Please be patient as the page loads
|
GDIR_HUMAN
|
||||||
| SwissProt Accessions | P52565 | Gene names | ARHGDIA, GDIA1 | |||
|
Domain Architecture |
|
|||||
| Description | Rho GDP-dissociation inhibitor 1 (Rho GDI 1) (Rho-GDI alpha). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
GDIR_HUMAN
|
||||||
| θ value | 3.1986e-114 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P52565 | Gene names | ARHGDIA, GDIA1 | |||
|
Domain Architecture |
|
|||||
| Description | Rho GDP-dissociation inhibitor 1 (Rho GDI 1) (Rho-GDI alpha). | |||||
|
GDIR_MOUSE
|
||||||
| θ value | 1.13764e-111 (rank : 2) | NC score | 0.998689 (rank : 2) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q99PT1, Q99KC4 | Gene names | Arhgdia, C87222, Gdi1 | |||
|
Domain Architecture |
|
|||||
| Description | Rho GDP-dissociation inhibitor 1 (Rho GDI 1) (Rho-GDI alpha) (GDI-1). | |||||
|
GDIS_MOUSE
|
||||||
| θ value | 2.82823e-78 (rank : 3) | NC score | 0.992211 (rank : 4) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61599, Q3TEB3 | Gene names | Arhgdib, Gdid4 | |||
|
Domain Architecture |
|
|||||
| Description | Rho GDP-dissociation inhibitor 2 (Rho GDI 2) (Rho-GDI beta) (D4). | |||||
|
GDIS_HUMAN
|
||||||
| θ value | 4.08394e-77 (rank : 4) | NC score | 0.992739 (rank : 3) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P52566 | Gene names | ARHGDIB, GDID4 | |||
|
Domain Architecture |

|
|||||
| Description | Rho GDP-dissociation inhibitor 2 (Rho GDI 2) (Rho-GDI beta) (Ly-GDI). | |||||
|
GDIT_HUMAN
|
||||||
| θ value | 1.04338e-64 (rank : 5) | NC score | 0.985054 (rank : 5) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99819, Q4TT69, Q96S29 | Gene names | ARHGDIG | |||
|
Domain Architecture |
|
|||||
| Description | Rho GDP-dissociation inhibitor 3 (Rho GDI 3) (Rho-GDI gamma). | |||||
|
GDIT_MOUSE
|
||||||
| θ value | 8.26713e-62 (rank : 6) | NC score | 0.983592 (rank : 6) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q62160 | Gene names | Arhgdig, Gdi5 | |||
|
Domain Architecture |
|
|||||
| Description | Rho GDP-dissociation inhibitor 3 (Rho GDI 3) (Rho-GDI gamma) (Rho- GDI2). | |||||
|
DC1I1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 7) | NC score | 0.037079 (rank : 8) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O14576, Q9Y2X1 | Gene names | DYNC1I1, DNCI1, DNCIC1 | |||
|
Domain Architecture |

|
|||||
| Description | Cytoplasmic dynein 1 intermediate chain 1 (Dynein intermediate chain 1, cytosolic) (DH IC-1) (Cytoplasmic dynein intermediate chain 1). | |||||
|
DC1I1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 8) | NC score | 0.037187 (rank : 7) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O88485, O88486 | Gene names | Dync1i1, Dnci1, Dncic1 | |||
|
Domain Architecture |

|
|||||
| Description | Cytoplasmic dynein 1 intermediate chain 1 (Dynein intermediate chain 1, cytosolic) (DH IC-1) (Cytoplasmic dynein intermediate chain 1). | |||||
|
AP3M2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 9) | NC score | 0.027481 (rank : 10) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P53677 | Gene names | AP3M2 | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit mu-2 (Adapter-related protein complex 3 mu-2 subunit) (Clathrin coat assembly protein AP47 homolog 2) (Clathrin coat-associated protein AP47 homolog 2) (Golgi adaptor AP-1 47 kDa protein homolog 2) (HA1 47 kDa subunit homolog 2) (Clathrin assembly protein assembly protein complex 1 medium chain homolog 2) (P47B). | |||||
|
RB6I2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 10) | NC score | 0.006854 (rank : 16) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 1584 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8IUD2, Q6NVK2, Q8IUD3, Q8IUD4, Q8IUD5, Q8NAS1, Q9NXN5, Q9UIK7, Q9UPS1 | Gene names | ERC1, ELKS, KIAA1081, RAB6IP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ELKS/RAB6-interacting/CAST family member 1 (RAB6-interacting protein 2) (ERC protein 1). | |||||
|
RB6I2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 11) | NC score | 0.007454 (rank : 15) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 1433 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99MI1, Q80TK7, Q8BPL1, Q8C7Y1, Q99MI2 | Gene names | Erc1, Cast2, Kiaa1081, Rab6ip2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ELKS/RAB6-interacting/CAST family member 1 (RAB6-interacting protein 2) (ERC protein 1) (ERC1) (CAZ-associated structural protein 2) (CAST2). | |||||
|
CCDC4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 12) | NC score | 0.032772 (rank : 9) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6ZU67, Q58A26, Q58A27 | Gene names | CCDC4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 4. | |||||
|
PYR5_MOUSE
|
||||||
| θ value | 5.27518 (rank : 13) | NC score | 0.023977 (rank : 11) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P13439, Q99L26 | Gene names | Umps | |||
|
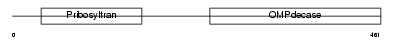
Domain Architecture |
|
|||||
| Description | Uridine 5'-monophosphate synthase (UMP synthase) [Includes: Orotate phosphoribosyltransferase (EC 2.4.2.10) (OPRTase); Orotidine 5'- phosphate decarboxylase (EC 4.1.1.23) (OMPdecase)]. | |||||
|
SSFA2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 14) | NC score | 0.020332 (rank : 12) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q922B9, Q8VEF7 | Gene names | Ssfa2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-specific antigen 2. | |||||
|
AP3M1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 15) | NC score | 0.018459 (rank : 14) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y2T2 | Gene names | AP3M1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit mu-1 (Adapter-related protein complex 3 mu-1 subunit) (Mu-adaptin 3A) (AP-3 adapter complex mu3A subunit). | |||||
|
AP3M1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 16) | NC score | 0.018470 (rank : 13) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9JKC8, Q5BKQ6 | Gene names | Ap3m1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit mu-1 (Adapter-related protein complex 3 mu-1 subunit) (Mu-adaptin 3A) (AP-3 adapter complex mu3A subunit). | |||||
|
GDIR_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 3.1986e-114 (rank : 1) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P52565 | Gene names | ARHGDIA, GDIA1 | |||
|
Domain Architecture |
|
|||||
| Description | Rho GDP-dissociation inhibitor 1 (Rho GDI 1) (Rho-GDI alpha). | |||||
|
GDIR_MOUSE
|
||||||
| NC score | 0.998689 (rank : 2) | θ value | 1.13764e-111 (rank : 2) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q99PT1, Q99KC4 | Gene names | Arhgdia, C87222, Gdi1 | |||
|
Domain Architecture |
|
|||||
| Description | Rho GDP-dissociation inhibitor 1 (Rho GDI 1) (Rho-GDI alpha) (GDI-1). | |||||
|
GDIS_HUMAN
|
||||||
| NC score | 0.992739 (rank : 3) | θ value | 4.08394e-77 (rank : 4) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P52566 | Gene names | ARHGDIB, GDID4 | |||
|
Domain Architecture |

|
|||||
| Description | Rho GDP-dissociation inhibitor 2 (Rho GDI 2) (Rho-GDI beta) (Ly-GDI). | |||||
|
GDIS_MOUSE
|
||||||
| NC score | 0.992211 (rank : 4) | θ value | 2.82823e-78 (rank : 3) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61599, Q3TEB3 | Gene names | Arhgdib, Gdid4 | |||
|
Domain Architecture |
|
|||||
| Description | Rho GDP-dissociation inhibitor 2 (Rho GDI 2) (Rho-GDI beta) (D4). | |||||
|
GDIT_HUMAN
|
||||||
| NC score | 0.985054 (rank : 5) | θ value | 1.04338e-64 (rank : 5) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99819, Q4TT69, Q96S29 | Gene names | ARHGDIG | |||
|
Domain Architecture |

|
|||||
| Description | Rho GDP-dissociation inhibitor 3 (Rho GDI 3) (Rho-GDI gamma). | |||||
|
GDIT_MOUSE
|
||||||
| NC score | 0.983592 (rank : 6) | θ value | 8.26713e-62 (rank : 6) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q62160 | Gene names | Arhgdig, Gdi5 | |||
|
Domain Architecture |

|
|||||
| Description | Rho GDP-dissociation inhibitor 3 (Rho GDI 3) (Rho-GDI gamma) (Rho- GDI2). | |||||
|
DC1I1_MOUSE
|
||||||
| NC score | 0.037187 (rank : 7) | θ value | 1.38821 (rank : 8) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O88485, O88486 | Gene names | Dync1i1, Dnci1, Dncic1 | |||
|
Domain Architecture |

|
|||||
| Description | Cytoplasmic dynein 1 intermediate chain 1 (Dynein intermediate chain 1, cytosolic) (DH IC-1) (Cytoplasmic dynein intermediate chain 1). | |||||
|
DC1I1_HUMAN
|
||||||
| NC score | 0.037079 (rank : 8) | θ value | 1.38821 (rank : 7) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O14576, Q9Y2X1 | Gene names | DYNC1I1, DNCI1, DNCIC1 | |||
|
Domain Architecture |

|
|||||
| Description | Cytoplasmic dynein 1 intermediate chain 1 (Dynein intermediate chain 1, cytosolic) (DH IC-1) (Cytoplasmic dynein intermediate chain 1). | |||||
|
CCDC4_HUMAN
|
||||||
| NC score | 0.032772 (rank : 9) | θ value | 5.27518 (rank : 12) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6ZU67, Q58A26, Q58A27 | Gene names | CCDC4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 4. | |||||
|
AP3M2_HUMAN
|
||||||
| NC score | 0.027481 (rank : 10) | θ value | 2.36792 (rank : 9) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P53677 | Gene names | AP3M2 | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit mu-2 (Adapter-related protein complex 3 mu-2 subunit) (Clathrin coat assembly protein AP47 homolog 2) (Clathrin coat-associated protein AP47 homolog 2) (Golgi adaptor AP-1 47 kDa protein homolog 2) (HA1 47 kDa subunit homolog 2) (Clathrin assembly protein assembly protein complex 1 medium chain homolog 2) (P47B). | |||||
|
PYR5_MOUSE
|
||||||
| NC score | 0.023977 (rank : 11) | θ value | 5.27518 (rank : 13) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P13439, Q99L26 | Gene names | Umps | |||
|
Domain Architecture |

|
|||||
| Description | Uridine 5'-monophosphate synthase (UMP synthase) [Includes: Orotate phosphoribosyltransferase (EC 2.4.2.10) (OPRTase); Orotidine 5'- phosphate decarboxylase (EC 4.1.1.23) (OMPdecase)]. | |||||
|
SSFA2_MOUSE
|
||||||
| NC score | 0.020332 (rank : 12) | θ value | 5.27518 (rank : 14) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q922B9, Q8VEF7 | Gene names | Ssfa2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-specific antigen 2. | |||||
|
AP3M1_MOUSE
|
||||||
| NC score | 0.018470 (rank : 13) | θ value | 8.99809 (rank : 16) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9JKC8, Q5BKQ6 | Gene names | Ap3m1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit mu-1 (Adapter-related protein complex 3 mu-1 subunit) (Mu-adaptin 3A) (AP-3 adapter complex mu3A subunit). | |||||
|
AP3M1_HUMAN
|
||||||
| NC score | 0.018459 (rank : 14) | θ value | 8.99809 (rank : 15) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y2T2 | Gene names | AP3M1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit mu-1 (Adapter-related protein complex 3 mu-1 subunit) (Mu-adaptin 3A) (AP-3 adapter complex mu3A subunit). | |||||
|
RB6I2_MOUSE
|
||||||
| NC score | 0.007454 (rank : 15) | θ value | 4.03905 (rank : 11) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 1433 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99MI1, Q80TK7, Q8BPL1, Q8C7Y1, Q99MI2 | Gene names | Erc1, Cast2, Kiaa1081, Rab6ip2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ELKS/RAB6-interacting/CAST family member 1 (RAB6-interacting protein 2) (ERC protein 1) (ERC1) (CAZ-associated structural protein 2) (CAST2). | |||||
|
RB6I2_HUMAN
|
||||||
| NC score | 0.006854 (rank : 16) | θ value | 4.03905 (rank : 10) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 1584 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8IUD2, Q6NVK2, Q8IUD3, Q8IUD4, Q8IUD5, Q8NAS1, Q9NXN5, Q9UIK7, Q9UPS1 | Gene names | ERC1, ELKS, KIAA1081, RAB6IP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ELKS/RAB6-interacting/CAST family member 1 (RAB6-interacting protein 2) (ERC protein 1). | |||||