Please be patient as the page loads
|
NUFP2_HUMAN
|
||||||
| SwissProt Accessions | Q7Z417, Q9P2M5 | Gene names | NUFIP2, KIAA1321, PIG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear fragile X mental retardation-interacting protein 2 (FMRP- interacting protein 2) (82 kDa FMRP-interacting protein) (82-FIP) (Proliferation-inducing gene 1 protein). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
NUFP2_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q7Z417, Q9P2M5 | Gene names | NUFIP2, KIAA1321, PIG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear fragile X mental retardation-interacting protein 2 (FMRP- interacting protein 2) (82 kDa FMRP-interacting protein) (82-FIP) (Proliferation-inducing gene 1 protein). | |||||
|
NUFP2_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.986251 (rank : 2) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q5F2E7, Q3TCE2, Q3V195, Q80TF1 | Gene names | Nufip2, Kiaa1321 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear fragile X mental retardation-interacting protein 2 (FMRP- interacting protein 2) (82 kDa FMRP-interacting protein) (82-FIP). | |||||
|
CYLC1_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 3) | NC score | 0.086877 (rank : 3) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P35663, Q5JQQ9 | Gene names | CYLC1, CYL, CYL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-1 (Cylicin I) (Multiple-band polypeptide I). | |||||
|
PGCA_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 4) | NC score | 0.030038 (rank : 18) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 643 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q61282, Q64021 | Gene names | Agc1, Agc | |||
|
Domain Architecture |

|
|||||
| Description | Aggrecan core protein precursor (Cartilage-specific proteoglycan core protein) (CSPCP). | |||||
|
RRBP1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 5) | NC score | 0.037995 (rank : 13) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1306 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9P2E9, O75300, O75301, Q5W165, Q96SB2, Q9BWP1, Q9H476 | Gene names | RRBP1, KIAA1398 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (180 kDa ribosome receptor homolog) (ES/130-related protein). | |||||
|
RRBP1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 6) | NC score | 0.040059 (rank : 12) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
AFF4_MOUSE
|
||||||
| θ value | 0.62314 (rank : 7) | NC score | 0.040839 (rank : 10) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9ESC8, Q8C6K4, Q8C6W3, Q8CCH3 | Gene names | Aff4, Alf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4. | |||||
|
PLD1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 8) | NC score | 0.052009 (rank : 7) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13393 | Gene names | PLD1 | |||
|
Domain Architecture |

|
|||||
| Description | Phospholipase D1 (EC 3.1.4.4) (PLD 1) (Choline phosphatase 1) (Phosphatidylcholine-hydrolyzing phospholipase D1) (hPLD1). | |||||
|
PCDBC_HUMAN
|
||||||
| θ value | 0.813845 (rank : 9) | NC score | 0.022150 (rank : 27) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Y5F1 | Gene names | PCDHB12 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin beta 12 precursor (PCDH-beta12). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 0.813845 (rank : 10) | NC score | 0.031287 (rank : 16) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
CKAP2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 11) | NC score | 0.063860 (rank : 4) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8WWK9, Q3KRA5, Q5VXB4, Q8IWV5, Q8IWV6, Q96FH9, Q9H012, Q9H0D0, Q9H988, Q9HC49, Q9NVG4 | Gene names | CKAP2, LB1, TMAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-associated protein 2 (Tumor-associated microtubule- associated protein) (CTCL tumor antigen se20-10). | |||||
|
COAA1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 12) | NC score | 0.020819 (rank : 29) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 316 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q03692 | Gene names | COL10A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(X) chain precursor. | |||||
|
EGR1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 13) | NC score | 0.007988 (rank : 56) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 816 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P08046, Q61777 | Gene names | Egr1, Egr-1, Krox-24 | |||
|
Domain Architecture |

|
|||||
| Description | Early growth response protein 1 (EGR-1) (Krox-24 protein) (Transcription factor Zif268) (Nerve growth factor-induced protein A) (NGFI-A). | |||||
|
CENPF_HUMAN
|
||||||
| θ value | 1.81305 (rank : 14) | NC score | 0.013814 (rank : 49) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1932 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P49454, Q13171, Q13246 | Gene names | CENPF | |||
|
Domain Architecture |

|
|||||
| Description | Centromere protein F (Kinetochore protein CENP-F) (Mitosin) (AH antigen). | |||||
|
EMSY_HUMAN
|
||||||
| θ value | 1.81305 (rank : 15) | NC score | 0.030015 (rank : 19) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q7Z589, Q4G109, Q8NBU6, Q8TE50, Q9H8I9, Q9NRH0 | Gene names | EMSY, C11orf30 | |||
|
Domain Architecture |
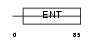
|
|||||
| Description | Protein EMSY. | |||||
|
PCDB3_HUMAN
|
||||||
| θ value | 1.81305 (rank : 16) | NC score | 0.021368 (rank : 28) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Y5E6 | Gene names | PCDHB3 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin beta 3 precursor (PCDH-beta3). | |||||
|
ANK2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 17) | NC score | 0.010574 (rank : 52) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
MAGB2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 18) | NC score | 0.018606 (rank : 41) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O15479, O75860 | Gene names | MAGEB2 | |||
|
Domain Architecture |
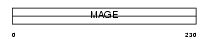
|
|||||
| Description | Melanoma-associated antigen B2 (MAGE-B2 antigen) (DSS-AHC critical interval MAGE superfamily 6) (DAM6) (MAGE XP-2). | |||||
|
ZCHC6_MOUSE
|
||||||
| θ value | 2.36792 (rank : 19) | NC score | 0.024583 (rank : 23) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5BLK4, Q8CIH3 | Gene names | Zcchc6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 6. | |||||
|
CXA5_MOUSE
|
||||||
| θ value | 3.0926 (rank : 20) | NC score | 0.014943 (rank : 47) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q01231 | Gene names | Gja5, Cxn-40 | |||
|
Domain Architecture |

|
|||||
| Description | Gap junction alpha-5 protein (Connexin-40) (Cx40). | |||||
|
MAGD1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 21) | NC score | 0.015933 (rank : 45) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y5V3, Q5VSH6, Q8IZ84, Q8WY92, Q9H352, Q9HBT4, Q9UF36 | Gene names | MAGED1, NRAGE | |||
|
Domain Architecture |

|
|||||
| Description | Melanoma-associated antigen D1 (MAGE-D1 antigen) (Neurotrophin receptor-interacting MAGE homolog) (MAGE tumor antigen CCF). | |||||
|
NEIL2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 22) | NC score | 0.030968 (rank : 17) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q969S2, Q7Z3Q7, Q8N842, Q8NG52 | Gene names | NEIL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endonuclease VIII-like 2 (EC 3.2.2.-) (EC 4.2.99.18) (Nei-like 2) (DNA glycosylase/AP lyase Neil2) (DNA-(apurinic or apyrimidinic site) lyase Neil2) (NEH2). | |||||
|
PME17_MOUSE
|
||||||
| θ value | 3.0926 (rank : 23) | NC score | 0.026578 (rank : 20) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60696 | Gene names | Silv, D10H12S53E, Pmel17, Si | |||
|
Domain Architecture |

|
|||||
| Description | Melanocyte protein Pmel 17 precursor (Silver locus protein). | |||||
|
AP3B1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 24) | NC score | 0.019028 (rank : 40) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Z1T1, Q91YR4 | Gene names | Ap3b1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-1 (Adapter-related protein complex 3 beta-1 subunit) (Beta3A-adaptin) (Adaptor protein complex AP-3 beta-1 subunit) (Clathrin assembly protein complex 3 beta-1 large chain). | |||||
|
BMPR2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 25) | NC score | 0.006193 (rank : 58) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 736 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q13873, Q16569 | Gene names | BMPR2, PPH1 | |||
|
Domain Architecture |

|
|||||
| Description | Bone morphogenetic protein receptor type-2 precursor (EC 2.7.11.30) (Bone morphogenetic protein receptor type II) (BMP type II receptor) (BMPR-II). | |||||
|
CHIC2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 26) | NC score | 0.059464 (rank : 6) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UKJ5 | Gene names | CHIC2, BTL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cysteine-rich hydrophobic domain 2 protein (BrX-like translocated in leukemia). | |||||
|
CHIC2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 27) | NC score | 0.059528 (rank : 5) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D9G3, Q3T9U2, Q80ZT7 | Gene names | Chic2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cysteine-rich hydrophobic domain 2 protein. | |||||
|
CLSPN_HUMAN
|
||||||
| θ value | 4.03905 (rank : 28) | NC score | 0.022613 (rank : 26) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9HAW4, Q5VYG0, Q6P6H5, Q8IWI1 | Gene names | CLSPN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Claspin (hClaspin) (Hu-Claspin). | |||||
|
DSPP_HUMAN
|
||||||
| θ value | 4.03905 (rank : 29) | NC score | 0.033155 (rank : 15) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
PEG3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 30) | NC score | 0.007763 (rank : 57) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1101 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q3URU2, O54978, Q3TQ69, Q5EBP7, Q61138, Q6GQS0, Q80U47, Q8R5N0, Q9QX53 | Gene names | Peg3, Kiaa0287, Pw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Paternally expressed gene 3 protein (ASF-1). | |||||
|
ZN638_MOUSE
|
||||||
| θ value | 4.03905 (rank : 31) | NC score | 0.034604 (rank : 14) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q61464, Q6DFV9, Q8C941 | Gene names | Znf638, Np220, Zfml | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 638 (Nuclear protein 220) (Zinc-finger matrin-like protein). | |||||
|
CPNE7_HUMAN
|
||||||
| θ value | 5.27518 (rank : 32) | NC score | 0.014977 (rank : 46) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UBL6 | Gene names | CPNE7 | |||
|
Domain Architecture |

|
|||||
| Description | Copine-7 (Copine VII). | |||||
|
CT111_MOUSE
|
||||||
| θ value | 5.27518 (rank : 33) | NC score | 0.026410 (rank : 21) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D722, Q3U5N3, Q9CXJ2, Q9D9G1 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | Uncharacterized protein C20orf111 homolog. | |||||
|
FILA_HUMAN
|
||||||
| θ value | 5.27518 (rank : 34) | NC score | 0.040618 (rank : 11) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 685 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P20930, Q01720, Q5T583 | Gene names | FLG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filaggrin. | |||||
|
MARK4_MOUSE
|
||||||
| θ value | 5.27518 (rank : 35) | NC score | 0.002697 (rank : 60) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 898 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8CIP4, Q80T81 | Gene names | Mark4, Kiaa1860 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MAP/microtubule affinity-regulating kinase 4 (EC 2.7.11.1). | |||||
|
PCDB7_HUMAN
|
||||||
| θ value | 5.27518 (rank : 36) | NC score | 0.019947 (rank : 31) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Y5E2 | Gene names | PCDHB7 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin beta 7 precursor (PCDH-beta7). | |||||
|
TNC6B_HUMAN
|
||||||
| θ value | 5.27518 (rank : 37) | NC score | 0.042432 (rank : 9) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9UPQ9, Q5TH52, Q8TBX2 | Gene names | TNRC6B, KIAA1093 | |||
|
Domain Architecture |

|
|||||
| Description | Trinucleotide repeat-containing 6B protein. | |||||
|
DDX21_MOUSE
|
||||||
| θ value | 6.88961 (rank : 38) | NC score | 0.022976 (rank : 25) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9JIK5, Q9WV45 | Gene names | Ddx21 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar RNA helicase 2 (EC 3.6.1.-) (Nucleolar RNA helicase II) (Nucleolar RNA helicase Gu) (RH II/Gu) (Gu-alpha) (DEAD box protein 21). | |||||
|
EEA1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 39) | NC score | 0.008357 (rank : 55) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1796 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q15075, Q14221 | Gene names | EEA1, ZFYVE2 | |||
|
Domain Architecture |

|
|||||
| Description | Early endosome antigen 1 (Endosome-associated protein p162) (Zinc finger FYVE domain-containing protein 2). | |||||
|
MARK4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 40) | NC score | 0.002440 (rank : 62) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 886 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96L34, Q8NG37, Q96JG7, Q96SQ2, Q9BYD8 | Gene names | MARK4, KIAA1860, MARKL1 | |||
|
Domain Architecture |

|
|||||
| Description | MAP/microtubule affinity-regulating kinase 4 (EC 2.7.11.1) (MAP/microtubule affinity-regulating kinase-like 1). | |||||
|
MYCB2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 41) | NC score | 0.019200 (rank : 37) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q7TPH6, Q6PCM8 | Gene names | Mycbp2, Pam, Phr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ubiquitin ligase protein MYCBP2 (EC 6.3.2.-) (Myc-binding protein 2) (Protein associated with Myc) (Pam/highwire/rpm-1 protein). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 6.88961 (rank : 42) | NC score | 0.020362 (rank : 30) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
NIPBL_HUMAN
|
||||||
| θ value | 6.88961 (rank : 43) | NC score | 0.026087 (rank : 22) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
NOL8_MOUSE
|
||||||
| θ value | 6.88961 (rank : 44) | NC score | 0.016583 (rank : 42) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q3UHX0, Q504M4, Q8CDJ7, Q9CUR0 | Gene names | Nol8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleolar protein 8. | |||||
|
PHF14_HUMAN
|
||||||
| θ value | 6.88961 (rank : 45) | NC score | 0.012911 (rank : 50) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O94880 | Gene names | PHF14, KIAA0783 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 14. | |||||
|
SRCH_HUMAN
|
||||||
| θ value | 6.88961 (rank : 46) | NC score | 0.048585 (rank : 8) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P23327, Q504Y6 | Gene names | HRC, HCP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcoplasmic reticulum histidine-rich calcium-binding protein precursor. | |||||
|
AP4E1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 47) | NC score | 0.010473 (rank : 53) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UPM8, Q9Y588 | Gene names | AP4E1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-4 complex subunit epsilon-1 (Adapter-related protein complex 4 epsilon-1 subunit) (Epsilon subunit of AP-4) (AP-4 adapter complex epsilon subunit). | |||||
|
KIF22_HUMAN
|
||||||
| θ value | 8.99809 (rank : 48) | NC score | 0.004856 (rank : 59) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q14807, O94814, Q9BT46 | Gene names | KIF22, KID, KNSL4 | |||
|
Domain Architecture |
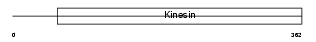
|
|||||
| Description | Kinesin-like protein KIF22 (Kinesin-like DNA-binding protein) (Kinesin-like protein 4). | |||||
|
PCDB2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 49) | NC score | 0.019308 (rank : 32) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Y5E7 | Gene names | PCDHB2 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin beta 2 precursor (PCDH-beta2). | |||||
|
PCDB4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 50) | NC score | 0.019274 (rank : 35) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Y5E5 | Gene names | PCDHB4 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin beta 4 precursor (PCDH-beta4). | |||||
|
PCDB5_HUMAN
|
||||||
| θ value | 8.99809 (rank : 51) | NC score | 0.019295 (rank : 34) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y5E4, Q9UFU9 | Gene names | PCDHB5 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin beta 5 precursor (PCDH-beta5). | |||||
|
PCDB8_HUMAN
|
||||||
| θ value | 8.99809 (rank : 52) | NC score | 0.019192 (rank : 38) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9UN66 | Gene names | PCDHB8, PCDH3I | |||
|
Domain Architecture |
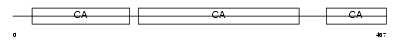
|
|||||
| Description | Protocadherin beta 8 precursor (PCDH-beta8) (Protocadherin 3I). | |||||
|
PCDB9_HUMAN
|
||||||
| θ value | 8.99809 (rank : 53) | NC score | 0.019225 (rank : 36) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y5E1, Q9NRJ8 | Gene names | PCDHB9, PCDH3H | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin beta 9 precursor (PCDH-beta9) (Protocadherin 3H). | |||||
|
PCDBA_HUMAN
|
||||||
| θ value | 8.99809 (rank : 54) | NC score | 0.019299 (rank : 33) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9UN67, Q96T99 | Gene names | PCDHB10 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin beta 10 precursor (PCDH-beta10). | |||||
|
PCDBG_HUMAN
|
||||||
| θ value | 8.99809 (rank : 55) | NC score | 0.019173 (rank : 39) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NRJ7, Q8IYD5, Q96SE9, Q9HCF1 | Gene names | PCDHB16, KIAA1621, PCDH3X | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin beta 16 precursor (PCDH-beta16) (Protocadherin 3X). | |||||
|
PF21A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 56) | NC score | 0.010342 (rank : 54) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96BD5, Q6AWA2, Q9C0G7, Q9H8V9, Q9HAK6, Q9NZE9 | Gene names | PHF21A, BHC80, KIAA1696 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (BHC80a). | |||||
|
SPG21_HUMAN
|
||||||
| θ value | 8.99809 (rank : 57) | NC score | 0.016549 (rank : 43) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NZD8, Q6ZMB6 | Gene names | SPG21, ACP33 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Maspardin (Spastic paraplegia 21 autosomal recessive Mast syndrome protein) (Acid cluster protein 33). | |||||
|
TDRD7_MOUSE
|
||||||
| θ value | 8.99809 (rank : 58) | NC score | 0.023481 (rank : 24) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8K1H1, Q8R181 | Gene names | Tdrd7, Pctaire2bp | |||
|
Domain Architecture |

|
|||||
| Description | Tudor domain-containing protein 7 (Tudor repeat associator with PCTAIRE 2) (Trap). | |||||
|
TITIN_HUMAN
|
||||||
| θ value | 8.99809 (rank : 59) | NC score | 0.002563 (rank : 61) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
TXD11_HUMAN
|
||||||
| θ value | 8.99809 (rank : 60) | NC score | 0.014143 (rank : 48) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6PKC3, O95887, Q6PJA6, Q8N2Q4, Q96K45, Q96K53 | Gene names | TXNDC11, EFP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 11 (EF-hand-binding protein 1). | |||||
|
USP9X_HUMAN
|
||||||
| θ value | 8.99809 (rank : 61) | NC score | 0.012298 (rank : 51) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q93008, O75550, Q8WWT3, Q8WX12 | Gene names | USP9X, DFFRX, USP9 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase FAF-X (EC 3.1.2.15) (Ubiquitin thioesterase FAF-X) (Ubiquitin-specific-processing protease FAF-X) (Deubiquitinating enzyme FAF-X) (Fat facets protein-related, X- linked) (Ubiquitin-specific protease 9, X chromosome). | |||||
|
YETS2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 62) | NC score | 0.016368 (rank : 44) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q3TUF7, Q6PGF8, Q80TI2, Q8CG86 | Gene names | Yeats2, Kiaa1197 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YEATS domain-containing protein 2. | |||||
|
NUFP2_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q7Z417, Q9P2M5 | Gene names | NUFIP2, KIAA1321, PIG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear fragile X mental retardation-interacting protein 2 (FMRP- interacting protein 2) (82 kDa FMRP-interacting protein) (82-FIP) (Proliferation-inducing gene 1 protein). | |||||
|
NUFP2_MOUSE
|
||||||
| NC score | 0.986251 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q5F2E7, Q3TCE2, Q3V195, Q80TF1 | Gene names | Nufip2, Kiaa1321 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear fragile X mental retardation-interacting protein 2 (FMRP- interacting protein 2) (82 kDa FMRP-interacting protein) (82-FIP). | |||||
|
CYLC1_HUMAN
|
||||||
| NC score | 0.086877 (rank : 3) | θ value | 0.0961366 (rank : 3) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P35663, Q5JQQ9 | Gene names | CYLC1, CYL, CYL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-1 (Cylicin I) (Multiple-band polypeptide I). | |||||
|
CKAP2_HUMAN
|
||||||
| NC score | 0.063860 (rank : 4) | θ value | 1.38821 (rank : 11) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8WWK9, Q3KRA5, Q5VXB4, Q8IWV5, Q8IWV6, Q96FH9, Q9H012, Q9H0D0, Q9H988, Q9HC49, Q9NVG4 | Gene names | CKAP2, LB1, TMAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-associated protein 2 (Tumor-associated microtubule- associated protein) (CTCL tumor antigen se20-10). | |||||
|
CHIC2_MOUSE
|
||||||
| NC score | 0.059528 (rank : 5) | θ value | 4.03905 (rank : 27) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D9G3, Q3T9U2, Q80ZT7 | Gene names | Chic2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cysteine-rich hydrophobic domain 2 protein. | |||||
|
CHIC2_HUMAN
|
||||||
| NC score | 0.059464 (rank : 6) | θ value | 4.03905 (rank : 26) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UKJ5 | Gene names | CHIC2, BTL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cysteine-rich hydrophobic domain 2 protein (BrX-like translocated in leukemia). | |||||
|
PLD1_HUMAN
|
||||||
| NC score | 0.052009 (rank : 7) | θ value | 0.62314 (rank : 8) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13393 | Gene names | PLD1 | |||
|
Domain Architecture |

|
|||||
| Description | Phospholipase D1 (EC 3.1.4.4) (PLD 1) (Choline phosphatase 1) (Phosphatidylcholine-hydrolyzing phospholipase D1) (hPLD1). | |||||
|
SRCH_HUMAN
|
||||||
| NC score | 0.048585 (rank : 8) | θ value | 6.88961 (rank : 46) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P23327, Q504Y6 | Gene names | HRC, HCP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcoplasmic reticulum histidine-rich calcium-binding protein precursor. | |||||
|
TNC6B_HUMAN
|
||||||
| NC score | 0.042432 (rank : 9) | θ value | 5.27518 (rank : 37) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9UPQ9, Q5TH52, Q8TBX2 | Gene names | TNRC6B, KIAA1093 | |||
|
Domain Architecture |

|
|||||
| Description | Trinucleotide repeat-containing 6B protein. | |||||
|
AFF4_MOUSE
|
||||||
| NC score | 0.040839 (rank : 10) | θ value | 0.62314 (rank : 7) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9ESC8, Q8C6K4, Q8C6W3, Q8CCH3 | Gene names | Aff4, Alf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4. | |||||
|
FILA_HUMAN
|
||||||
| NC score | 0.040618 (rank : 11) | θ value | 5.27518 (rank : 34) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 685 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P20930, Q01720, Q5T583 | Gene names | FLG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filaggrin. | |||||
|
RRBP1_MOUSE
|
||||||
| NC score | 0.040059 (rank : 12) | θ value | 0.47712 (rank : 6) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
RRBP1_HUMAN
|
||||||
| NC score | 0.037995 (rank : 13) | θ value | 0.47712 (rank : 5) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1306 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9P2E9, O75300, O75301, Q5W165, Q96SB2, Q9BWP1, Q9H476 | Gene names | RRBP1, KIAA1398 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (180 kDa ribosome receptor homolog) (ES/130-related protein). | |||||
|
ZN638_MOUSE
|
||||||
| NC score | 0.034604 (rank : 14) | θ value | 4.03905 (rank : 31) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q61464, Q6DFV9, Q8C941 | Gene names | Znf638, Np220, Zfml | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 638 (Nuclear protein 220) (Zinc-finger matrin-like protein). | |||||
|
DSPP_HUMAN
|
||||||
| NC score | 0.033155 (rank : 15) | θ value | 4.03905 (rank : 29) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.031287 (rank : 16) | θ value | 0.813845 (rank : 10) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
NEIL2_HUMAN
|
||||||
| NC score | 0.030968 (rank : 17) | θ value | 3.0926 (rank : 22) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q969S2, Q7Z3Q7, Q8N842, Q8NG52 | Gene names | NEIL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endonuclease VIII-like 2 (EC 3.2.2.-) (EC 4.2.99.18) (Nei-like 2) (DNA glycosylase/AP lyase Neil2) (DNA-(apurinic or apyrimidinic site) lyase Neil2) (NEH2). | |||||
|
PGCA_MOUSE
|
||||||
| NC score | 0.030038 (rank : 18) | θ value | 0.0961366 (rank : 4) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 643 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q61282, Q64021 | Gene names | Agc1, Agc | |||
|
Domain Architecture |

|
|||||
| Description | Aggrecan core protein precursor (Cartilage-specific proteoglycan core protein) (CSPCP). | |||||
|
EMSY_HUMAN
|
||||||
| NC score | 0.030015 (rank : 19) | θ value | 1.81305 (rank : 15) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q7Z589, Q4G109, Q8NBU6, Q8TE50, Q9H8I9, Q9NRH0 | Gene names | EMSY, C11orf30 | |||
|
Domain Architecture |

|
|||||
| Description | Protein EMSY. | |||||
|
PME17_MOUSE
|
||||||
| NC score | 0.026578 (rank : 20) | θ value | 3.0926 (rank : 23) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60696 | Gene names | Silv, D10H12S53E, Pmel17, Si | |||
|
Domain Architecture |

|
|||||
| Description | Melanocyte protein Pmel 17 precursor (Silver locus protein). | |||||
|
CT111_MOUSE
|
||||||
| NC score | 0.026410 (rank : 21) | θ value | 5.27518 (rank : 33) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D722, Q3U5N3, Q9CXJ2, Q9D9G1 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | Uncharacterized protein C20orf111 homolog. | |||||
|
NIPBL_HUMAN
|
||||||
| NC score | 0.026087 (rank : 22) | θ value | 6.88961 (rank : 43) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
ZCHC6_MOUSE
|
||||||
| NC score | 0.024583 (rank : 23) | θ value | 2.36792 (rank : 19) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5BLK4, Q8CIH3 | Gene names | Zcchc6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 6. | |||||
|
TDRD7_MOUSE
|
||||||
| NC score | 0.023481 (rank : 24) | θ value | 8.99809 (rank : 58) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8K1H1, Q8R181 | Gene names | Tdrd7, Pctaire2bp | |||
|
Domain Architecture |

|
|||||
| Description | Tudor domain-containing protein 7 (Tudor repeat associator with PCTAIRE 2) (Trap). | |||||
|
DDX21_MOUSE
|
||||||
| NC score | 0.022976 (rank : 25) | θ value | 6.88961 (rank : 38) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9JIK5, Q9WV45 | Gene names | Ddx21 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar RNA helicase 2 (EC 3.6.1.-) (Nucleolar RNA helicase II) (Nucleolar RNA helicase Gu) (RH II/Gu) (Gu-alpha) (DEAD box protein 21). | |||||
|
CLSPN_HUMAN
|
||||||
| NC score | 0.022613 (rank : 26) | θ value | 4.03905 (rank : 28) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9HAW4, Q5VYG0, Q6P6H5, Q8IWI1 | Gene names | CLSPN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Claspin (hClaspin) (Hu-Claspin). | |||||
|
PCDBC_HUMAN
|
||||||
| NC score | 0.022150 (rank : 27) | θ value | 0.813845 (rank : 9) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Y5F1 | Gene names | PCDHB12 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin beta 12 precursor (PCDH-beta12). | |||||
|
PCDB3_HUMAN
|
||||||
| NC score | 0.021368 (rank : 28) | θ value | 1.81305 (rank : 16) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Y5E6 | Gene names | PCDHB3 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin beta 3 precursor (PCDH-beta3). | |||||
|
COAA1_HUMAN
|
||||||
| NC score | 0.020819 (rank : 29) | θ value | 1.38821 (rank : 12) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 316 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q03692 | Gene names | COL10A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(X) chain precursor. | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.020362 (rank : 30) | θ value | 6.88961 (rank : 42) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
PCDB7_HUMAN
|
||||||
| NC score | 0.019947 (rank : 31) | θ value | 5.27518 (rank : 36) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Y5E2 | Gene names | PCDHB7 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin beta 7 precursor (PCDH-beta7). | |||||
|
PCDB2_HUMAN
|
||||||
| NC score | 0.019308 (rank : 32) | θ value | 8.99809 (rank : 49) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Y5E7 | Gene names | PCDHB2 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin beta 2 precursor (PCDH-beta2). | |||||
|
PCDBA_HUMAN
|
||||||
| NC score | 0.019299 (rank : 33) | θ value | 8.99809 (rank : 54) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9UN67, Q96T99 | Gene names | PCDHB10 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin beta 10 precursor (PCDH-beta10). | |||||
|
PCDB5_HUMAN
|
||||||
| NC score | 0.019295 (rank : 34) | θ value | 8.99809 (rank : 51) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y5E4, Q9UFU9 | Gene names | PCDHB5 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin beta 5 precursor (PCDH-beta5). | |||||
|
PCDB4_HUMAN
|
||||||
| NC score | 0.019274 (rank : 35) | θ value | 8.99809 (rank : 50) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Y5E5 | Gene names | PCDHB4 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin beta 4 precursor (PCDH-beta4). | |||||
|
PCDB9_HUMAN
|
||||||
| NC score | 0.019225 (rank : 36) | θ value | 8.99809 (rank : 53) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y5E1, Q9NRJ8 | Gene names | PCDHB9, PCDH3H | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin beta 9 precursor (PCDH-beta9) (Protocadherin 3H). | |||||
|
MYCB2_MOUSE
|
||||||
| NC score | 0.019200 (rank : 37) | θ value | 6.88961 (rank : 41) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q7TPH6, Q6PCM8 | Gene names | Mycbp2, Pam, Phr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ubiquitin ligase protein MYCBP2 (EC 6.3.2.-) (Myc-binding protein 2) (Protein associated with Myc) (Pam/highwire/rpm-1 protein). | |||||
|
PCDB8_HUMAN
|
||||||
| NC score | 0.019192 (rank : 38) | θ value | 8.99809 (rank : 52) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9UN66 | Gene names | PCDHB8, PCDH3I | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin beta 8 precursor (PCDH-beta8) (Protocadherin 3I). | |||||
|
PCDBG_HUMAN
|
||||||
| NC score | 0.019173 (rank : 39) | θ value | 8.99809 (rank : 55) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NRJ7, Q8IYD5, Q96SE9, Q9HCF1 | Gene names | PCDHB16, KIAA1621, PCDH3X | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin beta 16 precursor (PCDH-beta16) (Protocadherin 3X). | |||||
|
AP3B1_MOUSE
|
||||||
| NC score | 0.019028 (rank : 40) | θ value | 4.03905 (rank : 24) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Z1T1, Q91YR4 | Gene names | Ap3b1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-1 (Adapter-related protein complex 3 beta-1 subunit) (Beta3A-adaptin) (Adaptor protein complex AP-3 beta-1 subunit) (Clathrin assembly protein complex 3 beta-1 large chain). | |||||
|
MAGB2_HUMAN
|
||||||
| NC score | 0.018606 (rank : 41) | θ value | 2.36792 (rank : 18) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O15479, O75860 | Gene names | MAGEB2 | |||
|
Domain Architecture |

|
|||||
| Description | Melanoma-associated antigen B2 (MAGE-B2 antigen) (DSS-AHC critical interval MAGE superfamily 6) (DAM6) (MAGE XP-2). | |||||
|
NOL8_MOUSE
|
||||||
| NC score | 0.016583 (rank : 42) | θ value | 6.88961 (rank : 44) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q3UHX0, Q504M4, Q8CDJ7, Q9CUR0 | Gene names | Nol8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleolar protein 8. | |||||
|
SPG21_HUMAN
|
||||||
| NC score | 0.016549 (rank : 43) | θ value | 8.99809 (rank : 57) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NZD8, Q6ZMB6 | Gene names | SPG21, ACP33 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Maspardin (Spastic paraplegia 21 autosomal recessive Mast syndrome protein) (Acid cluster protein 33). | |||||
|
YETS2_MOUSE
|
||||||
| NC score | 0.016368 (rank : 44) | θ value | 8.99809 (rank : 62) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q3TUF7, Q6PGF8, Q80TI2, Q8CG86 | Gene names | Yeats2, Kiaa1197 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YEATS domain-containing protein 2. | |||||
|
MAGD1_HUMAN
|
||||||
| NC score | 0.015933 (rank : 45) | θ value | 3.0926 (rank : 21) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y5V3, Q5VSH6, Q8IZ84, Q8WY92, Q9H352, Q9HBT4, Q9UF36 | Gene names | MAGED1, NRAGE | |||
|
Domain Architecture |

|
|||||
| Description | Melanoma-associated antigen D1 (MAGE-D1 antigen) (Neurotrophin receptor-interacting MAGE homolog) (MAGE tumor antigen CCF). | |||||
|
CPNE7_HUMAN
|
||||||
| NC score | 0.014977 (rank : 46) | θ value | 5.27518 (rank : 32) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UBL6 | Gene names | CPNE7 | |||
|
Domain Architecture |

|
|||||
| Description | Copine-7 (Copine VII). | |||||
|
CXA5_MOUSE
|
||||||
| NC score | 0.014943 (rank : 47) | θ value | 3.0926 (rank : 20) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q01231 | Gene names | Gja5, Cxn-40 | |||
|
Domain Architecture |

|
|||||
| Description | Gap junction alpha-5 protein (Connexin-40) (Cx40). | |||||
|
TXD11_HUMAN
|
||||||
| NC score | 0.014143 (rank : 48) | θ value | 8.99809 (rank : 60) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6PKC3, O95887, Q6PJA6, Q8N2Q4, Q96K45, Q96K53 | Gene names | TXNDC11, EFP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 11 (EF-hand-binding protein 1). | |||||
|
CENPF_HUMAN
|
||||||
| NC score | 0.013814 (rank : 49) | θ value | 1.81305 (rank : 14) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1932 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P49454, Q13171, Q13246 | Gene names | CENPF | |||
|
Domain Architecture |

|
|||||
| Description | Centromere protein F (Kinetochore protein CENP-F) (Mitosin) (AH antigen). | |||||
|
PHF14_HUMAN
|
||||||
| NC score | 0.012911 (rank : 50) | θ value | 6.88961 (rank : 45) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O94880 | Gene names | PHF14, KIAA0783 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 14. | |||||
|
USP9X_HUMAN
|
||||||
| NC score | 0.012298 (rank : 51) | θ value | 8.99809 (rank : 61) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q93008, O75550, Q8WWT3, Q8WX12 | Gene names | USP9X, DFFRX, USP9 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase FAF-X (EC 3.1.2.15) (Ubiquitin thioesterase FAF-X) (Ubiquitin-specific-processing protease FAF-X) (Deubiquitinating enzyme FAF-X) (Fat facets protein-related, X- linked) (Ubiquitin-specific protease 9, X chromosome). | |||||
|
ANK2_HUMAN
|
||||||
| NC score | 0.010574 (rank : 52) | θ value | 2.36792 (rank : 17) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
AP4E1_HUMAN
|
||||||
| NC score | 0.010473 (rank : 53) | θ value | 8.99809 (rank : 47) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UPM8, Q9Y588 | Gene names | AP4E1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-4 complex subunit epsilon-1 (Adapter-related protein complex 4 epsilon-1 subunit) (Epsilon subunit of AP-4) (AP-4 adapter complex epsilon subunit). | |||||
|
PF21A_HUMAN
|
||||||
| NC score | 0.010342 (rank : 54) | θ value | 8.99809 (rank : 56) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96BD5, Q6AWA2, Q9C0G7, Q9H8V9, Q9HAK6, Q9NZE9 | Gene names | PHF21A, BHC80, KIAA1696 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (BHC80a). | |||||
|
EEA1_HUMAN
|
||||||
| NC score | 0.008357 (rank : 55) | θ value | 6.88961 (rank : 39) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1796 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q15075, Q14221 | Gene names | EEA1, ZFYVE2 | |||
|
Domain Architecture |

|
|||||
| Description | Early endosome antigen 1 (Endosome-associated protein p162) (Zinc finger FYVE domain-containing protein 2). | |||||
|
EGR1_MOUSE
|
||||||
| NC score | 0.007988 (rank : 56) | θ value | 1.38821 (rank : 13) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 816 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P08046, Q61777 | Gene names | Egr1, Egr-1, Krox-24 | |||
|
Domain Architecture |

|
|||||
| Description | Early growth response protein 1 (EGR-1) (Krox-24 protein) (Transcription factor Zif268) (Nerve growth factor-induced protein A) (NGFI-A). | |||||
|
PEG3_MOUSE
|
||||||
| NC score | 0.007763 (rank : 57) | θ value | 4.03905 (rank : 30) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1101 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q3URU2, O54978, Q3TQ69, Q5EBP7, Q61138, Q6GQS0, Q80U47, Q8R5N0, Q9QX53 | Gene names | Peg3, Kiaa0287, Pw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Paternally expressed gene 3 protein (ASF-1). | |||||
|
BMPR2_HUMAN
|
||||||
| NC score | 0.006193 (rank : 58) | θ value | 4.03905 (rank : 25) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 736 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q13873, Q16569 | Gene names | BMPR2, PPH1 | |||
|
Domain Architecture |

|
|||||
| Description | Bone morphogenetic protein receptor type-2 precursor (EC 2.7.11.30) (Bone morphogenetic protein receptor type II) (BMP type II receptor) (BMPR-II). | |||||
|
KIF22_HUMAN
|
||||||
| NC score | 0.004856 (rank : 59) | θ value | 8.99809 (rank : 48) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q14807, O94814, Q9BT46 | Gene names | KIF22, KID, KNSL4 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF22 (Kinesin-like DNA-binding protein) (Kinesin-like protein 4). | |||||
|
MARK4_MOUSE
|
||||||
| NC score | 0.002697 (rank : 60) | θ value | 5.27518 (rank : 35) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 898 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8CIP4, Q80T81 | Gene names | Mark4, Kiaa1860 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MAP/microtubule affinity-regulating kinase 4 (EC 2.7.11.1). | |||||
|
TITIN_HUMAN
|
||||||
| NC score | 0.002563 (rank : 61) | θ value | 8.99809 (rank : 59) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
MARK4_HUMAN
|
||||||
| NC score | 0.002440 (rank : 62) | θ value | 6.88961 (rank : 40) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 886 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96L34, Q8NG37, Q96JG7, Q96SQ2, Q9BYD8 | Gene names | MARK4, KIAA1860, MARKL1 | |||
|
Domain Architecture |

|
|||||
| Description | MAP/microtubule affinity-regulating kinase 4 (EC 2.7.11.1) (MAP/microtubule affinity-regulating kinase-like 1). | |||||