Please be patient as the page loads
|
NDST2_HUMAN
|
||||||
| SwissProt Accessions | P52849, Q2TB32, Q59H89 | Gene names | NDST2, HSST2 | |||
|
Domain Architecture |

|
|||||
| Description | Bifunctional heparan sulfate N-deacetylase/N-sulfotransferase 2 (EC 2.8.2.8) (Glucosaminyl N-deacetylase/N-sulfotransferase 2) (NDST- 2) (N-heparan sulfate sulfotransferase 2) (N-HSST 2) [Includes: Heparan sulfate N-deacetylase 2 (EC 3.-.-.-); Heparan sulfate N- sulfotransferase 2 (EC 2.8.2.-)]. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
NDST1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.997673 (rank : 4) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P52848, Q96E57 | Gene names | NDST1, HSST, HSST1 | |||
|
Domain Architecture |

|
|||||
| Description | Bifunctional heparan sulfate N-deacetylase/N-sulfotransferase 1 (EC 2.8.2.8) (Glucosaminyl N-deacetylase/N-sulfotransferase 1) (NDST- 1) ([Heparan sulfate]-glucosamine N-sulfotransferase 1) (HSNST 1) (N- heparan sulfate sulfotransferase 1) (N-HSST 1) [Includes: Heparan sulfate N-deacetylase 1 (EC 3.-.-.-); Heparan sulfate N- sulfotransferase 1 (EC 2.8.2.-)]. | |||||
|
NDST1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.997781 (rank : 3) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q3UHN9, O70353, Q3TBX3, Q3TDS3, Q8BZE5, Q9R206 | Gene names | Ndst1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bifunctional heparan sulfate N-deacetylase/N-sulfotransferase 1 (EC 2.8.2.8) (Glucosaminyl N-deacetylase/N-sulfotransferase 1) (NDST- 1) ([Heparan sulfate]-glucosamine N-sulfotransferase 1) (HSNST 1) (N- heparan sulfate sulfotransferase 1) (N-HSST 1) [Includes: Heparan sulfate N-deacetylase 1 (EC 3.-.-.-); Heparan sulfate N- sulfotransferase 1 (EC 2.8.2.-)]. | |||||
|
NDST2_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P52849, Q2TB32, Q59H89 | Gene names | NDST2, HSST2 | |||
|
Domain Architecture |

|
|||||
| Description | Bifunctional heparan sulfate N-deacetylase/N-sulfotransferase 2 (EC 2.8.2.8) (Glucosaminyl N-deacetylase/N-sulfotransferase 2) (NDST- 2) (N-heparan sulfate sulfotransferase 2) (N-HSST 2) [Includes: Heparan sulfate N-deacetylase 2 (EC 3.-.-.-); Heparan sulfate N- sulfotransferase 2 (EC 2.8.2.-)]. | |||||
|
NDST2_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.999349 (rank : 2) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P52850, Q3UDF4, Q549P5 | Gene names | Ndst2, Hsst2 | |||
|
Domain Architecture |

|
|||||
| Description | Bifunctional heparan sulfate N-deacetylase/N-sulfotransferase 2 (EC 2.8.2.8) (Glucosaminyl N-deacetylase/N-sulfotransferase 2) (NDST- 2) (N-heparan sulfate sulfotransferase 2) (N-HSST 2) (Mndns) [Includes: Heparan sulfate N-deacetylase 2 (EC 3.-.-.-); Heparan sulfate N-sulfotransferase 2 (EC 2.8.2.-)]. | |||||
|
NDST3_HUMAN
|
||||||
| θ value | 0 (rank : 5) | NC score | 0.997213 (rank : 6) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O95803, Q4W5C1, Q4W5D0, Q6UWC5, Q9UP21 | Gene names | NDST3, HSST3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bifunctional heparan sulfate N-deacetylase/N-sulfotransferase 3 (EC 2.8.2.8) (Glucosaminyl N-deacetylase/N-sulfotransferase 3) (NDST- 3) (hNDST-3) (N-heparan sulfate sulfotransferase 3) (N-HSST 3) [Includes: Heparan sulfate N-deacetylase 3 (EC 3.-.-.-); Heparan sulfate N-sulfotransferase 3 (EC 2.8.2.-)]. | |||||
|
NDST3_MOUSE
|
||||||
| θ value | 0 (rank : 6) | NC score | 0.997382 (rank : 5) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9EQH7, Q6AXE0, Q9D557 | Gene names | Ndst3, Hsst3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bifunctional heparan sulfate N-deacetylase/N-sulfotransferase 3 (EC 2.8.2.8) (Glucosaminyl N-deacetylase/N-sulfotransferase 3) (NDST- 3) (N-heparan sulfate sulfotransferase 3) (N-HSST 3) [Includes: Heparan sulfate N-deacetylase 3 (EC 3.-.-.-); Heparan sulfate N- sulfotransferase 3 (EC 2.8.2.-)]. | |||||
|
NDST4_HUMAN
|
||||||
| θ value | 0 (rank : 7) | NC score | 0.997156 (rank : 7) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9H3R1 | Gene names | NDST4, HSST4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bifunctional heparan sulfate N-deacetylase/N-sulfotransferase 4 (EC 2.8.2.8) (Glucosaminyl N-deacetylase/N-sulfotransferase 4) (NDST- 4) (N-heparan sulfate sulfotransferase 4) (N-HSST 4) [Includes: Heparan sulfate N-deacetylase 4 (EC 3.-.-.-); Heparan sulfate N- sulfotransferase 4 (EC 2.8.2.-)]. | |||||
|
NDST4_MOUSE
|
||||||
| θ value | 0 (rank : 8) | NC score | 0.997075 (rank : 8) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9EQW8 | Gene names | Ndst4, Hsst4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bifunctional heparan sulfate N-deacetylase/N-sulfotransferase 4 (EC 2.8.2.8) (Glucosaminyl N-deacetylase/N-sulfotransferase 4) (NDST- 4) (N-heparan sulfate sulfotransferase 4) (N-HSST 4) [Includes: Heparan sulfate N-deacetylase 4 (EC 3.-.-.-); Heparan sulfate N- sulfotransferase 4 (EC 2.8.2.-)]. | |||||
|
OST2_MOUSE
|
||||||
| θ value | 8.3442e-30 (rank : 9) | NC score | 0.763120 (rank : 13) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q673U1, Q8BLP1, Q8C055 | Gene names | Hs3st2, 3ost2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heparan sulfate glucosamine 3-O-sulfotransferase 2 (EC 2.8.2.29) (Heparan sulfate D-glucosaminyl 3-O-sulfotransferase 2) (Heparan sulfate 3-O-sulfotransferase 2). | |||||
|
OST5_MOUSE
|
||||||
| θ value | 1.4233e-29 (rank : 10) | NC score | 0.788558 (rank : 9) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BSL4 | Gene names | Hs3st5, Hs3ost5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heparan sulfate glucosamine 3-O-sulfotransferase 5 (EC 2.8.2.23) (Heparan sulfate D-glucosaminyl 3-O-sulfotransferase 5) (Heparan sulfate 3-O-sulfotransferase 5). | |||||
|
OST2_HUMAN
|
||||||
| θ value | 1.85889e-29 (rank : 11) | NC score | 0.762825 (rank : 14) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9Y278 | Gene names | HS3ST2, 3OST2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heparan sulfate glucosamine 3-O-sulfotransferase 2 (EC 2.8.2.29) (Heparan sulfate D-glucosaminyl 3-O-sulfotransferase 2) (Heparan sulfate 3-O-sulfotransferase 2) (h3-OST-2). | |||||
|
OST5_HUMAN
|
||||||
| θ value | 1.85889e-29 (rank : 12) | NC score | 0.788376 (rank : 11) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8IZT8, Q52LI2, Q8N285 | Gene names | HS3ST5, 3OST5, HS3OST5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heparan sulfate glucosamine 3-O-sulfotransferase 5 (EC 2.8.2.23) (Heparan sulfate D-glucosaminyl 3-O-sulfotransferase 5) (Heparan sulfate 3-O-sulfotransferase 5) (h3-OST-5). | |||||
|
OST3A_MOUSE
|
||||||
| θ value | 5.40856e-29 (rank : 13) | NC score | 0.760856 (rank : 15) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BKN6 | Gene names | Hs3st3a1, 3ost3a1, Hs3st3a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heparan sulfate glucosamine 3-O-sulfotransferase 3A1 (EC 2.8.2.23) (Heparan sulfate D-glucosaminyl 3-O-sulfotransferase 3A1) (Heparan sulfate 3-O-sulfotransferase 3A1). | |||||
|
OST3B_MOUSE
|
||||||
| θ value | 5.40856e-29 (rank : 14) | NC score | 0.758683 (rank : 17) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9QZS6 | Gene names | Hs3st3b1, 3ost3b1, Hs3st3b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heparan sulfate glucosamine 3-O-sulfotransferase 3B1 (EC 2.8.2.30) (Heparan sulfate D-glucosaminyl 3-O-sulfotransferase 3B1) (Heparan sulfate 3-O-sulfotransferase 3B1) (m3-OST-3B). | |||||
|
OST4_HUMAN
|
||||||
| θ value | 4.57862e-28 (rank : 15) | NC score | 0.758491 (rank : 18) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Y661, Q5QI42, Q8NDC2 | Gene names | HS3ST4, 3OST4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heparan sulfate glucosamine 3-O-sulfotransferase 4 (EC 2.8.2.23) (Heparan sulfate D-glucosaminyl 3-O-sulfotransferase 4) (Heparan sulfate 3-O-sulfotransferase 4) (h3-OST-4). | |||||
|
OST1_MOUSE
|
||||||
| θ value | 5.97985e-28 (rank : 16) | NC score | 0.786777 (rank : 12) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O35310 | Gene names | Hs3st1, 3ost, 3ost1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heparan sulfate glucosamine 3-O-sulfotransferase 1 precursor (EC 2.8.2.23) (Heparan sulfate D-glucosaminyl 3-O-sulfotransferase 1) (Heparan sulfate 3-O-sulfotransferase 1). | |||||
|
OST3B_HUMAN
|
||||||
| θ value | 5.97985e-28 (rank : 17) | NC score | 0.757022 (rank : 20) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Y662 | Gene names | HS3ST3B1, 3OST3B1, HS3ST3B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heparan sulfate glucosamine 3-O-sulfotransferase 3B1 (EC 2.8.2.30) (Heparan sulfate D-glucosaminyl 3-O-sulfotransferase 3B1) (Heparan sulfate 3-O-sulfotransferase 3B1) (h3-OST-3B). | |||||
|
OST3A_HUMAN
|
||||||
| θ value | 1.33217e-27 (rank : 18) | NC score | 0.759137 (rank : 16) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Y663 | Gene names | HS3ST3A1, 3OST3A1, HS3ST3A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heparan sulfate glucosamine 3-O-sulfotransferase 3A1 (EC 2.8.2.30) (Heparan sulfate D-glucosaminyl 3-O-sulfotransferase 3A1) (Heparan sulfate 3-O-sulfotransferase 3A1) (h3-OST-3A). | |||||
|
OST1_HUMAN
|
||||||
| θ value | 3.87602e-27 (rank : 19) | NC score | 0.788538 (rank : 10) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O14792, Q6PEY8 | Gene names | HS3ST1, 3OST, 3OST1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heparan sulfate glucosamine 3-O-sulfotransferase 1 precursor (EC 2.8.2.23) (Heparan sulfate D-glucosaminyl 3-O-sulfotransferase 1) (Heparan sulfate 3-O-sulfotransferase 1) (h3-OST-1). | |||||
|
OST6_HUMAN
|
||||||
| θ value | 1.47289e-26 (rank : 20) | NC score | 0.757338 (rank : 19) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q96QI5, Q96RX7 | Gene names | HS3ST6, HS3ST5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heparan sulfate glucosamine 3-O-sulfotransferase 6 (EC 2.8.2.23) (Heparan sulfate D-glucosaminyl 3-O-sulfotransferase 6) (Heparan sulfate 3-O-sulfotransferase 6) (h3-OST-6). | |||||
|
OST6_MOUSE
|
||||||
| θ value | 1.24688e-25 (rank : 21) | NC score | 0.754637 (rank : 21) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q5GFD5 | Gene names | Hs3st6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heparan sulfate glucosamine 3-O-sulfotransferase 6 (EC 2.8.2.23) (Heparan sulfate D-glucosaminyl 3-O-sulfotransferase 6) (Heparan sulfate 3-O-sulfotransferase 6) (3-OST-6). | |||||
|
ST4S6_MOUSE
|
||||||
| θ value | 0.00020696 (rank : 22) | NC score | 0.420720 (rank : 22) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q91XQ5, Q80TW4, Q8BLQ5, Q9D2N6 | Gene names | Galnac4s6st, Brag, Kiaa0598 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylgalactosamine 4-sulfate 6-O-sulfotransferase (EC 2.8.2.33) (GalNAc4S-6ST) (B-cell RAG-associated gene protein). | |||||
|
ST4S6_HUMAN
|
||||||
| θ value | 0.000270298 (rank : 23) | NC score | 0.410989 (rank : 23) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q7LFX5, O60338, O60474, Q86VM4 | Gene names | GALNAC4S6ST, BRAG, KIAA0598 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylgalactosamine 4-sulfate 6-O-sulfotransferase (EC 2.8.2.33) (GalNAc4S-6ST) (B-cell RAG-associated gene protein) (hBRAG). | |||||
|
ST1C3_HUMAN
|
||||||
| θ value | 0.125558 (rank : 24) | NC score | 0.050943 (rank : 24) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6IMI6, Q6IMI5 | Gene names | SULT1C3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative sulfotransferase 1C3 (EC 2.8.2.-). | |||||
|
CIC_HUMAN
|
||||||
| θ value | 0.279714 (rank : 25) | NC score | 0.017696 (rank : 28) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 992 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96RK0, Q7LGI1, Q9UEG5, Q9Y6T1 | Gene names | CIC, KIAA0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
ST1A3_HUMAN
|
||||||
| θ value | 0.813845 (rank : 26) | NC score | 0.042537 (rank : 25) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P50224, O95603, Q6ZWJ5 | Gene names | SULT1A3, STM | |||
|
Domain Architecture |

|
|||||
| Description | Monoamine-sulfating phenol sulfotransferase (EC 2.8.2.1) (Aryl sulfotransferase 1A3) (Sulfotransferase, monoamine-preferring) (M-PST) (Thermolabile phenol sulfotransferase) (TL-PST) (Placental estrogen sulfotransferase) (Catecholamine-sulfating phenol sulfotransferase) (HAST3). | |||||
|
RGS3_MOUSE
|
||||||
| θ value | 1.38821 (rank : 27) | NC score | 0.004227 (rank : 31) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
TECT1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 28) | NC score | 0.020708 (rank : 27) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BZ64, Q3UMP0, Q8BUE2 | Gene names | Tect1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tectonic-1 precursor. | |||||
|
NOTC2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 29) | NC score | -0.000798 (rank : 36) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 1069 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O35516, Q06008, Q60941 | Gene names | Notch2 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 2 precursor (Notch 2) (Motch B) [Contains: Notch 2 extracellular truncation; Notch 2 intracellular domain]. | |||||
|
PTPRU_HUMAN
|
||||||
| θ value | 4.03905 (rank : 30) | NC score | 0.001853 (rank : 34) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 237 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q92729 | Gene names | PTPRU, PCP2 | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase U precursor (EC 3.1.3.48) (R-PTP-U) (Protein-tyrosine phosphatase J) (PTP-J) (Pancreatic carcinoma phosphatase 2) (PCP-2). | |||||
|
CBPZ_HUMAN
|
||||||
| θ value | 5.27518 (rank : 31) | NC score | 0.003659 (rank : 32) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q66K79, O00520, Q96MX2 | Gene names | CPZ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxypeptidase Z precursor (EC 3.4.17.-) (CPZ). | |||||
|
ST4A1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 32) | NC score | 0.033719 (rank : 26) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BR01, O43728 | Gene names | SULT4A1, SULTX3 | |||
|
Domain Architecture |

|
|||||
| Description | Sulfotransferase 4A1 (EC 2.8.2.-) (Brain sulfotransferase-like protein) (hBR-STL) (hBR-STL-1) (Nervous system sulfotransferase) (NST). | |||||
|
ST4A1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 33) | NC score | 0.016210 (rank : 29) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P63046, O88872, Q3TXY5, Q91XS5, Q9CWY7, Q9DC97 | Gene names | Sult4a1, Sultx3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sulfotransferase 4A1 (EC 2.8.2.-) (Brain sulfotransferase-like protein) (mBR-STL) (Nervous system sulfotransferase) (NST). | |||||
|
GPDM_MOUSE
|
||||||
| θ value | 6.88961 (rank : 34) | NC score | 0.006455 (rank : 30) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q64521, Q61507 | Gene names | Gpd2, Gdm1 | |||
|
Domain Architecture |

|
|||||
| Description | Glycerol-3-phosphate dehydrogenase, mitochondrial precursor (EC 1.1.99.5) (GPD-M) (GPDH-M). | |||||
|
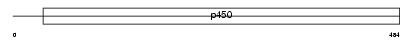
CP2CT_MOUSE
|
||||||
| θ value | 8.99809 (rank : 35) | NC score | 0.000258 (rank : 35) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q64458, Q8WUN8 | Gene names | Cyp2c29 | |||
|
Domain Architecture |
|
|||||
| Description | Cytochrome P450 2C29 (EC 1.14.14.1) (CYPIIC29) (P-450 MUT-2) (Aldehyde oxygenase). | |||||
|
SALL3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 36) | NC score | -0.002447 (rank : 37) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 1066 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BXA9, Q9UGH1 | Gene names | SALL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sal-like protein 3 (Zinc finger protein SALL3) (hSALL3). | |||||
|
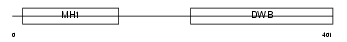
SMAD9_MOUSE
|
||||||
| θ value | 8.99809 (rank : 37) | NC score | 0.002525 (rank : 33) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JIW5, Q8CFF9 | Gene names | Smad9, Madh8, Madh9, Smad8 | |||
|
Domain Architecture |
|
|||||
| Description | Mothers against decapentaplegic homolog 9 (SMAD 9) (Mothers against DPP homolog 9) (Smad9) (Smad8). | |||||
|
NDST2_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P52849, Q2TB32, Q59H89 | Gene names | NDST2, HSST2 | |||
|
Domain Architecture |

|
|||||
| Description | Bifunctional heparan sulfate N-deacetylase/N-sulfotransferase 2 (EC 2.8.2.8) (Glucosaminyl N-deacetylase/N-sulfotransferase 2) (NDST- 2) (N-heparan sulfate sulfotransferase 2) (N-HSST 2) [Includes: Heparan sulfate N-deacetylase 2 (EC 3.-.-.-); Heparan sulfate N- sulfotransferase 2 (EC 2.8.2.-)]. | |||||
|
NDST2_MOUSE
|
||||||
| NC score | 0.999349 (rank : 2) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P52850, Q3UDF4, Q549P5 | Gene names | Ndst2, Hsst2 | |||
|
Domain Architecture |

|
|||||
| Description | Bifunctional heparan sulfate N-deacetylase/N-sulfotransferase 2 (EC 2.8.2.8) (Glucosaminyl N-deacetylase/N-sulfotransferase 2) (NDST- 2) (N-heparan sulfate sulfotransferase 2) (N-HSST 2) (Mndns) [Includes: Heparan sulfate N-deacetylase 2 (EC 3.-.-.-); Heparan sulfate N-sulfotransferase 2 (EC 2.8.2.-)]. | |||||
|
NDST1_MOUSE
|
||||||
| NC score | 0.997781 (rank : 3) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q3UHN9, O70353, Q3TBX3, Q3TDS3, Q8BZE5, Q9R206 | Gene names | Ndst1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bifunctional heparan sulfate N-deacetylase/N-sulfotransferase 1 (EC 2.8.2.8) (Glucosaminyl N-deacetylase/N-sulfotransferase 1) (NDST- 1) ([Heparan sulfate]-glucosamine N-sulfotransferase 1) (HSNST 1) (N- heparan sulfate sulfotransferase 1) (N-HSST 1) [Includes: Heparan sulfate N-deacetylase 1 (EC 3.-.-.-); Heparan sulfate N- sulfotransferase 1 (EC 2.8.2.-)]. | |||||
|
NDST1_HUMAN
|
||||||
| NC score | 0.997673 (rank : 4) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P52848, Q96E57 | Gene names | NDST1, HSST, HSST1 | |||
|
Domain Architecture |

|
|||||
| Description | Bifunctional heparan sulfate N-deacetylase/N-sulfotransferase 1 (EC 2.8.2.8) (Glucosaminyl N-deacetylase/N-sulfotransferase 1) (NDST- 1) ([Heparan sulfate]-glucosamine N-sulfotransferase 1) (HSNST 1) (N- heparan sulfate sulfotransferase 1) (N-HSST 1) [Includes: Heparan sulfate N-deacetylase 1 (EC 3.-.-.-); Heparan sulfate N- sulfotransferase 1 (EC 2.8.2.-)]. | |||||
|
NDST3_MOUSE
|
||||||
| NC score | 0.997382 (rank : 5) | θ value | 0 (rank : 6) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9EQH7, Q6AXE0, Q9D557 | Gene names | Ndst3, Hsst3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bifunctional heparan sulfate N-deacetylase/N-sulfotransferase 3 (EC 2.8.2.8) (Glucosaminyl N-deacetylase/N-sulfotransferase 3) (NDST- 3) (N-heparan sulfate sulfotransferase 3) (N-HSST 3) [Includes: Heparan sulfate N-deacetylase 3 (EC 3.-.-.-); Heparan sulfate N- sulfotransferase 3 (EC 2.8.2.-)]. | |||||
|
NDST3_HUMAN
|
||||||
| NC score | 0.997213 (rank : 6) | θ value | 0 (rank : 5) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O95803, Q4W5C1, Q4W5D0, Q6UWC5, Q9UP21 | Gene names | NDST3, HSST3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bifunctional heparan sulfate N-deacetylase/N-sulfotransferase 3 (EC 2.8.2.8) (Glucosaminyl N-deacetylase/N-sulfotransferase 3) (NDST- 3) (hNDST-3) (N-heparan sulfate sulfotransferase 3) (N-HSST 3) [Includes: Heparan sulfate N-deacetylase 3 (EC 3.-.-.-); Heparan sulfate N-sulfotransferase 3 (EC 2.8.2.-)]. | |||||
|
NDST4_HUMAN
|
||||||
| NC score | 0.997156 (rank : 7) | θ value | 0 (rank : 7) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9H3R1 | Gene names | NDST4, HSST4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bifunctional heparan sulfate N-deacetylase/N-sulfotransferase 4 (EC 2.8.2.8) (Glucosaminyl N-deacetylase/N-sulfotransferase 4) (NDST- 4) (N-heparan sulfate sulfotransferase 4) (N-HSST 4) [Includes: Heparan sulfate N-deacetylase 4 (EC 3.-.-.-); Heparan sulfate N- sulfotransferase 4 (EC 2.8.2.-)]. | |||||
|
NDST4_MOUSE
|
||||||
| NC score | 0.997075 (rank : 8) | θ value | 0 (rank : 8) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9EQW8 | Gene names | Ndst4, Hsst4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bifunctional heparan sulfate N-deacetylase/N-sulfotransferase 4 (EC 2.8.2.8) (Glucosaminyl N-deacetylase/N-sulfotransferase 4) (NDST- 4) (N-heparan sulfate sulfotransferase 4) (N-HSST 4) [Includes: Heparan sulfate N-deacetylase 4 (EC 3.-.-.-); Heparan sulfate N- sulfotransferase 4 (EC 2.8.2.-)]. | |||||
|
OST5_MOUSE
|
||||||
| NC score | 0.788558 (rank : 9) | θ value | 1.4233e-29 (rank : 10) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BSL4 | Gene names | Hs3st5, Hs3ost5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heparan sulfate glucosamine 3-O-sulfotransferase 5 (EC 2.8.2.23) (Heparan sulfate D-glucosaminyl 3-O-sulfotransferase 5) (Heparan sulfate 3-O-sulfotransferase 5). | |||||
|
OST1_HUMAN
|
||||||
| NC score | 0.788538 (rank : 10) | θ value | 3.87602e-27 (rank : 19) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O14792, Q6PEY8 | Gene names | HS3ST1, 3OST, 3OST1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heparan sulfate glucosamine 3-O-sulfotransferase 1 precursor (EC 2.8.2.23) (Heparan sulfate D-glucosaminyl 3-O-sulfotransferase 1) (Heparan sulfate 3-O-sulfotransferase 1) (h3-OST-1). | |||||
|
OST5_HUMAN
|
||||||
| NC score | 0.788376 (rank : 11) | θ value | 1.85889e-29 (rank : 12) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8IZT8, Q52LI2, Q8N285 | Gene names | HS3ST5, 3OST5, HS3OST5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heparan sulfate glucosamine 3-O-sulfotransferase 5 (EC 2.8.2.23) (Heparan sulfate D-glucosaminyl 3-O-sulfotransferase 5) (Heparan sulfate 3-O-sulfotransferase 5) (h3-OST-5). | |||||
|
OST1_MOUSE
|
||||||
| NC score | 0.786777 (rank : 12) | θ value | 5.97985e-28 (rank : 16) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O35310 | Gene names | Hs3st1, 3ost, 3ost1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heparan sulfate glucosamine 3-O-sulfotransferase 1 precursor (EC 2.8.2.23) (Heparan sulfate D-glucosaminyl 3-O-sulfotransferase 1) (Heparan sulfate 3-O-sulfotransferase 1). | |||||
|
OST2_MOUSE
|
||||||
| NC score | 0.763120 (rank : 13) | θ value | 8.3442e-30 (rank : 9) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q673U1, Q8BLP1, Q8C055 | Gene names | Hs3st2, 3ost2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heparan sulfate glucosamine 3-O-sulfotransferase 2 (EC 2.8.2.29) (Heparan sulfate D-glucosaminyl 3-O-sulfotransferase 2) (Heparan sulfate 3-O-sulfotransferase 2). | |||||
|
OST2_HUMAN
|
||||||
| NC score | 0.762825 (rank : 14) | θ value | 1.85889e-29 (rank : 11) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9Y278 | Gene names | HS3ST2, 3OST2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heparan sulfate glucosamine 3-O-sulfotransferase 2 (EC 2.8.2.29) (Heparan sulfate D-glucosaminyl 3-O-sulfotransferase 2) (Heparan sulfate 3-O-sulfotransferase 2) (h3-OST-2). | |||||
|
OST3A_MOUSE
|
||||||
| NC score | 0.760856 (rank : 15) | θ value | 5.40856e-29 (rank : 13) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BKN6 | Gene names | Hs3st3a1, 3ost3a1, Hs3st3a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heparan sulfate glucosamine 3-O-sulfotransferase 3A1 (EC 2.8.2.23) (Heparan sulfate D-glucosaminyl 3-O-sulfotransferase 3A1) (Heparan sulfate 3-O-sulfotransferase 3A1). | |||||
|
OST3A_HUMAN
|
||||||
| NC score | 0.759137 (rank : 16) | θ value | 1.33217e-27 (rank : 18) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Y663 | Gene names | HS3ST3A1, 3OST3A1, HS3ST3A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heparan sulfate glucosamine 3-O-sulfotransferase 3A1 (EC 2.8.2.30) (Heparan sulfate D-glucosaminyl 3-O-sulfotransferase 3A1) (Heparan sulfate 3-O-sulfotransferase 3A1) (h3-OST-3A). | |||||
|
OST3B_MOUSE
|
||||||
| NC score | 0.758683 (rank : 17) | θ value | 5.40856e-29 (rank : 14) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9QZS6 | Gene names | Hs3st3b1, 3ost3b1, Hs3st3b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heparan sulfate glucosamine 3-O-sulfotransferase 3B1 (EC 2.8.2.30) (Heparan sulfate D-glucosaminyl 3-O-sulfotransferase 3B1) (Heparan sulfate 3-O-sulfotransferase 3B1) (m3-OST-3B). | |||||
|
OST4_HUMAN
|
||||||
| NC score | 0.758491 (rank : 18) | θ value | 4.57862e-28 (rank : 15) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Y661, Q5QI42, Q8NDC2 | Gene names | HS3ST4, 3OST4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heparan sulfate glucosamine 3-O-sulfotransferase 4 (EC 2.8.2.23) (Heparan sulfate D-glucosaminyl 3-O-sulfotransferase 4) (Heparan sulfate 3-O-sulfotransferase 4) (h3-OST-4). | |||||
|
OST6_HUMAN
|
||||||
| NC score | 0.757338 (rank : 19) | θ value | 1.47289e-26 (rank : 20) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q96QI5, Q96RX7 | Gene names | HS3ST6, HS3ST5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heparan sulfate glucosamine 3-O-sulfotransferase 6 (EC 2.8.2.23) (Heparan sulfate D-glucosaminyl 3-O-sulfotransferase 6) (Heparan sulfate 3-O-sulfotransferase 6) (h3-OST-6). | |||||
|
OST3B_HUMAN
|
||||||
| NC score | 0.757022 (rank : 20) | θ value | 5.97985e-28 (rank : 17) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Y662 | Gene names | HS3ST3B1, 3OST3B1, HS3ST3B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heparan sulfate glucosamine 3-O-sulfotransferase 3B1 (EC 2.8.2.30) (Heparan sulfate D-glucosaminyl 3-O-sulfotransferase 3B1) (Heparan sulfate 3-O-sulfotransferase 3B1) (h3-OST-3B). | |||||
|
OST6_MOUSE
|
||||||
| NC score | 0.754637 (rank : 21) | θ value | 1.24688e-25 (rank : 21) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q5GFD5 | Gene names | Hs3st6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heparan sulfate glucosamine 3-O-sulfotransferase 6 (EC 2.8.2.23) (Heparan sulfate D-glucosaminyl 3-O-sulfotransferase 6) (Heparan sulfate 3-O-sulfotransferase 6) (3-OST-6). | |||||
|
ST4S6_MOUSE
|
||||||
| NC score | 0.420720 (rank : 22) | θ value | 0.00020696 (rank : 22) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q91XQ5, Q80TW4, Q8BLQ5, Q9D2N6 | Gene names | Galnac4s6st, Brag, Kiaa0598 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylgalactosamine 4-sulfate 6-O-sulfotransferase (EC 2.8.2.33) (GalNAc4S-6ST) (B-cell RAG-associated gene protein). | |||||
|
ST4S6_HUMAN
|
||||||
| NC score | 0.410989 (rank : 23) | θ value | 0.000270298 (rank : 23) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q7LFX5, O60338, O60474, Q86VM4 | Gene names | GALNAC4S6ST, BRAG, KIAA0598 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylgalactosamine 4-sulfate 6-O-sulfotransferase (EC 2.8.2.33) (GalNAc4S-6ST) (B-cell RAG-associated gene protein) (hBRAG). | |||||
|
ST1C3_HUMAN
|
||||||
| NC score | 0.050943 (rank : 24) | θ value | 0.125558 (rank : 24) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6IMI6, Q6IMI5 | Gene names | SULT1C3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative sulfotransferase 1C3 (EC 2.8.2.-). | |||||
|
ST1A3_HUMAN
|
||||||
| NC score | 0.042537 (rank : 25) | θ value | 0.813845 (rank : 26) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P50224, O95603, Q6ZWJ5 | Gene names | SULT1A3, STM | |||
|
Domain Architecture |

|
|||||
| Description | Monoamine-sulfating phenol sulfotransferase (EC 2.8.2.1) (Aryl sulfotransferase 1A3) (Sulfotransferase, monoamine-preferring) (M-PST) (Thermolabile phenol sulfotransferase) (TL-PST) (Placental estrogen sulfotransferase) (Catecholamine-sulfating phenol sulfotransferase) (HAST3). | |||||
|
ST4A1_HUMAN
|
||||||
| NC score | 0.033719 (rank : 26) | θ value | 5.27518 (rank : 32) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BR01, O43728 | Gene names | SULT4A1, SULTX3 | |||
|
Domain Architecture |

|
|||||
| Description | Sulfotransferase 4A1 (EC 2.8.2.-) (Brain sulfotransferase-like protein) (hBR-STL) (hBR-STL-1) (Nervous system sulfotransferase) (NST). | |||||
|
TECT1_MOUSE
|
||||||
| NC score | 0.020708 (rank : 27) | θ value | 2.36792 (rank : 28) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BZ64, Q3UMP0, Q8BUE2 | Gene names | Tect1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tectonic-1 precursor. | |||||
|
CIC_HUMAN
|
||||||
| NC score | 0.017696 (rank : 28) | θ value | 0.279714 (rank : 25) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 992 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96RK0, Q7LGI1, Q9UEG5, Q9Y6T1 | Gene names | CIC, KIAA0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
ST4A1_MOUSE
|
||||||
| NC score | 0.016210 (rank : 29) | θ value | 5.27518 (rank : 33) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P63046, O88872, Q3TXY5, Q91XS5, Q9CWY7, Q9DC97 | Gene names | Sult4a1, Sultx3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sulfotransferase 4A1 (EC 2.8.2.-) (Brain sulfotransferase-like protein) (mBR-STL) (Nervous system sulfotransferase) (NST). | |||||
|
GPDM_MOUSE
|
||||||
| NC score | 0.006455 (rank : 30) | θ value | 6.88961 (rank : 34) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q64521, Q61507 | Gene names | Gpd2, Gdm1 | |||
|
Domain Architecture |

|
|||||
| Description | Glycerol-3-phosphate dehydrogenase, mitochondrial precursor (EC 1.1.99.5) (GPD-M) (GPDH-M). | |||||
|
RGS3_MOUSE
|
||||||
| NC score | 0.004227 (rank : 31) | θ value | 1.38821 (rank : 27) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
CBPZ_HUMAN
|
||||||
| NC score | 0.003659 (rank : 32) | θ value | 5.27518 (rank : 31) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q66K79, O00520, Q96MX2 | Gene names | CPZ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Carboxypeptidase Z precursor (EC 3.4.17.-) (CPZ). | |||||
|
SMAD9_MOUSE
|
||||||
| NC score | 0.002525 (rank : 33) | θ value | 8.99809 (rank : 37) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JIW5, Q8CFF9 | Gene names | Smad9, Madh8, Madh9, Smad8 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 9 (SMAD 9) (Mothers against DPP homolog 9) (Smad9) (Smad8). | |||||
|
PTPRU_HUMAN
|
||||||
| NC score | 0.001853 (rank : 34) | θ value | 4.03905 (rank : 30) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 237 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q92729 | Gene names | PTPRU, PCP2 | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase U precursor (EC 3.1.3.48) (R-PTP-U) (Protein-tyrosine phosphatase J) (PTP-J) (Pancreatic carcinoma phosphatase 2) (PCP-2). | |||||
|
CP2CT_MOUSE
|
||||||
| NC score | 0.000258 (rank : 35) | θ value | 8.99809 (rank : 35) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q64458, Q8WUN8 | Gene names | Cyp2c29 | |||
|
Domain Architecture |

|
|||||
| Description | Cytochrome P450 2C29 (EC 1.14.14.1) (CYPIIC29) (P-450 MUT-2) (Aldehyde oxygenase). | |||||
|
NOTC2_MOUSE
|
||||||
| NC score | -0.000798 (rank : 36) | θ value | 3.0926 (rank : 29) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 1069 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O35516, Q06008, Q60941 | Gene names | Notch2 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 2 precursor (Notch 2) (Motch B) [Contains: Notch 2 extracellular truncation; Notch 2 intracellular domain]. | |||||
|
SALL3_HUMAN
|
||||||
| NC score | -0.002447 (rank : 37) | θ value | 8.99809 (rank : 36) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 1066 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BXA9, Q9UGH1 | Gene names | SALL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sal-like protein 3 (Zinc finger protein SALL3) (hSALL3). | |||||