Please be patient as the page loads
|
HPS3_MOUSE
|
||||||
| SwissProt Accessions | Q91VB4, Q8VDX2, Q8VHT1 | Gene names | Hps3, Coa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hermansky-Pudlak syndrome 3 protein homolog (Cocoa protein). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
HPS3_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.994380 (rank : 2) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q969F9, Q8WTV6, Q96AP1, Q96MR3, Q9H608 | Gene names | HPS3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hermansky-Pudlak syndrome 3 protein. | |||||
|
HPS3_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q91VB4, Q8VDX2, Q8VHT1 | Gene names | Hps3, Coa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hermansky-Pudlak syndrome 3 protein homolog (Cocoa protein). | |||||
|
CE005_HUMAN
|
||||||
| θ value | 0.365318 (rank : 3) | NC score | 0.030197 (rank : 7) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NYF5, Q6PGQ2, Q9P0I7 | Gene names | C5orf5 | |||
|
Domain Architecture |
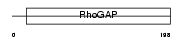
|
|||||
| Description | Protein C5orf5 (GAP-like protein N61). | |||||
|
UXS1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 4) | NC score | 0.046436 (rank : 4) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NBZ7, Q8NBX3, Q9H5C2 | Gene names | UXS1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UDP-glucuronic acid decarboxylase 1 (EC 4.1.1.35) (UDP-glucuronate decarboxylase 1) (UGD) (UXS-1). | |||||
|
UXS1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 5) | NC score | 0.046444 (rank : 3) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q91XL3, Q3U854 | Gene names | Uxs1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UDP-glucuronic acid decarboxylase 1 (EC 4.1.1.35) (UDP-glucuronate decarboxylase 1) (UGD) (UXS-1). | |||||
|
RFX2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 6) | NC score | 0.036027 (rank : 6) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P48378 | Gene names | RFX2 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-binding protein RFX2. | |||||
|
UROL1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 7) | NC score | 0.017672 (rank : 8) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5DID0, Q5DIC9, Q6LA40, Q6LA41, Q8N216 | Gene names | UMODL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uromodulin-like 1 precursor (Olfactorin). | |||||
|
LAX1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 8) | NC score | 0.036725 (rank : 5) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8IWV1, Q6NSZ6, Q9NXB4 | Gene names | LAX1, LAX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lymphocyte transmembrane adapter 1 (Membrane-associated adapter protein LAX) (Linker for activation of X cells). | |||||
|
SMUF1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 9) | NC score | 0.011733 (rank : 9) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9HCE7, O75853, Q9UJT8 | Gene names | SMURF1, KIAA1625 | |||
|
Domain Architecture |

|
|||||
| Description | Smad ubiquitination regulatory factor 1 (EC 6.3.2.-) (Ubiquitin-- protein ligase SMURF1) (Smad-specific E3 ubiquitin ligase 1) (hSMURF1). | |||||
|
SMUF1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 10) | NC score | 0.011638 (rank : 10) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9CUN6, Q3U412, Q8K300 | Gene names | Smurf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Smad ubiquitination regulatory factor 1 (EC 6.3.2.-) (Ubiquitin-- protein ligase SMURF1) (Smad-specific E3 ubiquitin ligase 1). | |||||
|
CEND2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 11) | NC score | 0.010102 (rank : 11) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96P48, O94879, Q59FI7, Q6PHS3, Q8WU51, Q96HP6, Q96L71 | Gene names | CENTD2, ARAP1, KIAA0782 | |||
|
Domain Architecture |

|
|||||
| Description | Centaurin-delta 2 (Cnt-d2) (Arf-GAP, Rho-GAP, ankyrin repeat and pleckstrin homology domain-containing protein 1). | |||||
|
AMGO2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 12) | NC score | 0.004752 (rank : 14) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q86SJ2, Q7Z4A0, Q96CN8 | Gene names | AMIGO2, ALI1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amphoterin-induced protein 2 precursor (AMIGO-2) (Alivin-1) (Differentially expressed in gastric adenocarcinomas) (DEGA). | |||||
|
PTN12_HUMAN
|
||||||
| θ value | 8.99809 (rank : 13) | NC score | 0.007030 (rank : 13) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q05209, Q16130, Q59FD6, Q75MN8, Q86XU4 | Gene names | PTPN12 | |||
|
Domain Architecture |
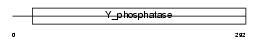
|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 12 (EC 3.1.3.48) (Protein-tyrosine phosphatase G1) (PTPG1). | |||||
|
PTN12_MOUSE
|
||||||
| θ value | 8.99809 (rank : 14) | NC score | 0.007049 (rank : 12) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P35831 | Gene names | Ptpn12 | |||
|
Domain Architecture |
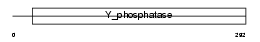
|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 12 (EC 3.1.3.48) (Protein-tyrosine phosphatase P19) (P19-PTP) (MPTP-PEST). | |||||
|
HPS3_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q91VB4, Q8VDX2, Q8VHT1 | Gene names | Hps3, Coa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hermansky-Pudlak syndrome 3 protein homolog (Cocoa protein). | |||||
|
HPS3_HUMAN
|
||||||
| NC score | 0.994380 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q969F9, Q8WTV6, Q96AP1, Q96MR3, Q9H608 | Gene names | HPS3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hermansky-Pudlak syndrome 3 protein. | |||||
|
UXS1_MOUSE
|
||||||
| NC score | 0.046444 (rank : 3) | θ value | 1.38821 (rank : 5) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q91XL3, Q3U854 | Gene names | Uxs1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UDP-glucuronic acid decarboxylase 1 (EC 4.1.1.35) (UDP-glucuronate decarboxylase 1) (UGD) (UXS-1). | |||||
|
UXS1_HUMAN
|
||||||
| NC score | 0.046436 (rank : 4) | θ value | 1.38821 (rank : 4) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NBZ7, Q8NBX3, Q9H5C2 | Gene names | UXS1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UDP-glucuronic acid decarboxylase 1 (EC 4.1.1.35) (UDP-glucuronate decarboxylase 1) (UGD) (UXS-1). | |||||
|
LAX1_HUMAN
|
||||||
| NC score | 0.036725 (rank : 5) | θ value | 5.27518 (rank : 8) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8IWV1, Q6NSZ6, Q9NXB4 | Gene names | LAX1, LAX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lymphocyte transmembrane adapter 1 (Membrane-associated adapter protein LAX) (Linker for activation of X cells). | |||||
|
RFX2_HUMAN
|
||||||
| NC score | 0.036027 (rank : 6) | θ value | 3.0926 (rank : 6) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P48378 | Gene names | RFX2 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-binding protein RFX2. | |||||
|
CE005_HUMAN
|
||||||
| NC score | 0.030197 (rank : 7) | θ value | 0.365318 (rank : 3) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NYF5, Q6PGQ2, Q9P0I7 | Gene names | C5orf5 | |||
|
Domain Architecture |

|
|||||
| Description | Protein C5orf5 (GAP-like protein N61). | |||||
|
UROL1_HUMAN
|
||||||
| NC score | 0.017672 (rank : 8) | θ value | 4.03905 (rank : 7) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5DID0, Q5DIC9, Q6LA40, Q6LA41, Q8N216 | Gene names | UMODL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uromodulin-like 1 precursor (Olfactorin). | |||||
|
SMUF1_HUMAN
|
||||||
| NC score | 0.011733 (rank : 9) | θ value | 5.27518 (rank : 9) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9HCE7, O75853, Q9UJT8 | Gene names | SMURF1, KIAA1625 | |||
|
Domain Architecture |

|
|||||
| Description | Smad ubiquitination regulatory factor 1 (EC 6.3.2.-) (Ubiquitin-- protein ligase SMURF1) (Smad-specific E3 ubiquitin ligase 1) (hSMURF1). | |||||
|
SMUF1_MOUSE
|
||||||
| NC score | 0.011638 (rank : 10) | θ value | 5.27518 (rank : 10) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9CUN6, Q3U412, Q8K300 | Gene names | Smurf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Smad ubiquitination regulatory factor 1 (EC 6.3.2.-) (Ubiquitin-- protein ligase SMURF1) (Smad-specific E3 ubiquitin ligase 1). | |||||
|
CEND2_HUMAN
|
||||||
| NC score | 0.010102 (rank : 11) | θ value | 6.88961 (rank : 11) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96P48, O94879, Q59FI7, Q6PHS3, Q8WU51, Q96HP6, Q96L71 | Gene names | CENTD2, ARAP1, KIAA0782 | |||
|
Domain Architecture |

|
|||||
| Description | Centaurin-delta 2 (Cnt-d2) (Arf-GAP, Rho-GAP, ankyrin repeat and pleckstrin homology domain-containing protein 1). | |||||
|
PTN12_MOUSE
|
||||||
| NC score | 0.007049 (rank : 12) | θ value | 8.99809 (rank : 14) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P35831 | Gene names | Ptpn12 | |||
|
Domain Architecture |
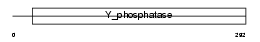
|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 12 (EC 3.1.3.48) (Protein-tyrosine phosphatase P19) (P19-PTP) (MPTP-PEST). | |||||
|
PTN12_HUMAN
|
||||||
| NC score | 0.007030 (rank : 13) | θ value | 8.99809 (rank : 13) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q05209, Q16130, Q59FD6, Q75MN8, Q86XU4 | Gene names | PTPN12 | |||
|
Domain Architecture |
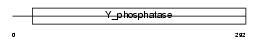
|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 12 (EC 3.1.3.48) (Protein-tyrosine phosphatase G1) (PTPG1). | |||||
|
AMGO2_HUMAN
|
||||||
| NC score | 0.004752 (rank : 14) | θ value | 8.99809 (rank : 12) | |||
| Query Neighborhood Hits | 14 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q86SJ2, Q7Z4A0, Q96CN8 | Gene names | AMIGO2, ALI1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amphoterin-induced protein 2 precursor (AMIGO-2) (Alivin-1) (Differentially expressed in gastric adenocarcinomas) (DEGA). | |||||