Please be patient as the page loads
|
MYBA_MOUSE
|
||||||
| SwissProt Accessions | P51960 | Gene names | Mybl1, Amyb | |||
|
Domain Architecture |
|
|||||
| Description | Myb-related protein A (A-Myb). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
MYBA_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.997161 (rank : 2) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P10243 | Gene names | MYBL1, AMYB | |||
|
Domain Architecture |
|
|||||
| Description | Myb-related protein A (A-Myb). | |||||
|
MYBA_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P51960 | Gene names | Mybl1, Amyb | |||
|
Domain Architecture |

|
|||||
| Description | Myb-related protein A (A-Myb). | |||||
|
MYBB_MOUSE
|
||||||
| θ value | 1.93603e-119 (rank : 3) | NC score | 0.939273 (rank : 5) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P48972 | Gene names | Mybl2, Bmyb | |||
|
Domain Architecture |

|
|||||
| Description | Myb-related protein B (B-Myb). | |||||
|
MYB_HUMAN
|
||||||
| θ value | 9.035e-101 (rank : 4) | NC score | 0.939843 (rank : 3) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P10242, P78391, P78392, P78525, P78526, Q14023, Q14024, Q9UE83 | Gene names | MYB | |||
|
Domain Architecture |
|
|||||
| Description | Myb proto-oncogene protein (C-myb). | |||||
|
MYB_MOUSE
|
||||||
| θ value | 8.4565e-99 (rank : 5) | NC score | 0.939399 (rank : 4) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P06876, Q61929 | Gene names | Myb | |||
|
Domain Architecture |

|
|||||
| Description | Myb proto-oncogene protein (C-myb). | |||||
|
MYBB_HUMAN
|
||||||
| θ value | 7.44272e-79 (rank : 6) | NC score | 0.916712 (rank : 6) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P10244 | Gene names | MYBL2, BMYB | |||
|
Domain Architecture |

|
|||||
| Description | Myb-related protein B (B-Myb). | |||||
|
SNPC4_HUMAN
|
||||||
| θ value | 7.0802e-21 (rank : 7) | NC score | 0.544948 (rank : 8) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q5SXM2, Q9Y6P7 | Gene names | SNAPC4, SNAP190 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | snRNA-activating protein complex subunit 4 (SNAPc subunit 4) (snRNA- activating protein complex 190 kDa subunit) (SNAPc 190 kDa subunit) (Proximal sequence element-binding transcription factor subunit alpha) (PSE-binding factor subunit alpha) (PTF subunit alpha). | |||||
|
SNPC4_MOUSE
|
||||||
| θ value | 1.5773e-20 (rank : 8) | NC score | 0.632167 (rank : 7) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 237 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BP86, Q6PGG7, Q80UG9, Q810L1 | Gene names | Snapc4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | snRNA-activating protein complex subunit 4 (SNAPc subunit 4) (snRNA- activating protein complex 190 kDa subunit) (SNAPc 190 kDa subunit). | |||||
|
CDC5L_MOUSE
|
||||||
| θ value | 1.2105e-12 (rank : 9) | NC score | 0.513870 (rank : 10) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6A068, Q8K1J9 | Gene names | Cdc5l, Kiaa0432 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle 5-related protein (Cdc5-like protein). | |||||
|
CDC5L_HUMAN
|
||||||
| θ value | 1.58096e-12 (rank : 10) | NC score | 0.516245 (rank : 9) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q99459, Q76N46, Q99974 | Gene names | CDC5L, KIAA0432, PCDC5RP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle 5-like protein (Cdc5-like protein) (Pombe cdc5- related protein). | |||||
|
MYSM1_MOUSE
|
||||||
| θ value | 0.00175202 (rank : 11) | NC score | 0.197564 (rank : 12) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q69Z66, Q3TPV7, Q8BRP5, Q8C4N1 | Gene names | Mysm1, Kiaa1915 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein MYSM1 (Myb-like, SWIRM and MPN domain-containing protein 1). | |||||
|
MYSM1_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 12) | NC score | 0.202272 (rank : 11) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q5VVJ2, Q7Z3G8, Q96PX3 | Gene names | MYSM1, KIAA1915 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein MYSM1 (Myb-like, SWIRM and MPN domain-containing protein 1). | |||||
|
CENPE_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 13) | NC score | 0.017144 (rank : 36) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 2060 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q02224 | Gene names | CENPE | |||
|
Domain Architecture |

|
|||||
| Description | Centromeric protein E (CENP-E). | |||||
|
ZRF1_HUMAN
|
||||||
| θ value | 0.163984 (rank : 14) | NC score | 0.107418 (rank : 19) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q99543 | Gene names | ZRF1, DNAJC2, MPHOSPH11, MPP11 | |||
|
Domain Architecture |

|
|||||
| Description | Zuotin-related factor 1 (M-phase phosphoprotein 11). | |||||
|
NCOR1_HUMAN
|
||||||
| θ value | 0.21417 (rank : 15) | NC score | 0.112034 (rank : 17) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 577 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O75376, Q9UPV5, Q9UQ18 | Gene names | NCOR1, KIAA1047 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR). | |||||
|
NCOR1_MOUSE
|
||||||
| θ value | 0.21417 (rank : 16) | NC score | 0.111389 (rank : 18) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 521 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q60974, Q60812 | Gene names | Ncor1, Rxrip13 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR) (Retinoid X receptor- interacting protein 13) (RIP13). | |||||
|
TERF1_MOUSE
|
||||||
| θ value | 0.21417 (rank : 17) | NC score | 0.181054 (rank : 13) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P70371, Q9CY71 | Gene names | Terf1, Trf1 | |||
|
Domain Architecture |

|
|||||
| Description | Telomeric repeat-binding factor 1 (TTAGGG repeat-binding factor 1). | |||||
|
NCOR2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 18) | NC score | 0.079942 (rank : 22) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9Y618, O00613, O15416, Q13354, Q9Y5U0 | Gene names | NCOR2, CTG26 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC) (CTG repeat protein 26) (SMAP270). | |||||
|
NCOR2_MOUSE
|
||||||
| θ value | 0.365318 (rank : 19) | NC score | 0.083956 (rank : 21) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 790 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9WU42, Q9WU43, Q9WUC1 | Gene names | Ncor2, Smrt | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC). | |||||
|
SGOL2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 20) | NC score | 0.036938 (rank : 27) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q562F6, Q53RR9, Q53T20, Q86XY4, Q8IWK2, Q8IZK1, Q8N1Q5, Q96LQ3 | Gene names | SGOL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Shugoshin-like 2 (Tripin). | |||||
|
TTF1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 21) | NC score | 0.094122 (rank : 20) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q62187, Q9JKK5 | Gene names | Ttf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription termination factor 1 (TTF-1) (TTF-I) (mTFF-I) (RNA polymerase I termination factor). | |||||
|
PRP40_HUMAN
|
||||||
| θ value | 0.62314 (rank : 22) | NC score | 0.046007 (rank : 24) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 442 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O75400, O43856, O75404, Q8TBQ1, Q9H782, Q9NWU9, Q9P0Q2, Q9Y5A8 | Gene names | PRPF40A, FLAF1, FNBP3, HYPA | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Huntingtin yeast partner A) (Huntingtin-interacting protein HYPA/FBP11) (Fas ligand-associated factor 1) (NY-REN-6 antigen). | |||||
|
ANXA8_MOUSE
|
||||||
| θ value | 0.813845 (rank : 23) | NC score | 0.046062 (rank : 23) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O35640 | Gene names | Anxa8, Anx8 | |||
|
Domain Architecture |
|
|||||
| Description | Annexin A8 (Annexin VIII). | |||||
|
PRP40_MOUSE
|
||||||
| θ value | 0.813845 (rank : 24) | NC score | 0.037852 (rank : 26) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9R1C7, Q61049, Q8BQ76, Q8BRW4, Q8C2U1 | Gene names | Prpf40a, Fbp11, Fnbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Formin-binding protein 11) (FBP-11). | |||||
|
TERF2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 25) | NC score | 0.154622 (rank : 15) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q15554 | Gene names | TERF2, TRF2 | |||
|
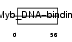
Domain Architecture |
|
|||||
| Description | Telomeric repeat-binding factor 2 (TTAGGG repeat-binding factor 2) (Telomeric DNA-binding protein). | |||||
|
ANXA8_HUMAN
|
||||||
| θ value | 1.38821 (rank : 26) | NC score | 0.040492 (rank : 25) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P13928, Q9BT34 | Gene names | ANXA8, ANX8 | |||
|
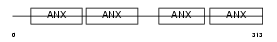
Domain Architecture |
|
|||||
| Description | Annexin A8 (Annexin VIII) (Vascular anticoagulant-beta) (VAC-beta). | |||||
|
TERF1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 27) | NC score | 0.156051 (rank : 14) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P54274, Q15553, Q93029 | Gene names | TERF1, PIN2, TRF, TRF1 | |||
|
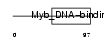
Domain Architecture |
|
|||||
| Description | Telomeric repeat-binding factor 1 (TTAGGG repeat-binding factor 1) (NIMA-interacting protein 2) (Telomeric protein Pin2/TRF1). | |||||
|
NC6IP_HUMAN
|
||||||
| θ value | 2.36792 (rank : 28) | NC score | 0.021199 (rank : 33) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96RS0, Q5GH23, Q8TDG9, Q96QU3, Q9H5V3 | Gene names | NCOA6IP, PIMT | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative RNA methyltransferase NCOA6IP (EC 2.1.1.-) (Nuclear receptor coactivator 6-interacting protein) (PRIP-interacting protein with methyltransferase motif) (PIPMT) (PIMT) (CLL-associated antigen KW-2) (Hepatocellular carcinoma-associated antigen 137) (HCA137). | |||||
|
NIN_HUMAN
|
||||||
| θ value | 2.36792 (rank : 29) | NC score | 0.017564 (rank : 35) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 1416 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8N4C6, Q6P0P6, Q9BWU6, Q9C012, Q9C013, Q9C014, Q9H5I6, Q9HAT7, Q9HBY5, Q9HCK7, Q9UH61 | Gene names | NIN, KIAA1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein (hNinein) (Glycogen synthase kinase 3 beta-interacting protein) (GSK3B-interacting protein). | |||||
|
PRG4_MOUSE
|
||||||
| θ value | 3.0926 (rank : 30) | NC score | 0.015078 (rank : 40) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9JM99, Q3UEL1, Q3V198 | Gene names | Prg4, Msf, Szp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
RERE_MOUSE
|
||||||
| θ value | 3.0926 (rank : 31) | NC score | 0.026237 (rank : 31) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q80TZ9 | Gene names | Rere, Atr2, Kiaa0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-2). | |||||
|
TERF2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 32) | NC score | 0.128521 (rank : 16) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O35144 | Gene names | Terf2, Trf2 | |||
|
Domain Architecture |

|
|||||
| Description | Telomeric repeat-binding factor 2 (TTAGGG repeat-binding factor 2) (Telomeric DNA-binding protein). | |||||
|
ZEP1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 33) | NC score | 0.004801 (rank : 54) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 1036 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P15822, Q14122 | Gene names | HIVEP1, ZNF40 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 40 (Human immunodeficiency virus type I enhancer- binding protein 1) (HIV-EP1) (Major histocompatibility complex-binding protein 1) (MBP-1) (Positive regulatory domain II-binding factor 1) (PRDII-BF1). | |||||
|
CAD15_MOUSE
|
||||||
| θ value | 4.03905 (rank : 34) | NC score | 0.004628 (rank : 55) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P33146, Q9QYZ7 | Gene names | Cdh15, Cdh14 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-15 precursor (Muscle-cadherin) (M-cadherin) (Cadherin-14). | |||||
|
ITSN1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 35) | NC score | 0.015113 (rank : 39) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 1669 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Z0R4, Q9R143 | Gene names | Itsn1, Ese1, Itsn | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-1 (EH and SH3 domains protein 1). | |||||
|
KAP1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 36) | NC score | 0.009034 (rank : 44) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P31321 | Gene names | PRKAR1B | |||
|
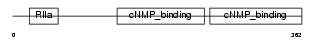
Domain Architecture |
|
|||||
| Description | cAMP-dependent protein kinase type I-beta regulatory subunit. | |||||
|
RERE_HUMAN
|
||||||
| θ value | 4.03905 (rank : 37) | NC score | 0.027503 (rank : 30) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9P2R6, O43393, O75046, O75359, Q5VXL9, Q6P6B9, Q9Y2W4 | Gene names | RERE, ARG, ARP, ATN1L, KIAA0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-1-like protein) (Atrophin-1-related protein). | |||||
|
DAXX_MOUSE
|
||||||
| θ value | 5.27518 (rank : 38) | NC score | 0.022914 (rank : 32) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O35613, Q9QWT8, Q9QWV3 | Gene names | Daxx | |||
|
Domain Architecture |

|
|||||
| Description | Death domain-associated protein 6 (Daxx). | |||||
|
KLH13_MOUSE
|
||||||
| θ value | 5.27518 (rank : 39) | NC score | 0.008628 (rank : 46) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80TF4, Q9DBY7 | Gene names | Klhl13, Bklhd2, Kiaa1309 | |||
|
Domain Architecture |

|
|||||
| Description | Kelch-like protein 13 (BTB and kelch domain-containing protein 2). | |||||
|
ORC2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 40) | NC score | 0.015263 (rank : 38) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q13416, Q13204 | Gene names | ORC2L, ORC2 | |||
|
Domain Architecture |

|
|||||
| Description | Origin recognition complex subunit 2. | |||||
|
PDE6A_MOUSE
|
||||||
| θ value | 5.27518 (rank : 41) | NC score | 0.005437 (rank : 53) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P27664 | Gene names | Pde6a, Mpa, Pdea | |||
|
Domain Architecture |

|
|||||
| Description | Rod cGMP-specific 3',5'-cyclic phosphodiesterase alpha-subunit (EC 3.1.4.35) (GMP-PDE alpha). | |||||
|
SMRC2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 42) | NC score | 0.034328 (rank : 28) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 499 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8TAQ2, Q92923, Q96E12, Q96GY4 | Gene names | SMARCC2, BAF170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
SMRC2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 43) | NC score | 0.032903 (rank : 29) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 487 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q6PDG5, Q6P6P3 | Gene names | Smarcc2, Baf170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
TRDN_HUMAN
|
||||||
| θ value | 5.27518 (rank : 44) | NC score | 0.013977 (rank : 41) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q13061 | Gene names | TRDN | |||
|
Domain Architecture |

|
|||||
| Description | Triadin. | |||||
|
CADH3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 45) | NC score | 0.004353 (rank : 56) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P10287, Q61465 | Gene names | Cdh3, Cdhp | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-3 precursor (Placental-cadherin) (P-cadherin). | |||||
|
CBP_MOUSE
|
||||||
| θ value | 6.88961 (rank : 46) | NC score | 0.009262 (rank : 43) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
KDEL2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 47) | NC score | 0.016461 (rank : 37) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7Z4H8, Q6UWW2, Q6ZUM9, Q8N7L8, Q8NE24 | Gene names | KDELC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | KDEL motif-containing protein 2 precursor. | |||||
|
KLHL9_HUMAN
|
||||||
| θ value | 6.88961 (rank : 48) | NC score | 0.007875 (rank : 48) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9P2J3, Q8TCQ2 | Gene names | KLHL9, KIAA1354 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kelch-like protein 9. | |||||
|
KLHL9_MOUSE
|
||||||
| θ value | 6.88961 (rank : 49) | NC score | 0.007828 (rank : 49) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 195 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6ZPT1 | Gene names | Klhl9, Kiaa1354 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kelch-like protein 9. | |||||
|
MORC1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 50) | NC score | 0.008662 (rank : 45) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q86VD1, Q7L8E2, Q9NSG7, Q9Y6D4 | Gene names | MORC1, MORC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORC family CW-type zinc finger protein 1. | |||||
|
SMC1A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 51) | NC score | 0.008426 (rank : 47) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 1346 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q14683, O14995, Q16351 | Gene names | SMC1L1, DXS423E, KIAA0178, SB1.8, SMC1 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Sb1.8). | |||||
|
CADH3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 52) | NC score | 0.003966 (rank : 57) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P22223 | Gene names | CDH3, CDHP | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-3 precursor (Placental-cadherin) (P-cadherin). | |||||
|
CO6A3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 53) | NC score | 0.005740 (rank : 52) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 317 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P12111, Q16501 | Gene names | COL6A3 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-3(VI) chain precursor. | |||||
|
COBA2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 54) | NC score | 0.002204 (rank : 58) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 501 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q64739, Q61432, Q9Z1W0 | Gene names | Col11a2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(XI) chain precursor. | |||||
|
EF1B_MOUSE
|
||||||
| θ value | 8.99809 (rank : 55) | NC score | 0.009767 (rank : 42) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O70251, Q3THP5, Q5SUH1, Q8CHS1, Q99L22, Q9CZD4 | Gene names | Eef1b, Eef1b2 | |||
|
Domain Architecture |
|
|||||
| Description | Elongation factor 1-beta (EF-1-beta). | |||||
|
MAP1B_HUMAN
|
||||||
| θ value | 8.99809 (rank : 56) | NC score | 0.018432 (rank : 34) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P46821 | Gene names | MAP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) [Contains: MAP1 light chain LC1]. | |||||
|
PEPL_MOUSE
|
||||||
| θ value | 8.99809 (rank : 57) | NC score | 0.007612 (rank : 50) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 1105 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9R269, O70231, Q9CUT1, Q9JLZ7 | Gene names | Ppl | |||
|
Domain Architecture |

|
|||||
| Description | Periplakin. | |||||
|
RASA1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 58) | NC score | 0.006823 (rank : 51) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P20936 | Gene names | RASA1, RASA | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 1 (GTPase-activating protein) (GAP) (Ras p21 protein activator) (p120GAP) (RasGAP). | |||||
|
MYBA_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P51960 | Gene names | Mybl1, Amyb | |||
|
Domain Architecture |

|
|||||
| Description | Myb-related protein A (A-Myb). | |||||
|
MYBA_HUMAN
|
||||||
| NC score | 0.997161 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P10243 | Gene names | MYBL1, AMYB | |||
|
Domain Architecture |

|
|||||
| Description | Myb-related protein A (A-Myb). | |||||
|
MYB_HUMAN
|
||||||
| NC score | 0.939843 (rank : 3) | θ value | 9.035e-101 (rank : 4) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P10242, P78391, P78392, P78525, P78526, Q14023, Q14024, Q9UE83 | Gene names | MYB | |||
|
Domain Architecture |
|
|||||
| Description | Myb proto-oncogene protein (C-myb). | |||||
|
MYB_MOUSE
|
||||||
| NC score | 0.939399 (rank : 4) | θ value | 8.4565e-99 (rank : 5) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P06876, Q61929 | Gene names | Myb | |||
|
Domain Architecture |

|
|||||
| Description | Myb proto-oncogene protein (C-myb). | |||||
|
MYBB_MOUSE
|
||||||
| NC score | 0.939273 (rank : 5) | θ value | 1.93603e-119 (rank : 3) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P48972 | Gene names | Mybl2, Bmyb | |||
|
Domain Architecture |

|
|||||
| Description | Myb-related protein B (B-Myb). | |||||
|
MYBB_HUMAN
|
||||||
| NC score | 0.916712 (rank : 6) | θ value | 7.44272e-79 (rank : 6) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P10244 | Gene names | MYBL2, BMYB | |||
|
Domain Architecture |

|
|||||
| Description | Myb-related protein B (B-Myb). | |||||
|
SNPC4_MOUSE
|
||||||
| NC score | 0.632167 (rank : 7) | θ value | 1.5773e-20 (rank : 8) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 237 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BP86, Q6PGG7, Q80UG9, Q810L1 | Gene names | Snapc4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | snRNA-activating protein complex subunit 4 (SNAPc subunit 4) (snRNA- activating protein complex 190 kDa subunit) (SNAPc 190 kDa subunit). | |||||
|
SNPC4_HUMAN
|
||||||
| NC score | 0.544948 (rank : 8) | θ value | 7.0802e-21 (rank : 7) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q5SXM2, Q9Y6P7 | Gene names | SNAPC4, SNAP190 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | snRNA-activating protein complex subunit 4 (SNAPc subunit 4) (snRNA- activating protein complex 190 kDa subunit) (SNAPc 190 kDa subunit) (Proximal sequence element-binding transcription factor subunit alpha) (PSE-binding factor subunit alpha) (PTF subunit alpha). | |||||
|
CDC5L_HUMAN
|
||||||
| NC score | 0.516245 (rank : 9) | θ value | 1.58096e-12 (rank : 10) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q99459, Q76N46, Q99974 | Gene names | CDC5L, KIAA0432, PCDC5RP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle 5-like protein (Cdc5-like protein) (Pombe cdc5- related protein). | |||||
|
CDC5L_MOUSE
|
||||||
| NC score | 0.513870 (rank : 10) | θ value | 1.2105e-12 (rank : 9) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6A068, Q8K1J9 | Gene names | Cdc5l, Kiaa0432 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle 5-related protein (Cdc5-like protein). | |||||
|
MYSM1_HUMAN
|
||||||
| NC score | 0.202272 (rank : 11) | θ value | 0.00509761 (rank : 12) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q5VVJ2, Q7Z3G8, Q96PX3 | Gene names | MYSM1, KIAA1915 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein MYSM1 (Myb-like, SWIRM and MPN domain-containing protein 1). | |||||
|
MYSM1_MOUSE
|
||||||
| NC score | 0.197564 (rank : 12) | θ value | 0.00175202 (rank : 11) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q69Z66, Q3TPV7, Q8BRP5, Q8C4N1 | Gene names | Mysm1, Kiaa1915 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein MYSM1 (Myb-like, SWIRM and MPN domain-containing protein 1). | |||||
|
TERF1_MOUSE
|
||||||
| NC score | 0.181054 (rank : 13) | θ value | 0.21417 (rank : 17) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P70371, Q9CY71 | Gene names | Terf1, Trf1 | |||
|
Domain Architecture |

|
|||||
| Description | Telomeric repeat-binding factor 1 (TTAGGG repeat-binding factor 1). | |||||
|
TERF1_HUMAN
|
||||||
| NC score | 0.156051 (rank : 14) | θ value | 1.38821 (rank : 27) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P54274, Q15553, Q93029 | Gene names | TERF1, PIN2, TRF, TRF1 | |||
|
Domain Architecture |

|
|||||
| Description | Telomeric repeat-binding factor 1 (TTAGGG repeat-binding factor 1) (NIMA-interacting protein 2) (Telomeric protein Pin2/TRF1). | |||||
|
TERF2_HUMAN
|
||||||
| NC score | 0.154622 (rank : 15) | θ value | 1.06291 (rank : 25) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q15554 | Gene names | TERF2, TRF2 | |||
|
Domain Architecture |

|
|||||
| Description | Telomeric repeat-binding factor 2 (TTAGGG repeat-binding factor 2) (Telomeric DNA-binding protein). | |||||
|
TERF2_MOUSE
|
||||||
| NC score | 0.128521 (rank : 16) | θ value | 3.0926 (rank : 32) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O35144 | Gene names | Terf2, Trf2 | |||
|
Domain Architecture |

|
|||||
| Description | Telomeric repeat-binding factor 2 (TTAGGG repeat-binding factor 2) (Telomeric DNA-binding protein). | |||||
|
NCOR1_HUMAN
|
||||||
| NC score | 0.112034 (rank : 17) | θ value | 0.21417 (rank : 15) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 577 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O75376, Q9UPV5, Q9UQ18 | Gene names | NCOR1, KIAA1047 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR). | |||||
|
NCOR1_MOUSE
|
||||||
| NC score | 0.111389 (rank : 18) | θ value | 0.21417 (rank : 16) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 521 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q60974, Q60812 | Gene names | Ncor1, Rxrip13 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR) (Retinoid X receptor- interacting protein 13) (RIP13). | |||||
|
ZRF1_HUMAN
|
||||||
| NC score | 0.107418 (rank : 19) | θ value | 0.163984 (rank : 14) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q99543 | Gene names | ZRF1, DNAJC2, MPHOSPH11, MPP11 | |||
|
Domain Architecture |

|
|||||
| Description | Zuotin-related factor 1 (M-phase phosphoprotein 11). | |||||
|
TTF1_MOUSE
|
||||||
| NC score | 0.094122 (rank : 20) | θ value | 0.47712 (rank : 21) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q62187, Q9JKK5 | Gene names | Ttf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription termination factor 1 (TTF-1) (TTF-I) (mTFF-I) (RNA polymerase I termination factor). | |||||
|
NCOR2_MOUSE
|
||||||
| NC score | 0.083956 (rank : 21) | θ value | 0.365318 (rank : 19) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 790 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9WU42, Q9WU43, Q9WUC1 | Gene names | Ncor2, Smrt | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC). | |||||
|
NCOR2_HUMAN
|
||||||
| NC score | 0.079942 (rank : 22) | θ value | 0.365318 (rank : 18) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9Y618, O00613, O15416, Q13354, Q9Y5U0 | Gene names | NCOR2, CTG26 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC) (CTG repeat protein 26) (SMAP270). | |||||
|
ANXA8_MOUSE
|
||||||
| NC score | 0.046062 (rank : 23) | θ value | 0.813845 (rank : 23) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O35640 | Gene names | Anxa8, Anx8 | |||
|
Domain Architecture |

|
|||||
| Description | Annexin A8 (Annexin VIII). | |||||
|
PRP40_HUMAN
|
||||||
| NC score | 0.046007 (rank : 24) | θ value | 0.62314 (rank : 22) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 442 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O75400, O43856, O75404, Q8TBQ1, Q9H782, Q9NWU9, Q9P0Q2, Q9Y5A8 | Gene names | PRPF40A, FLAF1, FNBP3, HYPA | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Huntingtin yeast partner A) (Huntingtin-interacting protein HYPA/FBP11) (Fas ligand-associated factor 1) (NY-REN-6 antigen). | |||||
|
ANXA8_HUMAN
|
||||||
| NC score | 0.040492 (rank : 25) | θ value | 1.38821 (rank : 26) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P13928, Q9BT34 | Gene names | ANXA8, ANX8 | |||
|
Domain Architecture |

|
|||||
| Description | Annexin A8 (Annexin VIII) (Vascular anticoagulant-beta) (VAC-beta). | |||||
|
PRP40_MOUSE
|
||||||
| NC score | 0.037852 (rank : 26) | θ value | 0.813845 (rank : 24) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9R1C7, Q61049, Q8BQ76, Q8BRW4, Q8C2U1 | Gene names | Prpf40a, Fbp11, Fnbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Formin-binding protein 11) (FBP-11). | |||||
|
SGOL2_HUMAN
|
||||||
| NC score | 0.036938 (rank : 27) | θ value | 0.365318 (rank : 20) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q562F6, Q53RR9, Q53T20, Q86XY4, Q8IWK2, Q8IZK1, Q8N1Q5, Q96LQ3 | Gene names | SGOL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Shugoshin-like 2 (Tripin). | |||||
|
SMRC2_HUMAN
|
||||||
| NC score | 0.034328 (rank : 28) | θ value | 5.27518 (rank : 42) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 499 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8TAQ2, Q92923, Q96E12, Q96GY4 | Gene names | SMARCC2, BAF170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
SMRC2_MOUSE
|
||||||
| NC score | 0.032903 (rank : 29) | θ value | 5.27518 (rank : 43) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 487 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q6PDG5, Q6P6P3 | Gene names | Smarcc2, Baf170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
RERE_HUMAN
|
||||||
| NC score | 0.027503 (rank : 30) | θ value | 4.03905 (rank : 37) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9P2R6, O43393, O75046, O75359, Q5VXL9, Q6P6B9, Q9Y2W4 | Gene names | RERE, ARG, ARP, ATN1L, KIAA0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-1-like protein) (Atrophin-1-related protein). | |||||
|
RERE_MOUSE
|
||||||
| NC score | 0.026237 (rank : 31) | θ value | 3.0926 (rank : 31) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q80TZ9 | Gene names | Rere, Atr2, Kiaa0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-2). | |||||
|
DAXX_MOUSE
|
||||||
| NC score | 0.022914 (rank : 32) | θ value | 5.27518 (rank : 38) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O35613, Q9QWT8, Q9QWV3 | Gene names | Daxx | |||
|
Domain Architecture |

|
|||||
| Description | Death domain-associated protein 6 (Daxx). | |||||
|
NC6IP_HUMAN
|
||||||
| NC score | 0.021199 (rank : 33) | θ value | 2.36792 (rank : 28) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96RS0, Q5GH23, Q8TDG9, Q96QU3, Q9H5V3 | Gene names | NCOA6IP, PIMT | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative RNA methyltransferase NCOA6IP (EC 2.1.1.-) (Nuclear receptor coactivator 6-interacting protein) (PRIP-interacting protein with methyltransferase motif) (PIPMT) (PIMT) (CLL-associated antigen KW-2) (Hepatocellular carcinoma-associated antigen 137) (HCA137). | |||||
|
MAP1B_HUMAN
|
||||||
| NC score | 0.018432 (rank : 34) | θ value | 8.99809 (rank : 56) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P46821 | Gene names | MAP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) [Contains: MAP1 light chain LC1]. | |||||
|
NIN_HUMAN
|
||||||
| NC score | 0.017564 (rank : 35) | θ value | 2.36792 (rank : 29) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 1416 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8N4C6, Q6P0P6, Q9BWU6, Q9C012, Q9C013, Q9C014, Q9H5I6, Q9HAT7, Q9HBY5, Q9HCK7, Q9UH61 | Gene names | NIN, KIAA1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein (hNinein) (Glycogen synthase kinase 3 beta-interacting protein) (GSK3B-interacting protein). | |||||
|
CENPE_HUMAN
|
||||||
| NC score | 0.017144 (rank : 36) | θ value | 0.0252991 (rank : 13) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 2060 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q02224 | Gene names | CENPE | |||
|
Domain Architecture |

|
|||||
| Description | Centromeric protein E (CENP-E). | |||||
|
KDEL2_HUMAN
|
||||||
| NC score | 0.016461 (rank : 37) | θ value | 6.88961 (rank : 47) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7Z4H8, Q6UWW2, Q6ZUM9, Q8N7L8, Q8NE24 | Gene names | KDELC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | KDEL motif-containing protein 2 precursor. | |||||
|
ORC2_HUMAN
|
||||||
| NC score | 0.015263 (rank : 38) | θ value | 5.27518 (rank : 40) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q13416, Q13204 | Gene names | ORC2L, ORC2 | |||
|
Domain Architecture |

|
|||||
| Description | Origin recognition complex subunit 2. | |||||
|
ITSN1_MOUSE
|
||||||
| NC score | 0.015113 (rank : 39) | θ value | 4.03905 (rank : 35) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 1669 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Z0R4, Q9R143 | Gene names | Itsn1, Ese1, Itsn | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-1 (EH and SH3 domains protein 1). | |||||
|
PRG4_MOUSE
|
||||||
| NC score | 0.015078 (rank : 40) | θ value | 3.0926 (rank : 30) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9JM99, Q3UEL1, Q3V198 | Gene names | Prg4, Msf, Szp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
TRDN_HUMAN
|
||||||
| NC score | 0.013977 (rank : 41) | θ value | 5.27518 (rank : 44) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q13061 | Gene names | TRDN | |||
|
Domain Architecture |

|
|||||
| Description | Triadin. | |||||
|
EF1B_MOUSE
|
||||||
| NC score | 0.009767 (rank : 42) | θ value | 8.99809 (rank : 55) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O70251, Q3THP5, Q5SUH1, Q8CHS1, Q99L22, Q9CZD4 | Gene names | Eef1b, Eef1b2 | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor 1-beta (EF-1-beta). | |||||
|
CBP_MOUSE
|
||||||
| NC score | 0.009262 (rank : 43) | θ value | 6.88961 (rank : 46) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
KAP1_HUMAN
|
||||||
| NC score | 0.009034 (rank : 44) | θ value | 4.03905 (rank : 36) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P31321 | Gene names | PRKAR1B | |||
|
Domain Architecture |

|
|||||
| Description | cAMP-dependent protein kinase type I-beta regulatory subunit. | |||||
|
MORC1_HUMAN
|
||||||
| NC score | 0.008662 (rank : 45) | θ value | 6.88961 (rank : 50) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q86VD1, Q7L8E2, Q9NSG7, Q9Y6D4 | Gene names | MORC1, MORC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORC family CW-type zinc finger protein 1. | |||||
|
KLH13_MOUSE
|
||||||
| NC score | 0.008628 (rank : 46) | θ value | 5.27518 (rank : 39) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80TF4, Q9DBY7 | Gene names | Klhl13, Bklhd2, Kiaa1309 | |||
|
Domain Architecture |

|
|||||
| Description | Kelch-like protein 13 (BTB and kelch domain-containing protein 2). | |||||
|
SMC1A_HUMAN
|
||||||
| NC score | 0.008426 (rank : 47) | θ value | 6.88961 (rank : 51) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 1346 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q14683, O14995, Q16351 | Gene names | SMC1L1, DXS423E, KIAA0178, SB1.8, SMC1 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Sb1.8). | |||||
|
KLHL9_HUMAN
|
||||||
| NC score | 0.007875 (rank : 48) | θ value | 6.88961 (rank : 48) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9P2J3, Q8TCQ2 | Gene names | KLHL9, KIAA1354 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kelch-like protein 9. | |||||
|
KLHL9_MOUSE
|
||||||
| NC score | 0.007828 (rank : 49) | θ value | 6.88961 (rank : 49) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 195 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6ZPT1 | Gene names | Klhl9, Kiaa1354 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kelch-like protein 9. | |||||
|
PEPL_MOUSE
|
||||||
| NC score | 0.007612 (rank : 50) | θ value | 8.99809 (rank : 57) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 1105 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9R269, O70231, Q9CUT1, Q9JLZ7 | Gene names | Ppl | |||
|
Domain Architecture |

|
|||||
| Description | Periplakin. | |||||
|
RASA1_HUMAN
|
||||||
| NC score | 0.006823 (rank : 51) | θ value | 8.99809 (rank : 58) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P20936 | Gene names | RASA1, RASA | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 1 (GTPase-activating protein) (GAP) (Ras p21 protein activator) (p120GAP) (RasGAP). | |||||
|
CO6A3_HUMAN
|
||||||
| NC score | 0.005740 (rank : 52) | θ value | 8.99809 (rank : 53) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 317 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P12111, Q16501 | Gene names | COL6A3 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-3(VI) chain precursor. | |||||
|
PDE6A_MOUSE
|
||||||
| NC score | 0.005437 (rank : 53) | θ value | 5.27518 (rank : 41) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P27664 | Gene names | Pde6a, Mpa, Pdea | |||
|
Domain Architecture |

|
|||||
| Description | Rod cGMP-specific 3',5'-cyclic phosphodiesterase alpha-subunit (EC 3.1.4.35) (GMP-PDE alpha). | |||||
|
ZEP1_HUMAN
|
||||||
| NC score | 0.004801 (rank : 54) | θ value | 3.0926 (rank : 33) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 1036 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P15822, Q14122 | Gene names | HIVEP1, ZNF40 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 40 (Human immunodeficiency virus type I enhancer- binding protein 1) (HIV-EP1) (Major histocompatibility complex-binding protein 1) (MBP-1) (Positive regulatory domain II-binding factor 1) (PRDII-BF1). | |||||
|
CAD15_MOUSE
|
||||||
| NC score | 0.004628 (rank : 55) | θ value | 4.03905 (rank : 34) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P33146, Q9QYZ7 | Gene names | Cdh15, Cdh14 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-15 precursor (Muscle-cadherin) (M-cadherin) (Cadherin-14). | |||||
|
CADH3_MOUSE
|
||||||
| NC score | 0.004353 (rank : 56) | θ value | 6.88961 (rank : 45) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P10287, Q61465 | Gene names | Cdh3, Cdhp | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-3 precursor (Placental-cadherin) (P-cadherin). | |||||
|
CADH3_HUMAN
|
||||||
| NC score | 0.003966 (rank : 57) | θ value | 8.99809 (rank : 52) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P22223 | Gene names | CDH3, CDHP | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-3 precursor (Placental-cadherin) (P-cadherin). | |||||
|
COBA2_MOUSE
|
||||||
| NC score | 0.002204 (rank : 58) | θ value | 8.99809 (rank : 54) | |||
| Query Neighborhood Hits | 58 | Target Neighborhood Hits | 501 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q64739, Q61432, Q9Z1W0 | Gene names | Col11a2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(XI) chain precursor. | |||||