Please be patient as the page loads
|
FAM5B_HUMAN
|
||||||
| SwissProt Accessions | Q9C0B6, O95560, Q6ZWC1, Q7LCZ9, Q8N360 | Gene names | FAM5B, BRINP2, DBCCR1L2, KIAA1747 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM5B precursor (BMP/retinoic acid-inducible neural-specific protein 2) (DBCCR1-like protein 2). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
FAM5A_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.984089 (rank : 5) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O60477, Q6IPV6, Q6P1A0, Q8WU22 | Gene names | FAM5A, DBC1, DBCCR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM5A precursor (Deleted in bladder cancer 1 protein). | |||||
|
FAM5A_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.983929 (rank : 6) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q920P3, Q80ZL2, Q812F0, Q8C1C9, Q920P4, Q9QXL0 | Gene names | Fam5a, Brinp, Brinp1, Dbccr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM5A precursor (Deleted in bladder cancer 1 protein homolog) (BMP/retinoic acid-inducible neural-specific protein 1). | |||||
|
FAM5B_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9C0B6, O95560, Q6ZWC1, Q7LCZ9, Q8N360 | Gene names | FAM5B, BRINP2, DBCCR1L2, KIAA1747 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM5B precursor (BMP/retinoic acid-inducible neural-specific protein 2) (DBCCR1-like protein 2). | |||||
|
FAM5B_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.998797 (rank : 2) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q6DFY8, Q80T96 | Gene names | Fam5b, Kiaa1747 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM5B precursor. | |||||
|
FAM5C_HUMAN
|
||||||
| θ value | 0 (rank : 5) | NC score | 0.992908 (rank : 4) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q76B58, O95726 | Gene names | FAM5C, DBCCR1L, DBCCR1L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM5C precursor (DBCCR1-like protein 1). | |||||
|
FAM5C_MOUSE
|
||||||
| θ value | 0 (rank : 6) | NC score | 0.992942 (rank : 3) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q499E0, Q8K0R9 | Gene names | Fam5c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM5C precursor. | |||||
|
CO7_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 7) | NC score | 0.106077 (rank : 7) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P10643, Q6P3T5, Q92489 | Gene names | C7 | |||
|
Domain Architecture |

|
|||||
| Description | Complement component C7 precursor. | |||||
|
PGBM_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 8) | NC score | 0.030594 (rank : 25) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 1073 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q05793 | Gene names | Hspg2 | |||
|
Domain Architecture |

|
|||||
| Description | Basement membrane-specific heparan sulfate proteoglycan core protein precursor (HSPG) (Perlecan) (PLC). | |||||
|
PGBM_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 9) | NC score | 0.028196 (rank : 28) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 1113 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P98160, Q16287, Q9H3V5 | Gene names | HSPG2 | |||
|
Domain Architecture |

|
|||||
| Description | Basement membrane-specific heparan sulfate proteoglycan core protein precursor (HSPG) (Perlecan) (PLC). | |||||
|
SREC_HUMAN
|
||||||
| θ value | 0.125558 (rank : 10) | NC score | 0.037483 (rank : 20) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q14162, O43701 | Gene names | SCARF1, KIAA0149, SREC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endothelial cells scavenger receptor precursor (Acetyl LDL receptor) (Scavenger receptor class F member 1). | |||||
|
FRAS1_HUMAN
|
||||||
| θ value | 0.21417 (rank : 11) | NC score | 0.039089 (rank : 19) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 759 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q86XX4, Q86UZ4, Q8N3U9, Q8NAU7, Q96JW7, Q9H6N9, Q9P228 | Gene names | FRAS1, KIAA1500 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Extracellular matrix protein FRAS1 precursor. | |||||
|
FBN2_HUMAN
|
||||||
| θ value | 0.279714 (rank : 12) | NC score | 0.033928 (rank : 21) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P35556 | Gene names | FBN2 | |||
|
Domain Architecture |

|
|||||
| Description | Fibrillin-2 precursor. | |||||
|
FBN2_MOUSE
|
||||||
| θ value | 0.365318 (rank : 13) | NC score | 0.033640 (rank : 22) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 549 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q61555, Q63957 | Gene names | Fbn2, Fbn-2 | |||
|
Domain Architecture |

|
|||||
| Description | Fibrillin-2 precursor. | |||||
|
CO8A_MOUSE
|
||||||
| θ value | 0.47712 (rank : 14) | NC score | 0.065733 (rank : 10) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 182 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8K182, Q8CHJ9, Q8CHK1 | Gene names | C8a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Complement component C8 alpha chain precursor (Complement component 8 subunit alpha). | |||||
|
C1QR1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 15) | NC score | 0.030382 (rank : 27) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 493 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9NPY3, O00274 | Gene names | CD93, C1QR1, MXRA4 | |||
|
Domain Architecture |
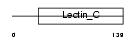
|
|||||
| Description | Complement component C1q receptor precursor (Complement component 1 q subcomponent receptor 1) (C1qR) (C1qRp) (C1qR(p)) (C1q/MBL/SPA receptor) (CD93 antigen) (CDw93). | |||||
|
ADAM8_HUMAN
|
||||||
| θ value | 1.06291 (rank : 16) | NC score | 0.013948 (rank : 49) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P78325 | Gene names | ADAM8, MS2 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 8 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 8) (Cell surface antigen MS2) (CD156a antigen) (CD156). | |||||
|
CELR2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 17) | NC score | 0.014808 (rank : 48) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9HCU4, Q92566 | Gene names | CELSR2, CDHF10, EGFL2, KIAA0279, MEGF3 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin EGF LAG seven-pass G-type receptor 2 precursor (Epidermal growth factor-like 2) (Multiple epidermal growth factor-like domains 3) (Flamingo 1). | |||||
|
PROC_HUMAN
|
||||||
| θ value | 1.38821 (rank : 18) | NC score | 0.013650 (rank : 50) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P04070, Q15189, Q15190, Q16001 | Gene names | PROC | |||
|
Domain Architecture |
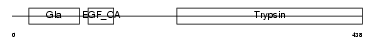
|
|||||
| Description | Vitamin K-dependent protein C precursor (EC 3.4.21.69) (Autoprothrombin IIA) (Anticoagulant protein C) (Blood coagulation factor XIV) [Contains: Vitamin K-dependent protein C light chain; Vitamin K-dependent protein C heavy chain; Activation peptide]. | |||||
|
KRA32_HUMAN
|
||||||
| θ value | 1.81305 (rank : 19) | NC score | 0.060782 (rank : 11) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BYR7 | Gene names | KRTAP3-2, KAP3.2, KRTAP3.2 | |||
|
Domain Architecture |
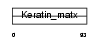
|
|||||
| Description | Keratin-associated protein 3-2 (Keratin-associated protein 3.2) (High sulfur keratin-associated protein 3.2). | |||||
|
PERF_MOUSE
|
||||||
| θ value | 1.81305 (rank : 20) | NC score | 0.082604 (rank : 8) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P10820 | Gene names | Prf1, Pfp | |||
|
Domain Architecture |

|
|||||
| Description | Perforin-1 precursor (P1) (Lymphocyte pore-forming protein) (Cytolysin). | |||||
|
CO8B_MOUSE
|
||||||
| θ value | 2.36792 (rank : 21) | NC score | 0.069279 (rank : 9) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 224 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8BH35, Q4VAH1, Q8CHJ5, Q8CHJ6, Q8VC14 | Gene names | C8b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Complement component C8 beta chain precursor (Complement component 8 subunit beta). | |||||
|
FBN1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 22) | NC score | 0.030726 (rank : 24) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 557 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P35555, Q15972, Q75N87 | Gene names | FBN1, FBN | |||
|
Domain Architecture |

|
|||||
| Description | Fibrillin-1 precursor. | |||||
|
FBN1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 23) | NC score | 0.031009 (rank : 23) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 563 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q61554, Q60826 | Gene names | Fbn1, Fbn-1 | |||
|
Domain Architecture |

|
|||||
| Description | Fibrillin-1 precursor. | |||||
|
KRA33_HUMAN
|
||||||
| θ value | 2.36792 (rank : 24) | NC score | 0.058150 (rank : 12) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BYR6, Q52LP0 | Gene names | KRTAP3-3, KAP3.3, KRTAP3.3 | |||
|
Domain Architecture |
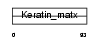
|
|||||
| Description | Keratin-associated protein 3-3 (Keratin-associated protein 3.3) (High sulfur keratin-associated protein 3.3). | |||||
|
LAMA3_MOUSE
|
||||||
| θ value | 2.36792 (rank : 25) | NC score | 0.025596 (rank : 34) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 859 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q61789, O08751, Q61788, Q61966, Q9JHQ7 | Gene names | Lama3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-3 chain precursor (Nicein subunit alpha). | |||||
|
MACOI_HUMAN
|
||||||
| θ value | 2.36792 (rank : 26) | NC score | 0.016355 (rank : 45) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8N5G2, Q2TLX5, Q2TLX6, Q9NVG6 | Gene names | TMEM57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Macoilin (Transmembrane protein 57). | |||||
|
TNR1B_HUMAN
|
||||||
| θ value | 2.36792 (rank : 27) | NC score | 0.014888 (rank : 47) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P20333, Q16042, Q6YI29, Q9UIH1 | Gene names | TNFRSF1B, TNFBR, TNFR2 | |||
|
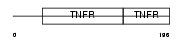
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 1B precursor (Tumor necrosis factor receptor 2) (TNF-R2) (Tumor necrosis factor receptor type II) (p75) (p80 TNF-alpha receptor) (CD120b antigen) (Etanercept) [Contains: Tumor necrosis factor receptor superfamily member 1b, membrane form; Tumor necrosis factor-binding protein 2 (TBPII) (TBP- 2)]. | |||||
|
CO8B_HUMAN
|
||||||
| θ value | 3.0926 (rank : 28) | NC score | 0.047421 (rank : 15) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P07358 | Gene names | C8B | |||
|
Domain Architecture |

|
|||||
| Description | Complement component C8 beta chain precursor (Complement component 8 subunit beta). | |||||
|
KI13A_HUMAN
|
||||||
| θ value | 3.0926 (rank : 29) | NC score | 0.008076 (rank : 54) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H1H9, Q9H1H8 | Gene names | KIF13A, RBKIN | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF13A (Kinesin-like protein RBKIN). | |||||
|
KRA31_HUMAN
|
||||||
| θ value | 3.0926 (rank : 30) | NC score | 0.054972 (rank : 14) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9BYR8 | Gene names | KRTAP3-1, KAP3.1, KRTAP3.1 | |||
|
Domain Architecture |
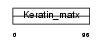
|
|||||
| Description | Keratin-associated protein 3-1 (Keratin-associated protein 3.1) (High sulfur keratin-associated protein 3.1). | |||||
|
NOTC1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 31) | NC score | 0.022989 (rank : 36) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 916 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q01705, Q06007, Q61905, Q99JC2, Q9QW58, Q9R0X7 | Gene names | Notch1, Motch | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 1 precursor (Notch 1) (Motch A) (mT14) (p300) [Contains: Notch 1 extracellular truncation; Notch 1 intracellular domain]. | |||||
|
PROC_MOUSE
|
||||||
| θ value | 4.03905 (rank : 32) | NC score | 0.012733 (rank : 51) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 426 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P33587, O35498, Q91WN8, Q99PC6 | Gene names | Proc | |||
|
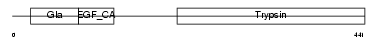
Domain Architecture |
|
|||||
| Description | Vitamin K-dependent protein C precursor (EC 3.4.21.69) (Autoprothrombin IIA) (Anticoagulant protein C) (Blood coagulation factor XIV) [Contains: Vitamin K-dependent protein C light chain; Vitamin K-dependent protein C heavy chain; Activation peptide]. | |||||
|
LAMA3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 33) | NC score | 0.026055 (rank : 32) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 760 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q16787, Q13679, Q13680, Q96TG0 | Gene names | LAMA3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-3 chain precursor (Epiligrin 170 kDa subunit) (E170) (Nicein subunit alpha). | |||||
|
LYAM2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 34) | NC score | 0.019946 (rank : 39) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P16581, P16111 | Gene names | SELE, ELAM1 | |||
|
Domain Architecture |

|
|||||
| Description | E-selectin precursor (Endothelial leukocyte adhesion molecule 1) (ELAM-1) (Leukocyte-endothelial cell adhesion molecule 2) (LECAM2) (CD62E antigen). | |||||
|
ODFP_MOUSE
|
||||||
| θ value | 5.27518 (rank : 35) | NC score | 0.019066 (rank : 43) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q61999 | Gene names | Odf1, Odfp | |||
|
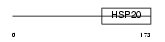
Domain Architecture |
|
|||||
| Description | Outer dense fiber protein. | |||||
|
ZAN_MOUSE
|
||||||
| θ value | 5.27518 (rank : 36) | NC score | 0.025733 (rank : 33) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 808 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O88799, O08647 | Gene names | Zan | |||
|
Domain Architecture |

|
|||||
| Description | Zonadhesin precursor. | |||||
|
CO8A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 37) | NC score | 0.045344 (rank : 17) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P07357, Q13668, Q9H130 | Gene names | C8A | |||
|
Domain Architecture |

|
|||||
| Description | Complement component C8 alpha chain precursor (Complement component 8 subunit alpha). | |||||
|
EGF_MOUSE
|
||||||
| θ value | 6.88961 (rank : 38) | NC score | 0.019660 (rank : 40) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 358 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P01132, Q6P9J2 | Gene names | Egf | |||
|
Domain Architecture |

|
|||||
| Description | Pro-epidermal growth factor precursor (EGF) [Contains: Epidermal growth factor]. | |||||
|
FRAS1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 39) | NC score | 0.030397 (rank : 26) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 747 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q80T14, Q80TC7, Q811H8, Q8BPZ4 | Gene names | Fras1, Kiaa1500 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Extracellular matrix protein FRAS1 precursor. | |||||
|
ITB5_MOUSE
|
||||||
| θ value | 6.88961 (rank : 40) | NC score | 0.018547 (rank : 44) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O70309, O70308, O88347 | Gene names | Itgb5 | |||
|
Domain Architecture |
|
|||||
| Description | Integrin beta-5 precursor. | |||||
|
LAMA5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 41) | NC score | 0.025483 (rank : 35) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 976 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O15230, Q8WZA7, Q9H1P1 | Gene names | LAMA5, KIAA0533, KIAA1907 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-5 chain precursor. | |||||
|
LAMB2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 42) | NC score | 0.022423 (rank : 37) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 692 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P55268, Q16321 | Gene names | LAMB2, LAMS | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-2 chain precursor (S-laminin) (Laminin B1s chain). | |||||
|
LYAM2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 43) | NC score | 0.019391 (rank : 42) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q00690 | Gene names | Sele, Elam-1 | |||
|
Domain Architecture |

|
|||||
| Description | E-selectin precursor (Endothelial leukocyte adhesion molecule 1) (ELAM-1) (Leukocyte-endothelial cell adhesion molecule 2) (LECAM2) (CD62E antigen). | |||||
|
MEGF6_HUMAN
|
||||||
| θ value | 6.88961 (rank : 44) | NC score | 0.026924 (rank : 30) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O75095, Q5VV39 | Gene names | MEGF6, EGFL3, KIAA0815 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Multiple epidermal growth factor-like domains 6 precursor (EGF-like domain-containing protein 3) (Multiple EGF-like domain protein 3). | |||||
|
PCSK6_HUMAN
|
||||||
| θ value | 6.88961 (rank : 45) | NC score | 0.019490 (rank : 41) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P29122, Q15099, Q15100, Q9UEG7, Q9UEJ1, Q9UEJ2, Q9UEJ7, Q9UEJ8, Q9UEJ9, Q9Y4G9, Q9Y4H0, Q9Y4H1 | Gene names | PCSK6, PACE4 | |||
|
Domain Architecture |

|
|||||
| Description | Proprotein convertase subtilisin/kexin type 6 precursor (EC 3.4.21.-) (Paired basic amino acid cleaving enzyme 4) (Subtilisin/kexin-like protease PACE4) (Subtilisin-like proprotein convertase 4) (SPC4). | |||||
|
PERF_HUMAN
|
||||||
| θ value | 6.88961 (rank : 46) | NC score | 0.055115 (rank : 13) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P14222, Q86WX7 | Gene names | PRF1, PFP | |||
|
Domain Architecture |

|
|||||
| Description | Perforin-1 precursor (P1) (Lymphocyte pore-forming protein) (PFP) (Cytolysin). | |||||
|
ZYGBL_MOUSE
|
||||||
| θ value | 6.88961 (rank : 47) | NC score | 0.010541 (rank : 53) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q80ZJ6, Q6PGH5, Q8BG38 | Gene names | Zyg11bl, Zyg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zyg-11 protein homolog (Zyg-11 homolog B-like). | |||||
|
A1AG1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 48) | NC score | 0.015799 (rank : 46) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60590 | Gene names | Orm1, Agp1, Orm-1 | |||
|
Domain Architecture |
|
|||||
| Description | Alpha-1-acid glycoprotein 1 precursor (AGP 1) (Orosomucoid-1) (OMD 1). | |||||
|
CO9_HUMAN
|
||||||
| θ value | 8.99809 (rank : 49) | NC score | 0.040070 (rank : 18) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P02748 | Gene names | C9 | |||
|
Domain Architecture |

|
|||||
| Description | Complement component C9 precursor [Contains: Complement component C9a; Complement component C9b]. | |||||
|
CO9_MOUSE
|
||||||
| θ value | 8.99809 (rank : 50) | NC score | 0.046068 (rank : 16) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P06683, Q91XA7 | Gene names | C9 | |||
|
Domain Architecture |

|
|||||
| Description | Complement component C9 precursor. | |||||
|
EXOC1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 51) | NC score | 0.007172 (rank : 55) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R3S6 | Gene names | Exoc1, Sec3, Sec3l1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Exocyst complex component 1 (Exocyst complex component Sec3). | |||||
|
LAMA5_MOUSE
|
||||||
| θ value | 8.99809 (rank : 52) | NC score | 0.026141 (rank : 31) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q61001, Q9JHQ6 | Gene names | Lama5 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-5 chain precursor. | |||||
|
MACOI_MOUSE
|
||||||
| θ value | 8.99809 (rank : 53) | NC score | 0.010579 (rank : 52) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 620 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7TQE6 | Gene names | Tmem57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Macoilin (Transmembrane protein 57) (Brain-specific adaptor protein C61). | |||||
|
NOTC3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 54) | NC score | 0.021092 (rank : 38) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 1062 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9UM47, Q9UEB3, Q9UPL3, Q9Y6L8 | Gene names | NOTCH3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 3 precursor (Notch 3) [Contains: Notch 3 extracellular truncation; Notch 3 intracellular domain]. | |||||
|
PCSK5_HUMAN
|
||||||
| θ value | 8.99809 (rank : 55) | NC score | 0.027542 (rank : 29) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q92824, Q13527 | Gene names | PCSK5, PC5, PC6 | |||
|
Domain Architecture |

|
|||||
| Description | Proprotein convertase subtilisin/kexin type 5 precursor (EC 3.4.21.-) (Proprotein convertase PC5) (Subtilisin/kexin-like protease PC5) (PC6) (hPC6). | |||||
|
FAM5B_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9C0B6, O95560, Q6ZWC1, Q7LCZ9, Q8N360 | Gene names | FAM5B, BRINP2, DBCCR1L2, KIAA1747 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM5B precursor (BMP/retinoic acid-inducible neural-specific protein 2) (DBCCR1-like protein 2). | |||||
|
FAM5B_MOUSE
|
||||||
| NC score | 0.998797 (rank : 2) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q6DFY8, Q80T96 | Gene names | Fam5b, Kiaa1747 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM5B precursor. | |||||
|
FAM5C_MOUSE
|
||||||
| NC score | 0.992942 (rank : 3) | θ value | 0 (rank : 6) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q499E0, Q8K0R9 | Gene names | Fam5c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM5C precursor. | |||||
|
FAM5C_HUMAN
|
||||||
| NC score | 0.992908 (rank : 4) | θ value | 0 (rank : 5) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q76B58, O95726 | Gene names | FAM5C, DBCCR1L, DBCCR1L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM5C precursor (DBCCR1-like protein 1). | |||||
|
FAM5A_HUMAN
|
||||||
| NC score | 0.984089 (rank : 5) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O60477, Q6IPV6, Q6P1A0, Q8WU22 | Gene names | FAM5A, DBC1, DBCCR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM5A precursor (Deleted in bladder cancer 1 protein). | |||||
|
FAM5A_MOUSE
|
||||||
| NC score | 0.983929 (rank : 6) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q920P3, Q80ZL2, Q812F0, Q8C1C9, Q920P4, Q9QXL0 | Gene names | Fam5a, Brinp, Brinp1, Dbccr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM5A precursor (Deleted in bladder cancer 1 protein homolog) (BMP/retinoic acid-inducible neural-specific protein 1). | |||||
|
CO7_HUMAN
|
||||||
| NC score | 0.106077 (rank : 7) | θ value | 0.0252991 (rank : 7) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P10643, Q6P3T5, Q92489 | Gene names | C7 | |||
|
Domain Architecture |

|
|||||
| Description | Complement component C7 precursor. | |||||
|
PERF_MOUSE
|
||||||
| NC score | 0.082604 (rank : 8) | θ value | 1.81305 (rank : 20) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P10820 | Gene names | Prf1, Pfp | |||
|
Domain Architecture |

|
|||||
| Description | Perforin-1 precursor (P1) (Lymphocyte pore-forming protein) (Cytolysin). | |||||
|
CO8B_MOUSE
|
||||||
| NC score | 0.069279 (rank : 9) | θ value | 2.36792 (rank : 21) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 224 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8BH35, Q4VAH1, Q8CHJ5, Q8CHJ6, Q8VC14 | Gene names | C8b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Complement component C8 beta chain precursor (Complement component 8 subunit beta). | |||||
|
CO8A_MOUSE
|
||||||
| NC score | 0.065733 (rank : 10) | θ value | 0.47712 (rank : 14) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 182 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8K182, Q8CHJ9, Q8CHK1 | Gene names | C8a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Complement component C8 alpha chain precursor (Complement component 8 subunit alpha). | |||||
|
KRA32_HUMAN
|
||||||
| NC score | 0.060782 (rank : 11) | θ value | 1.81305 (rank : 19) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BYR7 | Gene names | KRTAP3-2, KAP3.2, KRTAP3.2 | |||
|
Domain Architecture |
|
|||||
| Description | Keratin-associated protein 3-2 (Keratin-associated protein 3.2) (High sulfur keratin-associated protein 3.2). | |||||
|
KRA33_HUMAN
|
||||||
| NC score | 0.058150 (rank : 12) | θ value | 2.36792 (rank : 24) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BYR6, Q52LP0 | Gene names | KRTAP3-3, KAP3.3, KRTAP3.3 | |||
|
Domain Architecture |
|
|||||
| Description | Keratin-associated protein 3-3 (Keratin-associated protein 3.3) (High sulfur keratin-associated protein 3.3). | |||||
|
PERF_HUMAN
|
||||||
| NC score | 0.055115 (rank : 13) | θ value | 6.88961 (rank : 46) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P14222, Q86WX7 | Gene names | PRF1, PFP | |||
|
Domain Architecture |

|
|||||
| Description | Perforin-1 precursor (P1) (Lymphocyte pore-forming protein) (PFP) (Cytolysin). | |||||
|
KRA31_HUMAN
|
||||||
| NC score | 0.054972 (rank : 14) | θ value | 3.0926 (rank : 30) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9BYR8 | Gene names | KRTAP3-1, KAP3.1, KRTAP3.1 | |||
|
Domain Architecture |
|
|||||
| Description | Keratin-associated protein 3-1 (Keratin-associated protein 3.1) (High sulfur keratin-associated protein 3.1). | |||||
|
CO8B_HUMAN
|
||||||
| NC score | 0.047421 (rank : 15) | θ value | 3.0926 (rank : 28) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P07358 | Gene names | C8B | |||
|
Domain Architecture |

|
|||||
| Description | Complement component C8 beta chain precursor (Complement component 8 subunit beta). | |||||
|
CO9_MOUSE
|
||||||
| NC score | 0.046068 (rank : 16) | θ value | 8.99809 (rank : 50) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P06683, Q91XA7 | Gene names | C9 | |||
|
Domain Architecture |

|
|||||
| Description | Complement component C9 precursor. | |||||
|
CO8A_HUMAN
|
||||||
| NC score | 0.045344 (rank : 17) | θ value | 6.88961 (rank : 37) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P07357, Q13668, Q9H130 | Gene names | C8A | |||
|
Domain Architecture |

|
|||||
| Description | Complement component C8 alpha chain precursor (Complement component 8 subunit alpha). | |||||
|
CO9_HUMAN
|
||||||
| NC score | 0.040070 (rank : 18) | θ value | 8.99809 (rank : 49) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P02748 | Gene names | C9 | |||
|
Domain Architecture |

|
|||||
| Description | Complement component C9 precursor [Contains: Complement component C9a; Complement component C9b]. | |||||
|
FRAS1_HUMAN
|
||||||
| NC score | 0.039089 (rank : 19) | θ value | 0.21417 (rank : 11) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 759 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q86XX4, Q86UZ4, Q8N3U9, Q8NAU7, Q96JW7, Q9H6N9, Q9P228 | Gene names | FRAS1, KIAA1500 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Extracellular matrix protein FRAS1 precursor. | |||||
|
SREC_HUMAN
|
||||||
| NC score | 0.037483 (rank : 20) | θ value | 0.125558 (rank : 10) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q14162, O43701 | Gene names | SCARF1, KIAA0149, SREC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endothelial cells scavenger receptor precursor (Acetyl LDL receptor) (Scavenger receptor class F member 1). | |||||
|
FBN2_HUMAN
|
||||||
| NC score | 0.033928 (rank : 21) | θ value | 0.279714 (rank : 12) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P35556 | Gene names | FBN2 | |||
|
Domain Architecture |

|
|||||
| Description | Fibrillin-2 precursor. | |||||
|
FBN2_MOUSE
|
||||||
| NC score | 0.033640 (rank : 22) | θ value | 0.365318 (rank : 13) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 549 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q61555, Q63957 | Gene names | Fbn2, Fbn-2 | |||
|
Domain Architecture |

|
|||||
| Description | Fibrillin-2 precursor. | |||||
|
FBN1_MOUSE
|
||||||
| NC score | 0.031009 (rank : 23) | θ value | 2.36792 (rank : 23) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 563 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q61554, Q60826 | Gene names | Fbn1, Fbn-1 | |||
|
Domain Architecture |

|
|||||
| Description | Fibrillin-1 precursor. | |||||
|
FBN1_HUMAN
|
||||||
| NC score | 0.030726 (rank : 24) | θ value | 2.36792 (rank : 22) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 557 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P35555, Q15972, Q75N87 | Gene names | FBN1, FBN | |||
|
Domain Architecture |

|
|||||
| Description | Fibrillin-1 precursor. | |||||
|
PGBM_MOUSE
|
||||||
| NC score | 0.030594 (rank : 25) | θ value | 0.0431538 (rank : 8) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 1073 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q05793 | Gene names | Hspg2 | |||
|
Domain Architecture |

|
|||||
| Description | Basement membrane-specific heparan sulfate proteoglycan core protein precursor (HSPG) (Perlecan) (PLC). | |||||
|
FRAS1_MOUSE
|
||||||
| NC score | 0.030397 (rank : 26) | θ value | 6.88961 (rank : 39) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 747 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q80T14, Q80TC7, Q811H8, Q8BPZ4 | Gene names | Fras1, Kiaa1500 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Extracellular matrix protein FRAS1 precursor. | |||||
|
C1QR1_HUMAN
|
||||||
| NC score | 0.030382 (rank : 27) | θ value | 0.62314 (rank : 15) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 493 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9NPY3, O00274 | Gene names | CD93, C1QR1, MXRA4 | |||
|
Domain Architecture |

|
|||||
| Description | Complement component C1q receptor precursor (Complement component 1 q subcomponent receptor 1) (C1qR) (C1qRp) (C1qR(p)) (C1q/MBL/SPA receptor) (CD93 antigen) (CDw93). | |||||
|
PGBM_HUMAN
|
||||||
| NC score | 0.028196 (rank : 28) | θ value | 0.0961366 (rank : 9) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 1113 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P98160, Q16287, Q9H3V5 | Gene names | HSPG2 | |||
|
Domain Architecture |

|
|||||
| Description | Basement membrane-specific heparan sulfate proteoglycan core protein precursor (HSPG) (Perlecan) (PLC). | |||||
|
PCSK5_HUMAN
|
||||||
| NC score | 0.027542 (rank : 29) | θ value | 8.99809 (rank : 55) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q92824, Q13527 | Gene names | PCSK5, PC5, PC6 | |||
|
Domain Architecture |

|
|||||
| Description | Proprotein convertase subtilisin/kexin type 5 precursor (EC 3.4.21.-) (Proprotein convertase PC5) (Subtilisin/kexin-like protease PC5) (PC6) (hPC6). | |||||
|
MEGF6_HUMAN
|
||||||
| NC score | 0.026924 (rank : 30) | θ value | 6.88961 (rank : 44) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O75095, Q5VV39 | Gene names | MEGF6, EGFL3, KIAA0815 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Multiple epidermal growth factor-like domains 6 precursor (EGF-like domain-containing protein 3) (Multiple EGF-like domain protein 3). | |||||
|
LAMA5_MOUSE
|
||||||
| NC score | 0.026141 (rank : 31) | θ value | 8.99809 (rank : 52) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q61001, Q9JHQ6 | Gene names | Lama5 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-5 chain precursor. | |||||
|
LAMA3_HUMAN
|
||||||
| NC score | 0.026055 (rank : 32) | θ value | 5.27518 (rank : 33) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 760 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q16787, Q13679, Q13680, Q96TG0 | Gene names | LAMA3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-3 chain precursor (Epiligrin 170 kDa subunit) (E170) (Nicein subunit alpha). | |||||
|
ZAN_MOUSE
|
||||||
| NC score | 0.025733 (rank : 33) | θ value | 5.27518 (rank : 36) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 808 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O88799, O08647 | Gene names | Zan | |||
|
Domain Architecture |

|
|||||
| Description | Zonadhesin precursor. | |||||
|
LAMA3_MOUSE
|
||||||
| NC score | 0.025596 (rank : 34) | θ value | 2.36792 (rank : 25) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 859 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q61789, O08751, Q61788, Q61966, Q9JHQ7 | Gene names | Lama3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-3 chain precursor (Nicein subunit alpha). | |||||
|
LAMA5_HUMAN
|
||||||
| NC score | 0.025483 (rank : 35) | θ value | 6.88961 (rank : 41) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 976 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O15230, Q8WZA7, Q9H1P1 | Gene names | LAMA5, KIAA0533, KIAA1907 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-5 chain precursor. | |||||
|
NOTC1_MOUSE
|
||||||
| NC score | 0.022989 (rank : 36) | θ value | 3.0926 (rank : 31) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 916 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q01705, Q06007, Q61905, Q99JC2, Q9QW58, Q9R0X7 | Gene names | Notch1, Motch | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 1 precursor (Notch 1) (Motch A) (mT14) (p300) [Contains: Notch 1 extracellular truncation; Notch 1 intracellular domain]. | |||||
|
LAMB2_HUMAN
|
||||||
| NC score | 0.022423 (rank : 37) | θ value | 6.88961 (rank : 42) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 692 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P55268, Q16321 | Gene names | LAMB2, LAMS | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-2 chain precursor (S-laminin) (Laminin B1s chain). | |||||
|
NOTC3_HUMAN
|
||||||
| NC score | 0.021092 (rank : 38) | θ value | 8.99809 (rank : 54) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 1062 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9UM47, Q9UEB3, Q9UPL3, Q9Y6L8 | Gene names | NOTCH3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 3 precursor (Notch 3) [Contains: Notch 3 extracellular truncation; Notch 3 intracellular domain]. | |||||
|
LYAM2_HUMAN
|
||||||
| NC score | 0.019946 (rank : 39) | θ value | 5.27518 (rank : 34) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P16581, P16111 | Gene names | SELE, ELAM1 | |||
|
Domain Architecture |

|
|||||
| Description | E-selectin precursor (Endothelial leukocyte adhesion molecule 1) (ELAM-1) (Leukocyte-endothelial cell adhesion molecule 2) (LECAM2) (CD62E antigen). | |||||
|
EGF_MOUSE
|
||||||
| NC score | 0.019660 (rank : 40) | θ value | 6.88961 (rank : 38) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 358 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P01132, Q6P9J2 | Gene names | Egf | |||
|
Domain Architecture |

|
|||||
| Description | Pro-epidermal growth factor precursor (EGF) [Contains: Epidermal growth factor]. | |||||
|
PCSK6_HUMAN
|
||||||
| NC score | 0.019490 (rank : 41) | θ value | 6.88961 (rank : 45) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P29122, Q15099, Q15100, Q9UEG7, Q9UEJ1, Q9UEJ2, Q9UEJ7, Q9UEJ8, Q9UEJ9, Q9Y4G9, Q9Y4H0, Q9Y4H1 | Gene names | PCSK6, PACE4 | |||
|
Domain Architecture |

|
|||||
| Description | Proprotein convertase subtilisin/kexin type 6 precursor (EC 3.4.21.-) (Paired basic amino acid cleaving enzyme 4) (Subtilisin/kexin-like protease PACE4) (Subtilisin-like proprotein convertase 4) (SPC4). | |||||
|
LYAM2_MOUSE
|
||||||
| NC score | 0.019391 (rank : 42) | θ value | 6.88961 (rank : 43) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q00690 | Gene names | Sele, Elam-1 | |||
|
Domain Architecture |

|
|||||
| Description | E-selectin precursor (Endothelial leukocyte adhesion molecule 1) (ELAM-1) (Leukocyte-endothelial cell adhesion molecule 2) (LECAM2) (CD62E antigen). | |||||
|
ODFP_MOUSE
|
||||||
| NC score | 0.019066 (rank : 43) | θ value | 5.27518 (rank : 35) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q61999 | Gene names | Odf1, Odfp | |||
|
Domain Architecture |

|
|||||
| Description | Outer dense fiber protein. | |||||
|
ITB5_MOUSE
|
||||||
| NC score | 0.018547 (rank : 44) | θ value | 6.88961 (rank : 40) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O70309, O70308, O88347 | Gene names | Itgb5 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-5 precursor. | |||||
|
MACOI_HUMAN
|
||||||
| NC score | 0.016355 (rank : 45) | θ value | 2.36792 (rank : 26) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8N5G2, Q2TLX5, Q2TLX6, Q9NVG6 | Gene names | TMEM57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Macoilin (Transmembrane protein 57). | |||||
|
A1AG1_MOUSE
|
||||||
| NC score | 0.015799 (rank : 46) | θ value | 8.99809 (rank : 48) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60590 | Gene names | Orm1, Agp1, Orm-1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-1-acid glycoprotein 1 precursor (AGP 1) (Orosomucoid-1) (OMD 1). | |||||
|
TNR1B_HUMAN
|
||||||
| NC score | 0.014888 (rank : 47) | θ value | 2.36792 (rank : 27) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P20333, Q16042, Q6YI29, Q9UIH1 | Gene names | TNFRSF1B, TNFBR, TNFR2 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 1B precursor (Tumor necrosis factor receptor 2) (TNF-R2) (Tumor necrosis factor receptor type II) (p75) (p80 TNF-alpha receptor) (CD120b antigen) (Etanercept) [Contains: Tumor necrosis factor receptor superfamily member 1b, membrane form; Tumor necrosis factor-binding protein 2 (TBPII) (TBP- 2)]. | |||||
|
CELR2_HUMAN
|
||||||
| NC score | 0.014808 (rank : 48) | θ value | 1.38821 (rank : 17) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9HCU4, Q92566 | Gene names | CELSR2, CDHF10, EGFL2, KIAA0279, MEGF3 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin EGF LAG seven-pass G-type receptor 2 precursor (Epidermal growth factor-like 2) (Multiple epidermal growth factor-like domains 3) (Flamingo 1). | |||||
|
ADAM8_HUMAN
|
||||||
| NC score | 0.013948 (rank : 49) | θ value | 1.06291 (rank : 16) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P78325 | Gene names | ADAM8, MS2 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 8 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 8) (Cell surface antigen MS2) (CD156a antigen) (CD156). | |||||
|
PROC_HUMAN
|
||||||
| NC score | 0.013650 (rank : 50) | θ value | 1.38821 (rank : 18) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P04070, Q15189, Q15190, Q16001 | Gene names | PROC | |||
|
Domain Architecture |

|
|||||
| Description | Vitamin K-dependent protein C precursor (EC 3.4.21.69) (Autoprothrombin IIA) (Anticoagulant protein C) (Blood coagulation factor XIV) [Contains: Vitamin K-dependent protein C light chain; Vitamin K-dependent protein C heavy chain; Activation peptide]. | |||||
|
PROC_MOUSE
|
||||||
| NC score | 0.012733 (rank : 51) | θ value | 4.03905 (rank : 32) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 426 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P33587, O35498, Q91WN8, Q99PC6 | Gene names | Proc | |||
|
Domain Architecture |

|
|||||
| Description | Vitamin K-dependent protein C precursor (EC 3.4.21.69) (Autoprothrombin IIA) (Anticoagulant protein C) (Blood coagulation factor XIV) [Contains: Vitamin K-dependent protein C light chain; Vitamin K-dependent protein C heavy chain; Activation peptide]. | |||||
|
MACOI_MOUSE
|
||||||
| NC score | 0.010579 (rank : 52) | θ value | 8.99809 (rank : 53) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 620 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7TQE6 | Gene names | Tmem57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Macoilin (Transmembrane protein 57) (Brain-specific adaptor protein C61). | |||||
|
ZYGBL_MOUSE
|
||||||
| NC score | 0.010541 (rank : 53) | θ value | 6.88961 (rank : 47) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q80ZJ6, Q6PGH5, Q8BG38 | Gene names | Zyg11bl, Zyg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zyg-11 protein homolog (Zyg-11 homolog B-like). | |||||
|
KI13A_HUMAN
|
||||||
| NC score | 0.008076 (rank : 54) | θ value | 3.0926 (rank : 29) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H1H9, Q9H1H8 | Gene names | KIF13A, RBKIN | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF13A (Kinesin-like protein RBKIN). | |||||
|
EXOC1_MOUSE
|
||||||
| NC score | 0.007172 (rank : 55) | θ value | 8.99809 (rank : 51) | |||
| Query Neighborhood Hits | 55 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R3S6 | Gene names | Exoc1, Sec3, Sec3l1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Exocyst complex component 1 (Exocyst complex component Sec3). | |||||