Please be patient as the page loads
|
CRBB3_MOUSE
|
||||||
| SwissProt Accessions | Q9JJU9 | Gene names | Crybb3 | |||
|
Domain Architecture |
|
|||||
| Description | Beta crystallin B3 (Beta-B3-crystallin). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
CRBB3_MOUSE
|
||||||
| θ value | 2.36663e-117 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9JJU9 | Gene names | Crybb3 | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin B3 (Beta-B3-crystallin). | |||||
|
CRBB3_HUMAN
|
||||||
| θ value | 5.28454e-109 (rank : 2) | NC score | 0.999564 (rank : 2) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P26998, Q92965, Q9UH09 | Gene names | CRYBB3, CRYB3 | |||
|
Domain Architecture |
|
|||||
| Description | Beta crystallin B3 (Beta-B3-crystallin). | |||||
|
CRBB1_MOUSE
|
||||||
| θ value | 7.98882e-65 (rank : 3) | NC score | 0.991236 (rank : 4) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9WVJ5 | Gene names | Crybb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta crystallin B1 [Contains: Beta crystallin B1B]. | |||||
|
CRBB1_HUMAN
|
||||||
| θ value | 1.27544e-62 (rank : 4) | NC score | 0.991255 (rank : 3) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P53674 | Gene names | CRYBB1 | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin B1. | |||||
|
CRBB2_MOUSE
|
||||||
| θ value | 1.31987e-59 (rank : 5) | NC score | 0.990564 (rank : 5) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P62696, P19942, P26775 | Gene names | Crybb2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta crystallin B2 (Beta-crystallin Bp). | |||||
|
CRBB2_HUMAN
|
||||||
| θ value | 3.84025e-59 (rank : 6) | NC score | 0.990457 (rank : 6) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P43320, Q9UCM8 | Gene names | CRYBB2, CRYB2, CRYB2A | |||
|
Domain Architecture |
|
|||||
| Description | Beta crystallin B2 (Beta-crystallin Bp). | |||||
|
CRBA1_HUMAN
|
||||||
| θ value | 2.76489e-41 (rank : 7) | NC score | 0.966832 (rank : 7) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P05813, Q13633 | Gene names | CRYBA1, CRYB1 | |||
|
Domain Architecture |
|
|||||
| Description | Beta crystallin A3 [Contains: Beta crystallin A3, isoform A1, Delta4 form; Beta crystallin A3, isoform A1, Delta7 form; Beta crystallin A3, isoform A1, Delta8 form]. | |||||
|
CRBA1_MOUSE
|
||||||
| θ value | 2.86122e-38 (rank : 8) | NC score | 0.964174 (rank : 8) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P02525 | Gene names | Cryba1 | |||
|
Domain Architecture |
|
|||||
| Description | Beta crystallin A1 (Beta-A1-crystallin). | |||||
|
CRBA4_HUMAN
|
||||||
| θ value | 1.32909e-35 (rank : 9) | NC score | 0.958765 (rank : 9) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P53673, Q4VB22, Q6ICE4 | Gene names | CRYBA4 | |||
|
Domain Architecture |
|
|||||
| Description | Beta crystallin A4 (Beta-A4-crystallin). | |||||
|
CRBA4_MOUSE
|
||||||
| θ value | 2.26708e-35 (rank : 10) | NC score | 0.957508 (rank : 10) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9JJV0 | Gene names | Cryba4 | |||
|
Domain Architecture |
|
|||||
| Description | Beta crystallin A4 (Beta-A4-crystallin). | |||||
|
CRBA2_MOUSE
|
||||||
| θ value | 8.06329e-33 (rank : 11) | NC score | 0.956671 (rank : 11) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9JJV1 | Gene names | Cryba2 | |||
|
Domain Architecture |
|
|||||
| Description | Beta crystallin A2 (Beta-A2-crystallin). | |||||
|
CRBA2_HUMAN
|
||||||
| θ value | 4.00176e-32 (rank : 12) | NC score | 0.953899 (rank : 12) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P53672, Q9Y562 | Gene names | CRYBA2 | |||
|
Domain Architecture |
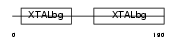
|
|||||
| Description | Beta crystallin A2 (Beta-A2-crystallin). | |||||
|
CRBS_HUMAN
|
||||||
| θ value | 3.17079e-29 (rank : 13) | NC score | 0.925480 (rank : 13) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P22914 | Gene names | CRYGS, GRYG8 | |||
|
Domain Architecture |
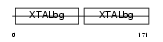
|
|||||
| Description | Beta crystallin S (Gamma crystallin S). | |||||
|
CRBS_MOUSE
|
||||||
| θ value | 1.2049e-28 (rank : 14) | NC score | 0.924438 (rank : 14) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O35486 | Gene names | Crygs | |||
|
Domain Architecture |
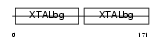
|
|||||
| Description | Beta crystallin S (Gamma crystallin S). | |||||
|
CRGB_MOUSE
|
||||||
| θ value | 1.73987e-27 (rank : 15) | NC score | 0.908760 (rank : 15) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P04344, Q61593 | Gene names | Crygb | |||
|
Domain Architecture |
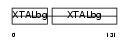
|
|||||
| Description | Gamma crystallin B (Gamma crystallin 3). | |||||
|
CRGA_MOUSE
|
||||||
| θ value | 6.61148e-27 (rank : 16) | NC score | 0.905064 (rank : 17) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P04345 | Gene names | Cryga | |||
|
Domain Architecture |
|
|||||
| Description | Gamma crystallin A (Gamma crystallin 4). | |||||
|
CRGC_HUMAN
|
||||||
| θ value | 6.61148e-27 (rank : 17) | NC score | 0.906971 (rank : 16) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P07315 | Gene names | CRYGC, CRYG3 | |||
|
Domain Architecture |
|
|||||
| Description | Gamma crystallin C (Gamma crystallin 2-1) (Gamma crystallin 3). | |||||
|
CRGB_HUMAN
|
||||||
| θ value | 9.54697e-26 (rank : 18) | NC score | 0.902027 (rank : 18) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P07316 | Gene names | CRYGB, CRYG2 | |||
|
Domain Architecture |
|
|||||
| Description | Gamma crystallin B (Gamma crystallin 1-2). | |||||
|
CRGA_HUMAN
|
||||||
| θ value | 3.62785e-25 (rank : 19) | NC score | 0.900832 (rank : 19) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P11844 | Gene names | CRYGA, CRYG1 | |||
|
Domain Architecture |
|
|||||
| Description | Gamma crystallin A (Gamma crystallin 5). | |||||
|
CRGC_MOUSE
|
||||||
| θ value | 3.62785e-25 (rank : 20) | NC score | 0.899525 (rank : 20) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q61597, Q03739 | Gene names | Crygc, Gammab-cry | |||
|
Domain Architecture |
|
|||||
| Description | Gamma crystallin C. | |||||
|
CRGD_MOUSE
|
||||||
| θ value | 1.05554e-24 (rank : 21) | NC score | 0.895953 (rank : 21) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P04342, O89027 | Gene names | Crygd | |||
|
Domain Architecture |
|
|||||
| Description | Gamma crystallin D (Gamma crystallin 1). | |||||
|
CRGF_MOUSE
|
||||||
| θ value | 3.07116e-24 (rank : 22) | NC score | 0.894648 (rank : 23) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9CXV3 | Gene names | Crygf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gamma crystallin F. | |||||
|
CRGE_MOUSE
|
||||||
| θ value | 5.23862e-24 (rank : 23) | NC score | 0.895675 (rank : 22) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q03740, O89028, P26999, Q9CXK5 | Gene names | Cryge | |||
|
Domain Architecture |
|
|||||
| Description | Gamma crystallin E. | |||||
|
CRGD_HUMAN
|
||||||
| θ value | 2.59989e-23 (rank : 24) | NC score | 0.892446 (rank : 24) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P07320, Q53R51, Q99681 | Gene names | CRYGD, CRYG4 | |||
|
Domain Architecture |
|
|||||
| Description | Gamma crystallin D (Gamma crystallin 4). | |||||
|
AIM1_HUMAN
|
||||||
| θ value | 1.74391e-19 (rank : 25) | NC score | 0.805081 (rank : 25) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9Y4K1, O00296, Q5VWJ2 | Gene names | AIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Absent in melanoma 1 protein. | |||||
|
TOP2B_HUMAN
|
||||||
| θ value | 1.81305 (rank : 26) | NC score | 0.017390 (rank : 26) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q02880, Q13600, Q9UMG8, Q9UQP8 | Gene names | TOP2B | |||
|
Domain Architecture |

|
|||||
| Description | DNA topoisomerase 2-beta (EC 5.99.1.3) (DNA topoisomerase II, beta isozyme). | |||||
|
ZKSC1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 27) | NC score | -0.002005 (rank : 32) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 755 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BGS3, Q7TS88, Q8BJ55, Q9CRN6 | Gene names | Zkscan1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger with KRAB and SCAN domain-containing protein 1. | |||||
|
TOP2B_MOUSE
|
||||||
| θ value | 4.03905 (rank : 28) | NC score | 0.016142 (rank : 27) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q64511, Q7TQG4 | Gene names | Top2b | |||
|
Domain Architecture |

|
|||||
| Description | DNA topoisomerase 2-beta (EC 5.99.1.3) (DNA topoisomerase II, beta isozyme). | |||||
|
ADA28_HUMAN
|
||||||
| θ value | 5.27518 (rank : 29) | NC score | 0.001725 (rank : 31) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UKQ2, Q9Y339, Q9Y3S0 | Gene names | ADAM28, MDCL | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 28 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 28) (Metalloproteinase-like, disintegrin-like, and cysteine- rich protein-L) (MDC-L) (eMDC II) (ADAM23). | |||||
|
NMNA3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 30) | NC score | 0.009183 (rank : 28) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96T66, Q8N4G1 | Gene names | NMNAT3 | |||
|
Domain Architecture |

|
|||||
| Description | Nicotinamide mononucleotide adenylyltransferase 3 (EC 2.7.7.1) (NMN adenylyltransferase 3). | |||||
|
DOCK4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 31) | NC score | 0.005930 (rank : 30) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 237 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8N1I0, O14584, O94824, Q8NB45 | Gene names | DOCK4, KIAA0716 | |||
|
Domain Architecture |

|
|||||
| Description | Dedicator of cytokinesis protein 4. | |||||
|
DOCK4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 32) | NC score | 0.005962 (rank : 29) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P59764 | Gene names | Dock4, Kiaa0716 | |||
|
Domain Architecture |

|
|||||
| Description | Dedicator of cytokinesis protein 4. | |||||
|
CRBB3_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 2.36663e-117 (rank : 1) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9JJU9 | Gene names | Crybb3 | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin B3 (Beta-B3-crystallin). | |||||
|
CRBB3_HUMAN
|
||||||
| NC score | 0.999564 (rank : 2) | θ value | 5.28454e-109 (rank : 2) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P26998, Q92965, Q9UH09 | Gene names | CRYBB3, CRYB3 | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin B3 (Beta-B3-crystallin). | |||||
|
CRBB1_HUMAN
|
||||||
| NC score | 0.991255 (rank : 3) | θ value | 1.27544e-62 (rank : 4) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P53674 | Gene names | CRYBB1 | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin B1. | |||||
|
CRBB1_MOUSE
|
||||||
| NC score | 0.991236 (rank : 4) | θ value | 7.98882e-65 (rank : 3) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9WVJ5 | Gene names | Crybb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta crystallin B1 [Contains: Beta crystallin B1B]. | |||||
|
CRBB2_MOUSE
|
||||||
| NC score | 0.990564 (rank : 5) | θ value | 1.31987e-59 (rank : 5) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P62696, P19942, P26775 | Gene names | Crybb2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta crystallin B2 (Beta-crystallin Bp). | |||||
|
CRBB2_HUMAN
|
||||||
| NC score | 0.990457 (rank : 6) | θ value | 3.84025e-59 (rank : 6) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P43320, Q9UCM8 | Gene names | CRYBB2, CRYB2, CRYB2A | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin B2 (Beta-crystallin Bp). | |||||
|
CRBA1_HUMAN
|
||||||
| NC score | 0.966832 (rank : 7) | θ value | 2.76489e-41 (rank : 7) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P05813, Q13633 | Gene names | CRYBA1, CRYB1 | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin A3 [Contains: Beta crystallin A3, isoform A1, Delta4 form; Beta crystallin A3, isoform A1, Delta7 form; Beta crystallin A3, isoform A1, Delta8 form]. | |||||
|
CRBA1_MOUSE
|
||||||
| NC score | 0.964174 (rank : 8) | θ value | 2.86122e-38 (rank : 8) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P02525 | Gene names | Cryba1 | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin A1 (Beta-A1-crystallin). | |||||
|
CRBA4_HUMAN
|
||||||
| NC score | 0.958765 (rank : 9) | θ value | 1.32909e-35 (rank : 9) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P53673, Q4VB22, Q6ICE4 | Gene names | CRYBA4 | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin A4 (Beta-A4-crystallin). | |||||
|
CRBA4_MOUSE
|
||||||
| NC score | 0.957508 (rank : 10) | θ value | 2.26708e-35 (rank : 10) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9JJV0 | Gene names | Cryba4 | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin A4 (Beta-A4-crystallin). | |||||
|
CRBA2_MOUSE
|
||||||
| NC score | 0.956671 (rank : 11) | θ value | 8.06329e-33 (rank : 11) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9JJV1 | Gene names | Cryba2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin A2 (Beta-A2-crystallin). | |||||
|
CRBA2_HUMAN
|
||||||
| NC score | 0.953899 (rank : 12) | θ value | 4.00176e-32 (rank : 12) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P53672, Q9Y562 | Gene names | CRYBA2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin A2 (Beta-A2-crystallin). | |||||
|
CRBS_HUMAN
|
||||||
| NC score | 0.925480 (rank : 13) | θ value | 3.17079e-29 (rank : 13) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P22914 | Gene names | CRYGS, GRYG8 | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin S (Gamma crystallin S). | |||||
|
CRBS_MOUSE
|
||||||
| NC score | 0.924438 (rank : 14) | θ value | 1.2049e-28 (rank : 14) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O35486 | Gene names | Crygs | |||
|
Domain Architecture |

|
|||||
| Description | Beta crystallin S (Gamma crystallin S). | |||||
|
CRGB_MOUSE
|
||||||
| NC score | 0.908760 (rank : 15) | θ value | 1.73987e-27 (rank : 15) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P04344, Q61593 | Gene names | Crygb | |||
|
Domain Architecture |

|
|||||
| Description | Gamma crystallin B (Gamma crystallin 3). | |||||
|
CRGC_HUMAN
|
||||||
| NC score | 0.906971 (rank : 16) | θ value | 6.61148e-27 (rank : 17) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P07315 | Gene names | CRYGC, CRYG3 | |||
|
Domain Architecture |

|
|||||
| Description | Gamma crystallin C (Gamma crystallin 2-1) (Gamma crystallin 3). | |||||
|
CRGA_MOUSE
|
||||||
| NC score | 0.905064 (rank : 17) | θ value | 6.61148e-27 (rank : 16) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P04345 | Gene names | Cryga | |||
|
Domain Architecture |

|
|||||
| Description | Gamma crystallin A (Gamma crystallin 4). | |||||
|
CRGB_HUMAN
|
||||||
| NC score | 0.902027 (rank : 18) | θ value | 9.54697e-26 (rank : 18) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P07316 | Gene names | CRYGB, CRYG2 | |||
|
Domain Architecture |

|
|||||
| Description | Gamma crystallin B (Gamma crystallin 1-2). | |||||
|
CRGA_HUMAN
|
||||||
| NC score | 0.900832 (rank : 19) | θ value | 3.62785e-25 (rank : 19) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P11844 | Gene names | CRYGA, CRYG1 | |||
|
Domain Architecture |

|
|||||
| Description | Gamma crystallin A (Gamma crystallin 5). | |||||
|
CRGC_MOUSE
|
||||||
| NC score | 0.899525 (rank : 20) | θ value | 3.62785e-25 (rank : 20) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q61597, Q03739 | Gene names | Crygc, Gammab-cry | |||
|
Domain Architecture |

|
|||||
| Description | Gamma crystallin C. | |||||
|
CRGD_MOUSE
|
||||||
| NC score | 0.895953 (rank : 21) | θ value | 1.05554e-24 (rank : 21) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P04342, O89027 | Gene names | Crygd | |||
|
Domain Architecture |

|
|||||
| Description | Gamma crystallin D (Gamma crystallin 1). | |||||
|
CRGE_MOUSE
|
||||||
| NC score | 0.895675 (rank : 22) | θ value | 5.23862e-24 (rank : 23) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q03740, O89028, P26999, Q9CXK5 | Gene names | Cryge | |||
|
Domain Architecture |

|
|||||
| Description | Gamma crystallin E. | |||||
|
CRGF_MOUSE
|
||||||
| NC score | 0.894648 (rank : 23) | θ value | 3.07116e-24 (rank : 22) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9CXV3 | Gene names | Crygf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gamma crystallin F. | |||||
|
CRGD_HUMAN
|
||||||
| NC score | 0.892446 (rank : 24) | θ value | 2.59989e-23 (rank : 24) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P07320, Q53R51, Q99681 | Gene names | CRYGD, CRYG4 | |||
|
Domain Architecture |

|
|||||
| Description | Gamma crystallin D (Gamma crystallin 4). | |||||
|
AIM1_HUMAN
|
||||||
| NC score | 0.805081 (rank : 25) | θ value | 1.74391e-19 (rank : 25) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9Y4K1, O00296, Q5VWJ2 | Gene names | AIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Absent in melanoma 1 protein. | |||||
|
TOP2B_HUMAN
|
||||||
| NC score | 0.017390 (rank : 26) | θ value | 1.81305 (rank : 26) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q02880, Q13600, Q9UMG8, Q9UQP8 | Gene names | TOP2B | |||
|
Domain Architecture |

|
|||||
| Description | DNA topoisomerase 2-beta (EC 5.99.1.3) (DNA topoisomerase II, beta isozyme). | |||||
|
TOP2B_MOUSE
|
||||||
| NC score | 0.016142 (rank : 27) | θ value | 4.03905 (rank : 28) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q64511, Q7TQG4 | Gene names | Top2b | |||
|
Domain Architecture |

|
|||||
| Description | DNA topoisomerase 2-beta (EC 5.99.1.3) (DNA topoisomerase II, beta isozyme). | |||||
|
NMNA3_HUMAN
|
||||||
| NC score | 0.009183 (rank : 28) | θ value | 5.27518 (rank : 30) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96T66, Q8N4G1 | Gene names | NMNAT3 | |||
|
Domain Architecture |

|
|||||
| Description | Nicotinamide mononucleotide adenylyltransferase 3 (EC 2.7.7.1) (NMN adenylyltransferase 3). | |||||
|
DOCK4_MOUSE
|
||||||
| NC score | 0.005962 (rank : 29) | θ value | 8.99809 (rank : 32) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P59764 | Gene names | Dock4, Kiaa0716 | |||
|
Domain Architecture |

|
|||||
| Description | Dedicator of cytokinesis protein 4. | |||||
|
DOCK4_HUMAN
|
||||||
| NC score | 0.005930 (rank : 30) | θ value | 8.99809 (rank : 31) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 237 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8N1I0, O14584, O94824, Q8NB45 | Gene names | DOCK4, KIAA0716 | |||
|
Domain Architecture |

|
|||||
| Description | Dedicator of cytokinesis protein 4. | |||||
|
ADA28_HUMAN
|
||||||
| NC score | 0.001725 (rank : 31) | θ value | 5.27518 (rank : 29) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UKQ2, Q9Y339, Q9Y3S0 | Gene names | ADAM28, MDCL | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 28 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 28) (Metalloproteinase-like, disintegrin-like, and cysteine- rich protein-L) (MDC-L) (eMDC II) (ADAM23). | |||||
|
ZKSC1_MOUSE
|
||||||
| NC score | -0.002005 (rank : 32) | θ value | 2.36792 (rank : 27) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 755 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BGS3, Q7TS88, Q8BJ55, Q9CRN6 | Gene names | Zkscan1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger with KRAB and SCAN domain-containing protein 1. | |||||