Please be patient as the page loads
|
IFI3_MOUSE
|
||||||
| SwissProt Accessions | O35368 | Gene names | Ifi203 | |||
|
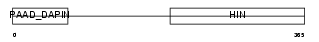
Domain Architecture |
|
|||||
| Description | Interferon-activable protein 203 (Ifi-203) (Interferon-inducible protein p203). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
IFI3_MOUSE
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O35368 | Gene names | Ifi203 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon-activable protein 203 (Ifi-203) (Interferon-inducible protein p203). | |||||
|
IFI4_MOUSE
|
||||||
| θ value | 1.18275e-92 (rank : 2) | NC score | 0.958266 (rank : 2) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P15092 | Gene names | Ifi204 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon-activable protein 204 (Ifi-204) (Interferon-inducible protein p204). | |||||
|
IF16_HUMAN
|
||||||
| θ value | 3.58603e-65 (rank : 3) | NC score | 0.919201 (rank : 8) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 182 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q16666, Q9UH78 | Gene names | IFI16, IFNGIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Gamma-interferon-inducible protein Ifi-16 (Interferon-inducible myeloid differentiation transcriptional activator) (IFI 16). | |||||
|
MNDA_HUMAN
|
||||||
| θ value | 6.76295e-64 (rank : 4) | NC score | 0.945860 (rank : 3) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P41218 | Gene names | MNDA | |||
|
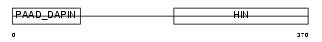
Domain Architecture |
|
|||||
| Description | Myeloid cell nuclear differentiation antigen. | |||||
|
II2B_MOUSE
|
||||||
| θ value | 3.60269e-49 (rank : 5) | NC score | 0.921500 (rank : 6) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9R002 | Gene names | Ifi202b | |||
|
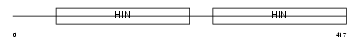
Domain Architecture |
|
|||||
| Description | Interferon-activable protein 202b (Ifi-202b) (Interferon-inducible protein p202b). | |||||
|
II2A_MOUSE
|
||||||
| θ value | 3.98324e-48 (rank : 6) | NC score | 0.921322 (rank : 7) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P15091 | Gene names | Ifi202a, Ifi202 | |||
|
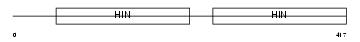
Domain Architecture |
|
|||||
| Description | Interferon-activable protein 202a (Ifi-202a) (Interferon-inducible protein p202a). | |||||
|
IFI5_MOUSE
|
||||||
| θ value | 1.67352e-46 (rank : 7) | NC score | 0.943587 (rank : 4) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q08619 | Gene names | Ifi205 | |||
|
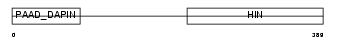
Domain Architecture |
|
|||||
| Description | Interferon-activable protein 205 (IFI-205) (D3 protein). | |||||
|
AIM2_HUMAN
|
||||||
| θ value | 1.05066e-40 (rank : 8) | NC score | 0.924964 (rank : 5) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O14862 | Gene names | AIM2 | |||
|
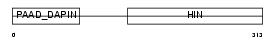
Domain Architecture |
|
|||||
| Description | Interferon-inducible protein AIM2 (Absent in melanoma 2). | |||||
|
REST_HUMAN
|
||||||
| θ value | 0.163984 (rank : 9) | NC score | 0.015776 (rank : 26) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 1605 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P30622 | Gene names | RSN, CYLN1 | |||
|
Domain Architecture |

|
|||||
| Description | Restin (Cytoplasmic linker protein 170 alpha-2) (CLIP-170) (Reed- Sternberg intermediate filament-associated protein) (Cytoplasmic linker protein 1). | |||||
|
CD008_HUMAN
|
||||||
| θ value | 0.279714 (rank : 10) | NC score | 0.052270 (rank : 9) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P78312, O43607, P78311, P78313, Q9UEG8 | Gene names | C4orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C4orf8 (Protein IT14). | |||||
|
NIPBL_HUMAN
|
||||||
| θ value | 0.365318 (rank : 11) | NC score | 0.023462 (rank : 22) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
NTBP1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 12) | NC score | 0.035385 (rank : 12) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 528 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BRM2, Q8CAA8 | Gene names | Scyl1bp1, Ntklbp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-terminal kinase-like-binding protein 1 (NTKL-binding protein 1) (NTKL-BP1) (mNTKL-BP1) (SCY1-like 1-binding protein 1) (SCYL1-binding protein 1) (SCYL1-BP1). | |||||
|
PHF20_MOUSE
|
||||||
| θ value | 0.47712 (rank : 13) | NC score | 0.039570 (rank : 10) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8BLG0, Q8BMA2, Q8BYR4, Q921N1 | Gene names | Phf20, Hca58 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58 homolog). | |||||
|
C8AP2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 14) | NC score | 0.033803 (rank : 13) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 405 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9WUF3, Q9CSW2 | Gene names | Casp8ap2, Flash | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
SMRC1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 15) | NC score | 0.026183 (rank : 20) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 459 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P97496, Q7TS80, Q7TT29 | Gene names | Smarcc1, Baf155, Srg3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 1 (SWI/SNF complex 155 kDa subunit) (BRG1-associated factor 155) (SWI3-related protein). | |||||
|
CE290_HUMAN
|
||||||
| θ value | 1.06291 (rank : 16) | NC score | 0.015869 (rank : 25) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 1553 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O15078, Q1PSK5, Q66GS8, Q9H2G6, Q9H6Q7, Q9H8I0 | Gene names | CEP290, KIAA0373, NPHP6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein Cep290 (Nephrocystin-6) (Tumor antigen se2-2). | |||||
|
CN145_HUMAN
|
||||||
| θ value | 1.06291 (rank : 17) | NC score | 0.014318 (rank : 29) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 1196 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6ZU80, Q96ML4 | Gene names | C14orf145 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf145. | |||||
|
RFC1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 18) | NC score | 0.028552 (rank : 16) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P35251, Q5XKF5, Q6PKU0, Q86V41, Q86V46 | Gene names | RFC1, RFC140 | |||
|
Domain Architecture |

|
|||||
| Description | Replication factor C subunit 1 (Replication factor C large subunit) (RF-C 140 kDa subunit) (Activator 1 140 kDa subunit) (Activator 1 large subunit) (A1 140 kDa subunit) (DNA-binding protein PO-GA). | |||||
|
K1210_HUMAN
|
||||||
| θ value | 1.38821 (rank : 19) | NC score | 0.033477 (rank : 14) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9ULL0, Q5JPN4 | Gene names | KIAA1210 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1210. | |||||
|
OBSCN_HUMAN
|
||||||
| θ value | 1.38821 (rank : 20) | NC score | 0.002175 (rank : 46) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 1931 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5VST9, Q2A664, Q5T7G8, Q5T7G9, Q5VSU2, Q86YC7, Q8NHN0, Q8NHN1, Q8NHN2, Q8NHN4, Q8NHN5, Q8NHN6, Q8NHN7, Q8NHN8, Q8NHN9, Q96AA2, Q9HCD3, Q9HCL6 | Gene names | OBSCN, KIAA1556, KIAA1639 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Obscurin (Obscurin-myosin light chain kinase) (Obscurin-MLCK) (Obscurin-RhoGEF). | |||||
|
MLZE_HUMAN
|
||||||
| θ value | 1.81305 (rank : 21) | NC score | 0.026835 (rank : 19) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BYG8, Q5XKF3, Q6P494 | Gene names | MLZE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-derived leucine zipper-containing extranuclear factor. | |||||
|
ICAL_HUMAN
|
||||||
| θ value | 2.36792 (rank : 22) | NC score | 0.027122 (rank : 18) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P20810, O95360, Q96D08, Q9H1Z5 | Gene names | CAST | |||
|
Domain Architecture |

|
|||||
| Description | Calpastatin (Calpain inhibitor) (Sperm BS-17 component). | |||||
|
MYO15_MOUSE
|
||||||
| θ value | 3.0926 (rank : 23) | NC score | 0.008473 (rank : 42) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9QZZ4, O70395, Q9QWL6 | Gene names | Myo15a, Myo15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
PHF20_HUMAN
|
||||||
| θ value | 3.0926 (rank : 24) | NC score | 0.035863 (rank : 11) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9BVI0, Q5JWY9, Q9BWV4, Q9BXA3, Q9BZW3, Q9H421, Q9H4J6, Q9NZ22 | Gene names | PHF20, C20orf104, GLEA2, HCA58, NZF, TZP | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58) (Glioma-expressed antigen 2) (Transcription factor TZP) (Novel zinc finger protein). | |||||
|
ZCPW1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 25) | NC score | 0.031767 (rank : 15) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6IR42 | Gene names | Zcwpw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CW-type PWWP domain protein 1 homolog. | |||||
|
BAZ1B_HUMAN
|
||||||
| θ value | 4.03905 (rank : 26) | NC score | 0.015028 (rank : 27) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 701 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9UIG0, O95039, O95247, O95277 | Gene names | BAZ1B, WBSC10, WBSCR9, WSTF | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein) (Williams syndrome transcription factor) (hWALP2). | |||||
|
BRCA2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 27) | NC score | 0.017773 (rank : 24) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P97929, O35922, P97383 | Gene names | Brca2, Fancd1 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 2 susceptibility protein homolog (Fanconi anemia group D1 protein homolog). | |||||
|
CCDC7_HUMAN
|
||||||
| θ value | 4.03905 (rank : 28) | NC score | 0.024641 (rank : 21) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 253 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96M83, Q8IVQ0, Q8NEQ0 | Gene names | CCDC7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 7. | |||||
|
CMGA_HUMAN
|
||||||
| θ value | 4.03905 (rank : 29) | NC score | 0.019497 (rank : 23) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P10645, Q96E84, Q96GL7, Q9BQB5 | Gene names | CHGA | |||
|
Domain Architecture |

|
|||||
| Description | Chromogranin A precursor (CgA) (Pituitary secretory protein I) (SP-I) [Contains: Vasostatin-1 (Vasostatin I); Vasostatin-2 (Vasostatin II); EA-92; ES-43; Pancreastatin; SS-18; WA-8; WE-14; LF-19; AL-11; GV-19; GR-44; ER-37]. | |||||
|
EYA2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 30) | NC score | 0.014531 (rank : 28) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O08575, P97925 | Gene names | Eya2, Eab1 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 2 (EC 3.1.3.48). | |||||
|
MBB1A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 31) | NC score | 0.028465 (rank : 17) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 665 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BQG0, Q86VM3, Q9BW49, Q9P0V5, Q9UF99 | Gene names | MYBBP1A, P160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myb-binding protein 1A. | |||||
|
ANR11_HUMAN
|
||||||
| θ value | 5.27518 (rank : 32) | NC score | 0.013557 (rank : 30) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 1289 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6UB99, Q6NTG1, Q6QMF8 | Gene names | ANKRD11, ANCO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 11 (Ankyrin repeat-containing cofactor 1). | |||||
|
FAK2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 33) | NC score | 0.000136 (rank : 47) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QVP9 | Gene names | Ptk2b, Fak2, Pyk2, Raftk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein tyrosine kinase 2 beta (EC 2.7.10.2) (Focal adhesion kinase 2) (FADK 2) (Proline-rich tyrosine kinase 2) (Cell adhesion kinase beta) (CAK beta) (Calcium-dependent tyrosine kinase) (CADTK) (Related adhesion focal tyrosine kinase) (RAFTK). | |||||
|
GOGA6_HUMAN
|
||||||
| θ value | 5.27518 (rank : 34) | NC score | 0.012529 (rank : 33) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 1007 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9NYA3, Q9NYA7 | Gene names | GOLGA6, GLP | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 6 (Golgin linked to PML) (Golgin-like protein). | |||||
|
LPIN2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 35) | NC score | 0.010020 (rank : 40) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q92539 | Gene names | LPIN2, KIAA0249 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipin-2. | |||||
|
RAD50_HUMAN
|
||||||
| θ value | 5.27518 (rank : 36) | NC score | 0.012944 (rank : 31) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 1317 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q92878, O43254, Q6GMT7, Q6P5X3, Q9UP86 | Gene names | RAD50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (hRAD50). | |||||
|
RAD50_MOUSE
|
||||||
| θ value | 5.27518 (rank : 37) | NC score | 0.012786 (rank : 32) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 1366 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P70388, Q6PEB0, Q8C2T7, Q9CU59 | Gene names | Rad50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (mRad50). | |||||
|
CAR10_MOUSE
|
||||||
| θ value | 6.88961 (rank : 38) | NC score | 0.010076 (rank : 39) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 407 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P58660 | Gene names | Card10, Bimp1 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 10 (Bcl10-interacting MAGUK protein 1) (Bimp1). | |||||
|
LYAM3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 39) | NC score | 0.004081 (rank : 45) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 378 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P16109, Q8IVD1 | Gene names | SELP, GMRP | |||
|
Domain Architecture |

|
|||||
| Description | P-selectin precursor (Granule membrane protein 140) (GMP-140) (PADGEM) (Leukocyte-endothelial cell adhesion molecule 3) (LECAM3) (CD62P antigen). | |||||
|
SMC4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 40) | NC score | 0.011449 (rank : 36) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 937 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9NTJ3, O95752, Q8NDL4, Q9UNT9 | Gene names | SMC4L1, CAPC, SMC4 | |||
|
Domain Architecture |
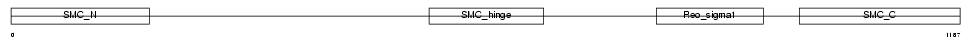
|
|||||
| Description | Structural maintenance of chromosomes 4-like 1 protein (Chromosome- associated polypeptide C) (hCAP-C) (XCAP-C homolog). | |||||
|
UBP1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 41) | NC score | 0.011041 (rank : 37) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O94782, Q9UFR0, Q9UNJ3 | Gene names | USP1 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 1 (EC 3.1.2.15) (Ubiquitin thioesterase 1) (Ubiquitin-specific-processing protease 1) (Deubiquitinating enzyme 1) (hUBP). | |||||
|
CCP2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 42) | NC score | 0.008652 (rank : 41) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JHG2 | Gene names | Dscr1l1 | |||
|
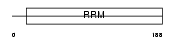
Domain Architecture |
|
|||||
| Description | Calcipressin-2 (Down syndrome candidate region 1-like protein 1) (Myocyte-enriched calcineurin-interacting protein 2) (MCIP2). | |||||
|
CEP70_MOUSE
|
||||||
| θ value | 8.99809 (rank : 43) | NC score | 0.012013 (rank : 35) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 532 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6IQY5, Q9CRL9, Q9CTS4, Q9JIC1 | Gene names | Cep70 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 70 kDa (Cep70 protein). | |||||
|
EYA2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 44) | NC score | 0.012346 (rank : 34) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O00167, Q5JSW8, Q96CV6, Q96H97, Q99503, Q99812, Q9BWF6, Q9H4S3, Q9H4S9, Q9NPZ4, Q9UIX7 | Gene names | EYA2, EAB1 | |||
|
Domain Architecture |
|
|||||
| Description | Eyes absent homolog 2 (EC 3.1.3.48). | |||||
|
MYO15_HUMAN
|
||||||
| θ value | 8.99809 (rank : 45) | NC score | 0.006648 (rank : 43) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UKN7 | Gene names | MYO15A, MYO15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
SMRA3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 46) | NC score | 0.005077 (rank : 44) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 265 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14527, Q14536, Q16051, Q7KYJ6, Q86YA5, Q92652, Q96KM9 | Gene names | SMARCA3, HIP116A, HLTF, SNF2L3, ZBU1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 3 (EC 3.6.1.-) (Sucrose nonfermenting protein 2-like 3) (DNA-binding protein/plasminogen activator inhibitor 1 regulator) (Helicase-like transcription factor) (HIP116). | |||||
|
TTC16_HUMAN
|
||||||
| θ value | 8.99809 (rank : 47) | NC score | 0.010633 (rank : 38) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NEE8, Q5JU66, Q96M72 | Gene names | TTC16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tetratricopeptide repeat protein 16 (TPR repeat protein 16). | |||||
|
IFI3_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O35368 | Gene names | Ifi203 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon-activable protein 203 (Ifi-203) (Interferon-inducible protein p203). | |||||
|
IFI4_MOUSE
|
||||||
| NC score | 0.958266 (rank : 2) | θ value | 1.18275e-92 (rank : 2) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P15092 | Gene names | Ifi204 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon-activable protein 204 (Ifi-204) (Interferon-inducible protein p204). | |||||
|
MNDA_HUMAN
|
||||||
| NC score | 0.945860 (rank : 3) | θ value | 6.76295e-64 (rank : 4) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P41218 | Gene names | MNDA | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid cell nuclear differentiation antigen. | |||||
|
IFI5_MOUSE
|
||||||
| NC score | 0.943587 (rank : 4) | θ value | 1.67352e-46 (rank : 7) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q08619 | Gene names | Ifi205 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon-activable protein 205 (IFI-205) (D3 protein). | |||||
|
AIM2_HUMAN
|
||||||
| NC score | 0.924964 (rank : 5) | θ value | 1.05066e-40 (rank : 8) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O14862 | Gene names | AIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon-inducible protein AIM2 (Absent in melanoma 2). | |||||
|
II2B_MOUSE
|
||||||
| NC score | 0.921500 (rank : 6) | θ value | 3.60269e-49 (rank : 5) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9R002 | Gene names | Ifi202b | |||
|
Domain Architecture |

|
|||||
| Description | Interferon-activable protein 202b (Ifi-202b) (Interferon-inducible protein p202b). | |||||
|
II2A_MOUSE
|
||||||
| NC score | 0.921322 (rank : 7) | θ value | 3.98324e-48 (rank : 6) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P15091 | Gene names | Ifi202a, Ifi202 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon-activable protein 202a (Ifi-202a) (Interferon-inducible protein p202a). | |||||
|
IF16_HUMAN
|
||||||
| NC score | 0.919201 (rank : 8) | θ value | 3.58603e-65 (rank : 3) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 182 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q16666, Q9UH78 | Gene names | IFI16, IFNGIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Gamma-interferon-inducible protein Ifi-16 (Interferon-inducible myeloid differentiation transcriptional activator) (IFI 16). | |||||
|
CD008_HUMAN
|
||||||
| NC score | 0.052270 (rank : 9) | θ value | 0.279714 (rank : 10) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P78312, O43607, P78311, P78313, Q9UEG8 | Gene names | C4orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C4orf8 (Protein IT14). | |||||
|
PHF20_MOUSE
|
||||||
| NC score | 0.039570 (rank : 10) | θ value | 0.47712 (rank : 13) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8BLG0, Q8BMA2, Q8BYR4, Q921N1 | Gene names | Phf20, Hca58 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58 homolog). | |||||
|
PHF20_HUMAN
|
||||||
| NC score | 0.035863 (rank : 11) | θ value | 3.0926 (rank : 24) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9BVI0, Q5JWY9, Q9BWV4, Q9BXA3, Q9BZW3, Q9H421, Q9H4J6, Q9NZ22 | Gene names | PHF20, C20orf104, GLEA2, HCA58, NZF, TZP | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58) (Glioma-expressed antigen 2) (Transcription factor TZP) (Novel zinc finger protein). | |||||
|
NTBP1_MOUSE
|
||||||
| NC score | 0.035385 (rank : 12) | θ value | 0.365318 (rank : 12) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 528 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BRM2, Q8CAA8 | Gene names | Scyl1bp1, Ntklbp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-terminal kinase-like-binding protein 1 (NTKL-binding protein 1) (NTKL-BP1) (mNTKL-BP1) (SCY1-like 1-binding protein 1) (SCYL1-binding protein 1) (SCYL1-BP1). | |||||
|
C8AP2_MOUSE
|
||||||
| NC score | 0.033803 (rank : 13) | θ value | 0.62314 (rank : 14) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 405 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9WUF3, Q9CSW2 | Gene names | Casp8ap2, Flash | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
K1210_HUMAN
|
||||||
| NC score | 0.033477 (rank : 14) | θ value | 1.38821 (rank : 19) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9ULL0, Q5JPN4 | Gene names | KIAA1210 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1210. | |||||
|
ZCPW1_MOUSE
|
||||||
| NC score | 0.031767 (rank : 15) | θ value | 3.0926 (rank : 25) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6IR42 | Gene names | Zcwpw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CW-type PWWP domain protein 1 homolog. | |||||
|
RFC1_HUMAN
|
||||||
| NC score | 0.028552 (rank : 16) | θ value | 1.06291 (rank : 18) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P35251, Q5XKF5, Q6PKU0, Q86V41, Q86V46 | Gene names | RFC1, RFC140 | |||
|
Domain Architecture |

|
|||||
| Description | Replication factor C subunit 1 (Replication factor C large subunit) (RF-C 140 kDa subunit) (Activator 1 140 kDa subunit) (Activator 1 large subunit) (A1 140 kDa subunit) (DNA-binding protein PO-GA). | |||||
|
MBB1A_HUMAN
|
||||||
| NC score | 0.028465 (rank : 17) | θ value | 4.03905 (rank : 31) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 665 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BQG0, Q86VM3, Q9BW49, Q9P0V5, Q9UF99 | Gene names | MYBBP1A, P160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myb-binding protein 1A. | |||||
|
ICAL_HUMAN
|
||||||
| NC score | 0.027122 (rank : 18) | θ value | 2.36792 (rank : 22) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P20810, O95360, Q96D08, Q9H1Z5 | Gene names | CAST | |||
|
Domain Architecture |

|
|||||
| Description | Calpastatin (Calpain inhibitor) (Sperm BS-17 component). | |||||
|
MLZE_HUMAN
|
||||||
| NC score | 0.026835 (rank : 19) | θ value | 1.81305 (rank : 21) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BYG8, Q5XKF3, Q6P494 | Gene names | MLZE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-derived leucine zipper-containing extranuclear factor. | |||||
|
SMRC1_MOUSE
|
||||||
| NC score | 0.026183 (rank : 20) | θ value | 0.62314 (rank : 15) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 459 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P97496, Q7TS80, Q7TT29 | Gene names | Smarcc1, Baf155, Srg3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 1 (SWI/SNF complex 155 kDa subunit) (BRG1-associated factor 155) (SWI3-related protein). | |||||
|
CCDC7_HUMAN
|
||||||
| NC score | 0.024641 (rank : 21) | θ value | 4.03905 (rank : 28) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 253 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96M83, Q8IVQ0, Q8NEQ0 | Gene names | CCDC7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 7. | |||||
|
NIPBL_HUMAN
|
||||||
| NC score | 0.023462 (rank : 22) | θ value | 0.365318 (rank : 11) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
CMGA_HUMAN
|
||||||
| NC score | 0.019497 (rank : 23) | θ value | 4.03905 (rank : 29) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P10645, Q96E84, Q96GL7, Q9BQB5 | Gene names | CHGA | |||
|
Domain Architecture |

|
|||||
| Description | Chromogranin A precursor (CgA) (Pituitary secretory protein I) (SP-I) [Contains: Vasostatin-1 (Vasostatin I); Vasostatin-2 (Vasostatin II); EA-92; ES-43; Pancreastatin; SS-18; WA-8; WE-14; LF-19; AL-11; GV-19; GR-44; ER-37]. | |||||
|
BRCA2_MOUSE
|
||||||
| NC score | 0.017773 (rank : 24) | θ value | 4.03905 (rank : 27) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P97929, O35922, P97383 | Gene names | Brca2, Fancd1 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 2 susceptibility protein homolog (Fanconi anemia group D1 protein homolog). | |||||
|
CE290_HUMAN
|
||||||
| NC score | 0.015869 (rank : 25) | θ value | 1.06291 (rank : 16) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 1553 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O15078, Q1PSK5, Q66GS8, Q9H2G6, Q9H6Q7, Q9H8I0 | Gene names | CEP290, KIAA0373, NPHP6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein Cep290 (Nephrocystin-6) (Tumor antigen se2-2). | |||||
|
REST_HUMAN
|
||||||
| NC score | 0.015776 (rank : 26) | θ value | 0.163984 (rank : 9) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 1605 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P30622 | Gene names | RSN, CYLN1 | |||
|
Domain Architecture |

|
|||||
| Description | Restin (Cytoplasmic linker protein 170 alpha-2) (CLIP-170) (Reed- Sternberg intermediate filament-associated protein) (Cytoplasmic linker protein 1). | |||||
|
BAZ1B_HUMAN
|
||||||
| NC score | 0.015028 (rank : 27) | θ value | 4.03905 (rank : 26) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 701 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9UIG0, O95039, O95247, O95277 | Gene names | BAZ1B, WBSC10, WBSCR9, WSTF | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein) (Williams syndrome transcription factor) (hWALP2). | |||||
|
EYA2_MOUSE
|
||||||
| NC score | 0.014531 (rank : 28) | θ value | 4.03905 (rank : 30) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O08575, P97925 | Gene names | Eya2, Eab1 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 2 (EC 3.1.3.48). | |||||
|
CN145_HUMAN
|
||||||
| NC score | 0.014318 (rank : 29) | θ value | 1.06291 (rank : 17) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 1196 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6ZU80, Q96ML4 | Gene names | C14orf145 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf145. | |||||
|
ANR11_HUMAN
|
||||||
| NC score | 0.013557 (rank : 30) | θ value | 5.27518 (rank : 32) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 1289 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6UB99, Q6NTG1, Q6QMF8 | Gene names | ANKRD11, ANCO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 11 (Ankyrin repeat-containing cofactor 1). | |||||
|
RAD50_HUMAN
|
||||||
| NC score | 0.012944 (rank : 31) | θ value | 5.27518 (rank : 36) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 1317 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q92878, O43254, Q6GMT7, Q6P5X3, Q9UP86 | Gene names | RAD50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (hRAD50). | |||||
|
RAD50_MOUSE
|
||||||
| NC score | 0.012786 (rank : 32) | θ value | 5.27518 (rank : 37) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 1366 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P70388, Q6PEB0, Q8C2T7, Q9CU59 | Gene names | Rad50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (mRad50). | |||||
|
GOGA6_HUMAN
|
||||||
| NC score | 0.012529 (rank : 33) | θ value | 5.27518 (rank : 34) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 1007 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9NYA3, Q9NYA7 | Gene names | GOLGA6, GLP | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 6 (Golgin linked to PML) (Golgin-like protein). | |||||
|
EYA2_HUMAN
|
||||||
| NC score | 0.012346 (rank : 34) | θ value | 8.99809 (rank : 44) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O00167, Q5JSW8, Q96CV6, Q96H97, Q99503, Q99812, Q9BWF6, Q9H4S3, Q9H4S9, Q9NPZ4, Q9UIX7 | Gene names | EYA2, EAB1 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 2 (EC 3.1.3.48). | |||||
|
CEP70_MOUSE
|
||||||
| NC score | 0.012013 (rank : 35) | θ value | 8.99809 (rank : 43) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 532 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6IQY5, Q9CRL9, Q9CTS4, Q9JIC1 | Gene names | Cep70 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 70 kDa (Cep70 protein). | |||||
|
SMC4_HUMAN
|
||||||
| NC score | 0.011449 (rank : 36) | θ value | 6.88961 (rank : 40) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 937 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9NTJ3, O95752, Q8NDL4, Q9UNT9 | Gene names | SMC4L1, CAPC, SMC4 | |||
|
Domain Architecture |
|
|||||
| Description | Structural maintenance of chromosomes 4-like 1 protein (Chromosome- associated polypeptide C) (hCAP-C) (XCAP-C homolog). | |||||
|
UBP1_HUMAN
|
||||||
| NC score | 0.011041 (rank : 37) | θ value | 6.88961 (rank : 41) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O94782, Q9UFR0, Q9UNJ3 | Gene names | USP1 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 1 (EC 3.1.2.15) (Ubiquitin thioesterase 1) (Ubiquitin-specific-processing protease 1) (Deubiquitinating enzyme 1) (hUBP). | |||||
|
TTC16_HUMAN
|
||||||
| NC score | 0.010633 (rank : 38) | θ value | 8.99809 (rank : 47) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NEE8, Q5JU66, Q96M72 | Gene names | TTC16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tetratricopeptide repeat protein 16 (TPR repeat protein 16). | |||||
|
CAR10_MOUSE
|
||||||
| NC score | 0.010076 (rank : 39) | θ value | 6.88961 (rank : 38) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 407 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P58660 | Gene names | Card10, Bimp1 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 10 (Bcl10-interacting MAGUK protein 1) (Bimp1). | |||||
|
LPIN2_HUMAN
|
||||||
| NC score | 0.010020 (rank : 40) | θ value | 5.27518 (rank : 35) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q92539 | Gene names | LPIN2, KIAA0249 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipin-2. | |||||
|
CCP2_MOUSE
|
||||||
| NC score | 0.008652 (rank : 41) | θ value | 8.99809 (rank : 42) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JHG2 | Gene names | Dscr1l1 | |||
|
Domain Architecture |

|
|||||
| Description | Calcipressin-2 (Down syndrome candidate region 1-like protein 1) (Myocyte-enriched calcineurin-interacting protein 2) (MCIP2). | |||||
|
MYO15_MOUSE
|
||||||
| NC score | 0.008473 (rank : 42) | θ value | 3.0926 (rank : 23) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9QZZ4, O70395, Q9QWL6 | Gene names | Myo15a, Myo15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
MYO15_HUMAN
|
||||||
| NC score | 0.006648 (rank : 43) | θ value | 8.99809 (rank : 45) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UKN7 | Gene names | MYO15A, MYO15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
SMRA3_HUMAN
|
||||||
| NC score | 0.005077 (rank : 44) | θ value | 8.99809 (rank : 46) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 265 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14527, Q14536, Q16051, Q7KYJ6, Q86YA5, Q92652, Q96KM9 | Gene names | SMARCA3, HIP116A, HLTF, SNF2L3, ZBU1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 3 (EC 3.6.1.-) (Sucrose nonfermenting protein 2-like 3) (DNA-binding protein/plasminogen activator inhibitor 1 regulator) (Helicase-like transcription factor) (HIP116). | |||||
|
LYAM3_HUMAN
|
||||||
| NC score | 0.004081 (rank : 45) | θ value | 6.88961 (rank : 39) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 378 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P16109, Q8IVD1 | Gene names | SELP, GMRP | |||
|
Domain Architecture |

|
|||||
| Description | P-selectin precursor (Granule membrane protein 140) (GMP-140) (PADGEM) (Leukocyte-endothelial cell adhesion molecule 3) (LECAM3) (CD62P antigen). | |||||
|
OBSCN_HUMAN
|
||||||
| NC score | 0.002175 (rank : 46) | θ value | 1.38821 (rank : 20) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 1931 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5VST9, Q2A664, Q5T7G8, Q5T7G9, Q5VSU2, Q86YC7, Q8NHN0, Q8NHN1, Q8NHN2, Q8NHN4, Q8NHN5, Q8NHN6, Q8NHN7, Q8NHN8, Q8NHN9, Q96AA2, Q9HCD3, Q9HCL6 | Gene names | OBSCN, KIAA1556, KIAA1639 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Obscurin (Obscurin-myosin light chain kinase) (Obscurin-MLCK) (Obscurin-RhoGEF). | |||||
|
FAK2_MOUSE
|
||||||
| NC score | 0.000136 (rank : 47) | θ value | 5.27518 (rank : 33) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QVP9 | Gene names | Ptk2b, Fak2, Pyk2, Raftk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein tyrosine kinase 2 beta (EC 2.7.10.2) (Focal adhesion kinase 2) (FADK 2) (Proline-rich tyrosine kinase 2) (Cell adhesion kinase beta) (CAK beta) (Calcium-dependent tyrosine kinase) (CADTK) (Related adhesion focal tyrosine kinase) (RAFTK). | |||||