1 protein precursor (CRIM-1) (Cysteine-rich repeat-containing protein S52). | |||||
|
ZAN_MOUSE
|
||||||
| NC score | 0.030318 (rank : 28) | θ value | 6.88961 (rank : 42) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 808 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O88799, O08647 | Gene names | Zan | |||
|
Domain Architecture |

|
|||||
| Description | Zonadhesin precursor. | |||||
|
ADA28_HUMAN
|
||||||
| NC score | 0.029688 (rank : 29) | θ value | 0.365318 (rank : 17) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9UKQ2, Q9Y339, Q9Y3S0 | Gene names | ADAM28, MDCL | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 28 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 28) (Metalloproteinase-like, disintegrin-like, and cysteine- rich protein-L) (MDC-L) (eMDC II) (ADAM23). | |||||
|
ADA32_MOUSE
|
||||||
| NC score | 0.028227 (rank : 30) | θ value | 3.0926 (rank : 29) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8K410, Q6P901, Q8BJ80 | Gene names | Adam32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAM 32 precursor (A disintegrin and metalloproteinase domain 32). | |||||
|
ADA28_MOUSE
|
||||||
| NC score | 0.027272 (rank : 31) | θ value | 1.81305 (rank : 24) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9JLN6, Q8K5D2, Q8K5D3 | Gene names | Adam28 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 28 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 28) (Thymic epithelial cell-ADAM) (TECADAM). | |||||
|
ADA11_HUMAN
|
||||||
| NC score | 0.026995 (rank : 32) | θ value | 6.88961 (rank : 36) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O75078, Q14808, Q14809, Q14810 | Gene names | ADAM11, MDC | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 11 precursor (A disintegrin and metalloproteinase domain 11) (Metalloproteinase-like, disintegrin-like, and cysteine-rich protein) (MDC). | |||||
|
ADA11_MOUSE
|
||||||
| NC score | 0.026822 (rank : 33) | θ value | 6.88961 (rank : 37) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9R1V4 | Gene names | Adam11, Mdc | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 11 precursor (A disintegrin and metalloproteinase domain 11) (Metalloproteinase-like, disintegrin-like, and cysteine-rich protein) (MDC). | |||||
|
PCSK5_HUMAN
|
||||||
| NC score | 0.022507 (rank : 34) | θ value | 0.62314 (rank : 19) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q92824, Q13527 | Gene names | PCSK5, PC5, PC6 | |||
|
Domain Architecture |

|
|||||
| Description | Proprotein convertase subtilisin/kexin type 5 precursor (EC 3.4.21.-) (Proprotein convertase PC5) (Subtilisin/kexin-like protease PC5) (PC6) (hPC6). | |||||
|
ADA22_HUMAN
|
||||||
| NC score | 0.021940 (rank : 35) | θ value | 6.88961 (rank : 38) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9P0K1, O75075, O75076, Q9P0K2, Q9UIA1, Q9UKK2 | Gene names | ADAM22, MDC2 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 22 precursor (A disintegrin and metalloproteinase domain 22) (Metalloproteinase-like, disintegrin-like, and cysteine-rich protein 2) (Metalloproteinase-disintegrin ADAM22-3). | |||||
|
ADA22_MOUSE
|
||||||
| NC score | 0.021616 (rank : 36) | θ value | 8.99809 (rank : 44) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9R1V6, Q5TLI8, Q5TLI9, Q5TLJ0, Q5TLJ1, Q5TLJ2, Q5TLJ3, Q5TLJ4, Q5TLJ5, Q5TLJ6, Q5TLJ7, Q5TLJ8, Q5TLJ9, Q5TLK0, Q5TLK1, Q5TLK2, Q5TLK3, Q5TLK4, Q5TLK5, Q5TLK6, Q5TLK7, Q5TLK8, Q8BSF2, Q9R1V5 | Gene names | Adam22 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 22 precursor (A disintegrin and metalloproteinase domain 22). | |||||
|
ACCN1_MOUSE
|
||||||
| NC score | 0.020650 (rank : 37) | θ value | 2.36792 (rank : 26) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q925H0, Q5SUU2, Q61203 | Gene names | Accn1, Asic2, Bnac1 | |||
|
Domain Architecture |

|
|||||
| Description | Amiloride-sensitive cation channel 1, neuronal (Amiloride-sensitive brain sodium channel) (Acid-sensing ion channel 2) (Brain sodium channel 1) (BNaC1). | |||||
|
ACCN1_HUMAN
|
||||||
| NC score | 0.020644 (rank : 38) | θ value | 2.36792 (rank : 25) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q16515, Q13553, Q8N3E2 | Gene names | ACCN1, ACCN, BNAC1, MDEG | |||
|
Domain Architecture |

|
|||||
| Description | Amiloride-sensitive cation channel 1, neuronal (Amiloride-sensitive brain sodium channel) (Amiloride-sensitive cation channel neuronal 1) (Acid-sensing ion channel 2) (ASIC2) (Brain sodium channel 1) (BNaC1) (BNC1) (Mammalian degenerin homolog). | |||||
|
MEGF6_HUMAN
|
||||||
| NC score | 0.018536 (rank : 39) | θ value | 4.03905 (rank : 34) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O75095, Q5VV39 | Gene names | MEGF6, EGFL3, KIAA0815 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Multiple epidermal growth factor-like domains 6 precursor (EGF-like domain-containing protein 3) (Multiple EGF-like domain protein 3). | |||||
|
ATRN_MOUSE
|
||||||
| NC score | 0.017356 (rank : 40) | θ value | 2.36792 (rank : 27) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9WU60, Q9R263, Q9WU77 | Gene names | Atrn, mg, Mgca | |||
|
Domain Architecture |

|
|||||
| Description | Attractin precursor (Mahogany protein). | |||||
|
OLR1_HUMAN
|
||||||
| NC score | 0.016444 (rank : 41) | θ value | 8.99809 (rank : 48) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P78380, Q2PP00, Q7Z484 | Gene names | OLR1, LOX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oxidized low-density lipoprotein receptor 1 (Ox-LDL receptor 1) (Lectin-type oxidized LDL receptor 1) (Lectin-like oxidized LDL receptor 1) (Lectin-like oxLDL receptor 1) (LOX-1) (hLOX-1) [Contains: Oxidized low-density lipoprotein receptor 1, soluble form]. | |||||
|
LTBP2_HUMAN
|
||||||
| NC score | 0.012700 (rank : 42) | θ value | 6.88961 (rank : 40) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 552 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q14767, Q99907, Q9NS51 | Gene names | LTBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein 2 precursor (LTBP-2). | |||||
|
LTBP2_MOUSE
|
||||||
| NC score | 0.012002 (rank : 43) | θ value | 8.99809 (rank : 47) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 617 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O08999, Q8C6W9 | Gene names | Ltbp2 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein 2 precursor (LTBP-2). | |||||
|
EGFL8_HUMAN
|
||||||
| NC score | 0.011980 (rank : 44) | θ value | 8.99809 (rank : 46) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 270 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q99944, Q8IV30 | Gene names | EGFL8, C6orf8, NG3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EGF-like domain-containing protein 8 precursor (Epidermal growth factor-like protein 8) (Multiple EGF-like domain protein 8) (Vascular endothelial-statin 2) (VE-statin-2). | |||||
|
DLL1_MOUSE
|
||||||
| NC score | 0.011676 (rank : 45) | θ value | 8.99809 (rank : 45) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 456 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q61483 | Gene names | Dll1 | |||
|
Domain Architecture |

|
|||||
| Description | Delta-like protein 1 precursor (Drosophila Delta homolog 1) (Delta1). | |||||
|
SLIT3_HUMAN
|
||||||
| NC score | 0.006571 (rank : 46) | θ value | 5.27518 (rank : 35) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O75094, O95804, Q9UFH5 | Gene names | SLIT3, KIAA0814, MEGF5, SLIL2 | |||
|
Domain Architecture |

|
|||||
| Description | Slit homolog 3 protein precursor (Slit-3) (Multiple epidermal growth factor-like domains 5). | |||||
|
FBLI1_MOUSE
|
||||||
| NC score | 0.005666 (rank : 47) | θ value | 6.88961 (rank : 39) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q71FD7, Q99J35 | Gene names | Fblim1, Cal | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-binding LIM protein 1 (CSX-associated LIM). | |||||
|
KLHL5_HUMAN
|
||||||
| NC score | 0.005635 (rank : 48) | θ value | 3.0926 (rank : 30) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96PQ7, Q86XW0, Q9NUK3, Q9NV27, Q9NVA9, Q9Y2X2 | Gene names | KLHL5 | |||
|
Domain Architecture |

|
|||||
| Description | Kelch-like protein 5. | |||||
|
ZN180_HUMAN
|
||||||
| NC score | -0.001943 (rank : 49) | θ value | 6.88961 (rank : 43) | |||
| Query Neighborhood Hits | 48 | Target Neighborhood Hits | 752 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UJW8, Q9P1U2 | Gene names | ZNF180 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 180 (HHZ168). | |||||