Please be patient as the page loads
|
NGLY1_HUMAN
|
||||||
| SwissProt Accessions | Q96IV0, Q59FB1, Q6PJD8, Q9BVR8, Q9NR70 | Gene names | NGLY1, PNG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peptide-N(4)-(N-acetyl-beta-glucosaminyl)asparagine amidase (EC 3.5.1.52) (PNGase) (hPNGase) (Peptide:N-glycanase) (N-glycanase 1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
NGLY1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q96IV0, Q59FB1, Q6PJD8, Q9BVR8, Q9NR70 | Gene names | NGLY1, PNG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peptide-N(4)-(N-acetyl-beta-glucosaminyl)asparagine amidase (EC 3.5.1.52) (PNGase) (hPNGase) (Peptide:N-glycanase) (N-glycanase 1). | |||||
|
NGLY1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.990688 (rank : 2) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9JI78, Q8K113, Q9CTK3 | Gene names | Ngly1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peptide-N(4)-(N-acetyl-beta-glucosaminyl)asparagine amidase (EC 3.5.1.52) (PNGase) (mPNGase) (Peptide:N-glycanase) (N-glycanase 1). | |||||
|
ADIP_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 3) | NC score | 0.084828 (rank : 8) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 470 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y2D8, Q6P2P8, Q6ULS1, Q7L168, Q9UIX0 | Gene names | SSX2IP, KIAA0923 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Afadin- and alpha-actinin-binding protein (ADIP) (Afadin DIL domain- interacting protein) (SSX2-interacting protein). | |||||
|
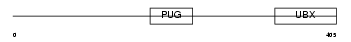
UBXD1_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 4) | NC score | 0.144792 (rank : 3) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BZV1, Q96AH1, Q96IK9, Q9BZV0 | Gene names | UBXD1, UBXDC2 | |||
|
Domain Architecture |
|
|||||
| Description | UBX domain-containing protein 1. | |||||
|
ADIP_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 5) | NC score | 0.085059 (rank : 7) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 415 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8VC66, Q8BG59, Q8C7X0, Q8K2F7 | Gene names | Ssx2ip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Afadin- and alpha-actinin-binding protein (ADIP) (Afadin DIL domain- interacting protein). | |||||
|
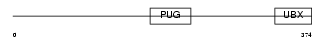
UBXD1_MOUSE
|
||||||
| θ value | 0.125558 (rank : 6) | NC score | 0.134317 (rank : 4) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99PL6, Q91W79 | Gene names | Ubxd1, Ubxdc2 | |||
|
Domain Architecture |
|
|||||
| Description | UBX domain-containing protein 1. | |||||
|
JAD1A_HUMAN
|
||||||
| θ value | 0.163984 (rank : 7) | NC score | 0.031309 (rank : 14) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P29375 | Gene names | JARID1A, RBBP2, RBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1A (Retinoblastoma-binding protein 2) (RBBP-2). | |||||
|
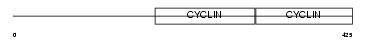
CCNA2_HUMAN
|
||||||
| θ value | 0.279714 (rank : 8) | NC score | 0.032499 (rank : 13) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P20248 | Gene names | CCNA2, CCN1, CCNA | |||
|
Domain Architecture |
|
|||||
| Description | Cyclin-A2 (Cyclin-A). | |||||
|
APC_HUMAN
|
||||||
| θ value | 0.365318 (rank : 9) | NC score | 0.039637 (rank : 11) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 603 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P25054, Q15162, Q15163, Q93042 | Gene names | APC, DP2.5 | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC). | |||||
|
REM1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 10) | NC score | 0.020693 (rank : 22) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O75628, Q9NP57 | Gene names | REM1, GES, REM | |||
|
Domain Architecture |

|
|||||
| Description | GTP-binding protein REM 1 (Rad and Gem-like GTP-binding protein 1) (GTPase-regulating endothelial cell sprouting). | |||||
|
XPC_HUMAN
|
||||||
| θ value | 0.62314 (rank : 11) | NC score | 0.093875 (rank : 6) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q01831, Q96AX0 | Gene names | XPC, XPCC | |||
|
Domain Architecture |

|
|||||
| Description | DNA-repair protein complementing XP-C cells (Xeroderma pigmentosum group C-complementing protein) (p125). | |||||
|
XPC_MOUSE
|
||||||
| θ value | 0.62314 (rank : 12) | NC score | 0.096369 (rank : 5) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P51612, P54732, Q3TKI2, Q920M1, Q9DBW7 | Gene names | Xpc | |||
|
Domain Architecture |

|
|||||
| Description | DNA-repair protein complementing XP-C cells homolog (Xeroderma pigmentosum group C-complementing protein homolog) (p125). | |||||
|
BRD3_HUMAN
|
||||||
| θ value | 1.06291 (rank : 13) | NC score | 0.027591 (rank : 16) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 369 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15059, Q5T1R7, Q8N5M3, Q92645 | Gene names | BRD3, KIAA0043, RING3L | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 3 (RING3-like protein). | |||||
|
REM1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 14) | NC score | 0.019522 (rank : 24) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O35929 | Gene names | Rem1, Rem | |||
|
Domain Architecture |

|
|||||
| Description | GTP-binding protein REM 1 (Rad and Gem-like GTP-binding protein 1). | |||||
|
K0460_HUMAN
|
||||||
| θ value | 1.81305 (rank : 15) | NC score | 0.028738 (rank : 15) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5VT52, O75048, Q5VT53, Q6MZL4, Q86XD2 | Gene names | KIAA0460 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0460. | |||||
|
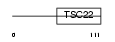
T22D1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 16) | NC score | 0.039733 (rank : 10) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q15714, O00666, Q96JS5 | Gene names | TSC22D1, TGFB1I4, TSC22 | |||
|
Domain Architecture |
|
|||||
| Description | TSC22 domain family protein 1 (Transforming growth factor beta-1- induced transcript 4 protein) (Regulatory protein TSC-22) (TGFB- stimulated clone 22 homolog) (Cerebral protein 2). | |||||
|
EP300_HUMAN
|
||||||
| θ value | 2.36792 (rank : 17) | NC score | 0.021698 (rank : 20) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q09472 | Gene names | EP300, P300 | |||
|
Domain Architecture |

|
|||||
| Description | E1A-associated protein p300 (EC 2.3.1.48). | |||||
|
NU214_HUMAN
|
||||||
| θ value | 3.0926 (rank : 18) | NC score | 0.025894 (rank : 17) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P35658 | Gene names | NUP214, CAIN, CAN | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup214 (Nucleoporin Nup214) (214 kDa nucleoporin) (CAN protein). | |||||
|
PO121_MOUSE
|
||||||
| θ value | 3.0926 (rank : 19) | NC score | 0.022327 (rank : 19) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8K3Z9, Q7TSH5 | Gene names | Pom121, Nup121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear envelope pore membrane protein POM 121 (Pore membrane protein of 121 kDa). | |||||
|
CF061_HUMAN
|
||||||
| θ value | 4.03905 (rank : 20) | NC score | 0.048021 (rank : 9) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NXL9, Q9HCV5 | Gene names | C6orf61 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C6orf61. | |||||
|
PCNT_MOUSE
|
||||||
| θ value | 4.03905 (rank : 21) | NC score | 0.020967 (rank : 21) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 1207 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P48725 | Gene names | Pcnt, Pcnt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pericentrin. | |||||
|
T22D1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 22) | NC score | 0.034039 (rank : 12) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P62500, Q00992 | Gene names | Tsc22d1, Tgfb1i4, Tsc22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TSC22 domain family protein 1 (Transforming growth factor beta-1- induced transcript 4 protein) (Regulatory protein TSC-22) (TGFB- stimulated clone 22 homolog). | |||||
|
TAGAP_HUMAN
|
||||||
| θ value | 4.03905 (rank : 23) | NC score | 0.016378 (rank : 28) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N103, Q2NKM8, Q8NI40, Q96KZ2, Q96QA2 | Gene names | TAGAP, TAGAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | T-cell activation Rho GTPase-activating protein (T-cell activation GTPase-activating protein). | |||||
|
BRD3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 24) | NC score | 0.022520 (rank : 18) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8K2F0, Q8C665, Q8CAX7, Q9JI25 | Gene names | Brd3, Fsrg2 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 3 (Bromodomain-containing FSH-like protein FSRG2). | |||||
|
MINT_HUMAN
|
||||||
| θ value | 5.27518 (rank : 25) | NC score | 0.012041 (rank : 32) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 1291 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q96T58, Q9H9A8, Q9NWH5, Q9UQ01, Q9Y556 | Gene names | SPEN, KIAA0929, MINT, SHARP | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
PHF2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 26) | NC score | 0.018611 (rank : 26) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9WTU0, Q80WA8 | Gene names | Phf2 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
RRMJ3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 27) | NC score | 0.018951 (rank : 25) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9DBE9, Q3ULI1, Q921I7 | Gene names | Ftsj3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative rRNA methyltransferase 3 (EC 2.1.1.-) (rRNA (uridine-2'-O-)- methyltransferase 3). | |||||
|
ZHX1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 28) | NC score | 0.012932 (rank : 31) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 334 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P70121, Q8BQ68, Q8C6T4, Q8CJG3 | Gene names | Zhx1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc fingers and homeoboxes protein 1. | |||||
|
PHF2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 29) | NC score | 0.017741 (rank : 27) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75151, Q8N3K2, Q9Y6N4 | Gene names | PHF2, KIAA0662 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
CUTA_MOUSE
|
||||||
| θ value | 8.99809 (rank : 30) | NC score | 0.020290 (rank : 23) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 3 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9CQ89, Q3TXB6, Q8CI44, Q9D1L4 | Gene names | Cuta | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CutA precursor (Brain acetylcholinesterase putative membrane anchor). | |||||
|
FBX43_MOUSE
|
||||||
| θ value | 8.99809 (rank : 31) | NC score | 0.014715 (rank : 30) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CDI2, Q4G0C8, Q8R0R1 | Gene names | Fbxo43, Emi2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 43 (Endogenous meiotic inhibitor 2). | |||||
|
OPTN_HUMAN
|
||||||
| θ value | 8.99809 (rank : 32) | NC score | 0.005455 (rank : 34) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 827 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96CV9, Q5T672, Q5T673, Q5T674, Q5T675, Q7LDL9, Q8N562, Q9UET9, Q9UEV4, Q9Y218 | Gene names | OPTN, FIP2, GLC1E, NRP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Optineurin (Optic neuropathy-inducing protein) (E3-14.7K-interacting protein) (FIP-2) (Huntingtin-interacting protein HYPL) (NEMO-related protein) (Transcription factor IIIA-interacting protein) (TFIIIA- IntP). | |||||
|
SFR16_HUMAN
|
||||||
| θ value | 8.99809 (rank : 33) | NC score | 0.015285 (rank : 29) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8N2M8, O96026, Q6UW71, Q96DX2 | Gene names | SFRS16, SWAP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 16 (Suppressor of white-apricot homolog 2). | |||||
|
UBQL2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 34) | NC score | 0.008725 (rank : 33) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UHD9, O94798, Q9H3W6, Q9HAZ4 | Gene names | UBQLN2, PLIC2 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquilin-2 (Protein linking IAP with cytoskeleton 2) (PLIC-2) (hPLIC- 2) (Ubiquitin-like product Chap1/Dsk2) (DSK2 homolog) (Chap1). | |||||
|
NGLY1_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q96IV0, Q59FB1, Q6PJD8, Q9BVR8, Q9NR70 | Gene names | NGLY1, PNG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peptide-N(4)-(N-acetyl-beta-glucosaminyl)asparagine amidase (EC 3.5.1.52) (PNGase) (hPNGase) (Peptide:N-glycanase) (N-glycanase 1). | |||||
|
NGLY1_MOUSE
|
||||||
| NC score | 0.990688 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9JI78, Q8K113, Q9CTK3 | Gene names | Ngly1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peptide-N(4)-(N-acetyl-beta-glucosaminyl)asparagine amidase (EC 3.5.1.52) (PNGase) (mPNGase) (Peptide:N-glycanase) (N-glycanase 1). | |||||
|
UBXD1_HUMAN
|
||||||
| NC score | 0.144792 (rank : 3) | θ value | 0.0431538 (rank : 4) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BZV1, Q96AH1, Q96IK9, Q9BZV0 | Gene names | UBXD1, UBXDC2 | |||
|
Domain Architecture |

|
|||||
| Description | UBX domain-containing protein 1. | |||||
|
UBXD1_MOUSE
|
||||||
| NC score | 0.134317 (rank : 4) | θ value | 0.125558 (rank : 6) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99PL6, Q91W79 | Gene names | Ubxd1, Ubxdc2 | |||
|
Domain Architecture |

|
|||||
| Description | UBX domain-containing protein 1. | |||||
|
XPC_MOUSE
|
||||||
| NC score | 0.096369 (rank : 5) | θ value | 0.62314 (rank : 12) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P51612, P54732, Q3TKI2, Q920M1, Q9DBW7 | Gene names | Xpc | |||
|
Domain Architecture |

|
|||||
| Description | DNA-repair protein complementing XP-C cells homolog (Xeroderma pigmentosum group C-complementing protein homolog) (p125). | |||||
|
XPC_HUMAN
|
||||||
| NC score | 0.093875 (rank : 6) | θ value | 0.62314 (rank : 11) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q01831, Q96AX0 | Gene names | XPC, XPCC | |||
|
Domain Architecture |

|
|||||
| Description | DNA-repair protein complementing XP-C cells (Xeroderma pigmentosum group C-complementing protein) (p125). | |||||
|
ADIP_MOUSE
|
||||||
| NC score | 0.085059 (rank : 7) | θ value | 0.0961366 (rank : 5) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 415 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8VC66, Q8BG59, Q8C7X0, Q8K2F7 | Gene names | Ssx2ip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Afadin- and alpha-actinin-binding protein (ADIP) (Afadin DIL domain- interacting protein). | |||||
|
ADIP_HUMAN
|
||||||
| NC score | 0.084828 (rank : 8) | θ value | 0.0330416 (rank : 3) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 470 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y2D8, Q6P2P8, Q6ULS1, Q7L168, Q9UIX0 | Gene names | SSX2IP, KIAA0923 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Afadin- and alpha-actinin-binding protein (ADIP) (Afadin DIL domain- interacting protein) (SSX2-interacting protein). | |||||
|
CF061_HUMAN
|
||||||
| NC score | 0.048021 (rank : 9) | θ value | 4.03905 (rank : 20) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NXL9, Q9HCV5 | Gene names | C6orf61 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C6orf61. | |||||
|
T22D1_HUMAN
|
||||||
| NC score | 0.039733 (rank : 10) | θ value | 1.81305 (rank : 16) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q15714, O00666, Q96JS5 | Gene names | TSC22D1, TGFB1I4, TSC22 | |||
|
Domain Architecture |

|
|||||
| Description | TSC22 domain family protein 1 (Transforming growth factor beta-1- induced transcript 4 protein) (Regulatory protein TSC-22) (TGFB- stimulated clone 22 homolog) (Cerebral protein 2). | |||||
|
APC_HUMAN
|
||||||
| NC score | 0.039637 (rank : 11) | θ value | 0.365318 (rank : 9) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 603 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P25054, Q15162, Q15163, Q93042 | Gene names | APC, DP2.5 | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC). | |||||
|
T22D1_MOUSE
|
||||||
| NC score | 0.034039 (rank : 12) | θ value | 4.03905 (rank : 22) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P62500, Q00992 | Gene names | Tsc22d1, Tgfb1i4, Tsc22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TSC22 domain family protein 1 (Transforming growth factor beta-1- induced transcript 4 protein) (Regulatory protein TSC-22) (TGFB- stimulated clone 22 homolog). | |||||
|
CCNA2_HUMAN
|
||||||
| NC score | 0.032499 (rank : 13) | θ value | 0.279714 (rank : 8) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P20248 | Gene names | CCNA2, CCN1, CCNA | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-A2 (Cyclin-A). | |||||
|
JAD1A_HUMAN
|
||||||
| NC score | 0.031309 (rank : 14) | θ value | 0.163984 (rank : 7) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P29375 | Gene names | JARID1A, RBBP2, RBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1A (Retinoblastoma-binding protein 2) (RBBP-2). | |||||
|
K0460_HUMAN
|
||||||
| NC score | 0.028738 (rank : 15) | θ value | 1.81305 (rank : 15) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5VT52, O75048, Q5VT53, Q6MZL4, Q86XD2 | Gene names | KIAA0460 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0460. | |||||
|
BRD3_HUMAN
|
||||||
| NC score | 0.027591 (rank : 16) | θ value | 1.06291 (rank : 13) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 369 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15059, Q5T1R7, Q8N5M3, Q92645 | Gene names | BRD3, KIAA0043, RING3L | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 3 (RING3-like protein). | |||||
|
NU214_HUMAN
|
||||||
| NC score | 0.025894 (rank : 17) | θ value | 3.0926 (rank : 18) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P35658 | Gene names | NUP214, CAIN, CAN | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup214 (Nucleoporin Nup214) (214 kDa nucleoporin) (CAN protein). | |||||
|
BRD3_MOUSE
|
||||||
| NC score | 0.022520 (rank : 18) | θ value | 5.27518 (rank : 24) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8K2F0, Q8C665, Q8CAX7, Q9JI25 | Gene names | Brd3, Fsrg2 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 3 (Bromodomain-containing FSH-like protein FSRG2). | |||||
|
PO121_MOUSE
|
||||||
| NC score | 0.022327 (rank : 19) | θ value | 3.0926 (rank : 19) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8K3Z9, Q7TSH5 | Gene names | Pom121, Nup121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear envelope pore membrane protein POM 121 (Pore membrane protein of 121 kDa). | |||||
|
EP300_HUMAN
|
||||||
| NC score | 0.021698 (rank : 20) | θ value | 2.36792 (rank : 17) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q09472 | Gene names | EP300, P300 | |||
|
Domain Architecture |

|
|||||
| Description | E1A-associated protein p300 (EC 2.3.1.48). | |||||
|
PCNT_MOUSE
|
||||||
| NC score | 0.020967 (rank : 21) | θ value | 4.03905 (rank : 21) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 1207 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P48725 | Gene names | Pcnt, Pcnt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pericentrin. | |||||
|
REM1_HUMAN
|
||||||
| NC score | 0.020693 (rank : 22) | θ value | 0.47712 (rank : 10) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O75628, Q9NP57 | Gene names | REM1, GES, REM | |||
|
Domain Architecture |

|
|||||
| Description | GTP-binding protein REM 1 (Rad and Gem-like GTP-binding protein 1) (GTPase-regulating endothelial cell sprouting). | |||||
|
CUTA_MOUSE
|
||||||
| NC score | 0.020290 (rank : 23) | θ value | 8.99809 (rank : 30) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 3 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9CQ89, Q3TXB6, Q8CI44, Q9D1L4 | Gene names | Cuta | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CutA precursor (Brain acetylcholinesterase putative membrane anchor). | |||||
|
REM1_MOUSE
|
||||||
| NC score | 0.019522 (rank : 24) | θ value | 1.06291 (rank : 14) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O35929 | Gene names | Rem1, Rem | |||
|
Domain Architecture |

|
|||||
| Description | GTP-binding protein REM 1 (Rad and Gem-like GTP-binding protein 1). | |||||
|
RRMJ3_MOUSE
|
||||||
| NC score | 0.018951 (rank : 25) | θ value | 5.27518 (rank : 27) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9DBE9, Q3ULI1, Q921I7 | Gene names | Ftsj3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative rRNA methyltransferase 3 (EC 2.1.1.-) (rRNA (uridine-2'-O-)- methyltransferase 3). | |||||
|
PHF2_MOUSE
|
||||||
| NC score | 0.018611 (rank : 26) | θ value | 5.27518 (rank : 26) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9WTU0, Q80WA8 | Gene names | Phf2 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
PHF2_HUMAN
|
||||||
| NC score | 0.017741 (rank : 27) | θ value | 6.88961 (rank : 29) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75151, Q8N3K2, Q9Y6N4 | Gene names | PHF2, KIAA0662 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
TAGAP_HUMAN
|
||||||
| NC score | 0.016378 (rank : 28) | θ value | 4.03905 (rank : 23) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N103, Q2NKM8, Q8NI40, Q96KZ2, Q96QA2 | Gene names | TAGAP, TAGAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | T-cell activation Rho GTPase-activating protein (T-cell activation GTPase-activating protein). | |||||
|
SFR16_HUMAN
|
||||||
| NC score | 0.015285 (rank : 29) | θ value | 8.99809 (rank : 33) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8N2M8, O96026, Q6UW71, Q96DX2 | Gene names | SFRS16, SWAP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 16 (Suppressor of white-apricot homolog 2). | |||||
|
FBX43_MOUSE
|
||||||
| NC score | 0.014715 (rank : 30) | θ value | 8.99809 (rank : 31) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CDI2, Q4G0C8, Q8R0R1 | Gene names | Fbxo43, Emi2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 43 (Endogenous meiotic inhibitor 2). | |||||
|
ZHX1_MOUSE
|
||||||
| NC score | 0.012932 (rank : 31) | θ value | 5.27518 (rank : 28) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 334 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P70121, Q8BQ68, Q8C6T4, Q8CJG3 | Gene names | Zhx1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc fingers and homeoboxes protein 1. | |||||
|
MINT_HUMAN
|
||||||
| NC score | 0.012041 (rank : 32) | θ value | 5.27518 (rank : 25) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 1291 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q96T58, Q9H9A8, Q9NWH5, Q9UQ01, Q9Y556 | Gene names | SPEN, KIAA0929, MINT, SHARP | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
UBQL2_HUMAN
|
||||||
| NC score | 0.008725 (rank : 33) | θ value | 8.99809 (rank : 34) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UHD9, O94798, Q9H3W6, Q9HAZ4 | Gene names | UBQLN2, PLIC2 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquilin-2 (Protein linking IAP with cytoskeleton 2) (PLIC-2) (hPLIC- 2) (Ubiquitin-like product Chap1/Dsk2) (DSK2 homolog) (Chap1). | |||||
|
OPTN_HUMAN
|
||||||
| NC score | 0.005455 (rank : 34) | θ value | 8.99809 (rank : 32) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 827 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96CV9, Q5T672, Q5T673, Q5T674, Q5T675, Q7LDL9, Q8N562, Q9UET9, Q9UEV4, Q9Y218 | Gene names | OPTN, FIP2, GLC1E, NRP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Optineurin (Optic neuropathy-inducing protein) (E3-14.7K-interacting protein) (FIP-2) (Huntingtin-interacting protein HYPL) (NEMO-related protein) (Transcription factor IIIA-interacting protein) (TFIIIA- IntP). | |||||