Please be patient as the page loads
|
CASB_HUMAN
|
||||||
| SwissProt Accessions | P05814, Q4VAZ9 | Gene names | CSN2, CASB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-casein precursor. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
CASB_HUMAN
|
||||||
| θ value | 4.83544e-70 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P05814, Q4VAZ9 | Gene names | CSN2, CASB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-casein precursor. | |||||
|
CASB_MOUSE
|
||||||
| θ value | 2.27234e-27 (rank : 2) | NC score | 0.856733 (rank : 2) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P10598, Q543D9, Q8VCT6, Q8VCU8, Q91VI5, Q922Y5, Q9D1U6, Q9D1U7 | Gene names | Csn2, Csnb | |||
|
Domain Architecture |

|
|||||
| Description | Beta-casein precursor. | |||||
|
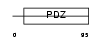
PRAX_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 3) | NC score | 0.151617 (rank : 4) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O55103, O55104 | Gene names | Prx | |||
|
Domain Architecture |
|
|||||
| Description | Periaxin. | |||||
|
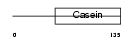
CASE_MOUSE
|
||||||
| θ value | 0.125558 (rank : 4) | NC score | 0.184800 (rank : 3) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P02664, Q542L5 | Gene names | Csne, Csnd | |||
|
Domain Architecture |
|
|||||
| Description | Epsilon-casein precursor. | |||||
|
MINT_MOUSE
|
||||||
| θ value | 0.163984 (rank : 5) | NC score | 0.078899 (rank : 6) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
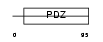
PRAX_HUMAN
|
||||||
| θ value | 0.163984 (rank : 6) | NC score | 0.139239 (rank : 5) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BXM0, Q9BXL9, Q9HCF2 | Gene names | PRX, KIAA1620 | |||
|
Domain Architecture |
|
|||||
| Description | Periaxin. | |||||
|
STAT2_MOUSE
|
||||||
| θ value | 0.365318 (rank : 7) | NC score | 0.053405 (rank : 9) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9WVL2, Q64188, Q64189, Q64250 | Gene names | Stat2 | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 2. | |||||
|
SON_HUMAN
|
||||||
| θ value | 0.813845 (rank : 8) | NC score | 0.051159 (rank : 10) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
TARSH_HUMAN
|
||||||
| θ value | 0.813845 (rank : 9) | NC score | 0.066316 (rank : 8) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 768 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q7Z7G0, Q6ZW20, Q6ZW22, Q9C082, Q9UFI6 | Gene names | ABI3BP, NESHBP, TARSH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Target of Nesh-SH3 precursor (Tarsh) (Nesh-binding protein) (NeshBP) (ABI gene family member 3-binding protein). | |||||
|
DCC_MOUSE
|
||||||
| θ value | 1.81305 (rank : 10) | NC score | 0.019015 (rank : 18) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P70211 | Gene names | Dcc | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC). | |||||
|
MINT_HUMAN
|
||||||
| θ value | 1.81305 (rank : 11) | NC score | 0.073948 (rank : 7) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 1291 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q96T58, Q9H9A8, Q9NWH5, Q9UQ01, Q9Y556 | Gene names | SPEN, KIAA0929, MINT, SHARP | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
PTN23_HUMAN
|
||||||
| θ value | 2.36792 (rank : 12) | NC score | 0.024717 (rank : 17) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 646 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9H3S7, Q7KZF8, Q8N6Z5, Q9BSR5, Q9P257, Q9UG03, Q9UMZ4 | Gene names | PTPN23, KIAA1471 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 23 (EC 3.1.3.48) (His- domain-containing protein tyrosine phosphatase) (HD-PTP) (Protein tyrosine phosphatase TD14) (PTP-TD14). | |||||
|
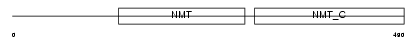
NMT1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 13) | NC score | 0.048334 (rank : 11) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O70310 | Gene names | Nmt1 | |||
|
Domain Architecture |
|
|||||
| Description | Glycylpeptide N-tetradecanoyltransferase 1 (EC 2.3.1.97) (Peptide N- myristoyltransferase 1) (Myristoyl-CoA:protein N-myristoyltransferase 1) (NMT 1) (Type I N-myristoyltransferase). | |||||
|
AFF2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 14) | NC score | 0.027534 (rank : 15) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P51816, O43786, O60215, P78407, Q13521, Q14323, Q9UNA5 | Gene names | AFF2, FMR2, OX19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation 2 protein) (Protein FMR-2) (FMR2P) (Protein Ox19) (Fragile X E mental retardation syndrome protein). | |||||
|
NMT1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 15) | NC score | 0.045291 (rank : 12) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P30419, Q9UE09 | Gene names | NMT1, NMT | |||
|
Domain Architecture |

|
|||||
| Description | Glycylpeptide N-tetradecanoyltransferase 1 (EC 2.3.1.97) (Peptide N- myristoyltransferase 1) (Myristoyl-CoA:protein N-myristoyltransferase 1) (NMT 1) (Type I N-myristoyltransferase). | |||||
|
TCRG1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 16) | NC score | 0.040733 (rank : 14) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 929 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O14776 | Gene names | TCERG1, CA150, TAF2S | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150). | |||||
|
TCRG1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 17) | NC score | 0.040754 (rank : 13) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8CGF7, Q61051, Q8C490, Q8CHT8, Q9R0R5 | Gene names | Tcerg1, Taf2s | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150) (p144) (Formin- binding protein 28) (FBP 28). | |||||
|
CD008_HUMAN
|
||||||
| θ value | 8.99809 (rank : 18) | NC score | 0.025334 (rank : 16) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P78312, O43607, P78311, P78313, Q9UEG8 | Gene names | C4orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C4orf8 (Protein IT14). | |||||
|
FBN1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 19) | NC score | 0.005024 (rank : 19) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 557 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P35555, Q15972, Q75N87 | Gene names | FBN1, FBN | |||
|
Domain Architecture |

|
|||||
| Description | Fibrillin-1 precursor. | |||||
|
CASB_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 4.83544e-70 (rank : 1) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P05814, Q4VAZ9 | Gene names | CSN2, CASB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-casein precursor. | |||||
|
CASB_MOUSE
|
||||||
| NC score | 0.856733 (rank : 2) | θ value | 2.27234e-27 (rank : 2) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P10598, Q543D9, Q8VCT6, Q8VCU8, Q91VI5, Q922Y5, Q9D1U6, Q9D1U7 | Gene names | Csn2, Csnb | |||
|
Domain Architecture |

|
|||||
| Description | Beta-casein precursor. | |||||
|
CASE_MOUSE
|
||||||
| NC score | 0.184800 (rank : 3) | θ value | 0.125558 (rank : 4) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P02664, Q542L5 | Gene names | Csne, Csnd | |||
|
Domain Architecture |

|
|||||
| Description | Epsilon-casein precursor. | |||||
|
PRAX_MOUSE
|
||||||
| NC score | 0.151617 (rank : 4) | θ value | 0.0431538 (rank : 3) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O55103, O55104 | Gene names | Prx | |||
|
Domain Architecture |

|
|||||
| Description | Periaxin. | |||||
|
PRAX_HUMAN
|
||||||
| NC score | 0.139239 (rank : 5) | θ value | 0.163984 (rank : 6) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BXM0, Q9BXL9, Q9HCF2 | Gene names | PRX, KIAA1620 | |||
|
Domain Architecture |

|
|||||
| Description | Periaxin. | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.078899 (rank : 6) | θ value | 0.163984 (rank : 5) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
MINT_HUMAN
|
||||||
| NC score | 0.073948 (rank : 7) | θ value | 1.81305 (rank : 11) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 1291 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q96T58, Q9H9A8, Q9NWH5, Q9UQ01, Q9Y556 | Gene names | SPEN, KIAA0929, MINT, SHARP | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
TARSH_HUMAN
|
||||||
| NC score | 0.066316 (rank : 8) | θ value | 0.813845 (rank : 9) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 768 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q7Z7G0, Q6ZW20, Q6ZW22, Q9C082, Q9UFI6 | Gene names | ABI3BP, NESHBP, TARSH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Target of Nesh-SH3 precursor (Tarsh) (Nesh-binding protein) (NeshBP) (ABI gene family member 3-binding protein). | |||||
|
STAT2_MOUSE
|
||||||
| NC score | 0.053405 (rank : 9) | θ value | 0.365318 (rank : 7) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9WVL2, Q64188, Q64189, Q64250 | Gene names | Stat2 | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 2. | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.051159 (rank : 10) | θ value | 0.813845 (rank : 8) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
NMT1_MOUSE
|
||||||
| NC score | 0.048334 (rank : 11) | θ value | 3.0926 (rank : 13) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O70310 | Gene names | Nmt1 | |||
|
Domain Architecture |

|
|||||
| Description | Glycylpeptide N-tetradecanoyltransferase 1 (EC 2.3.1.97) (Peptide N- myristoyltransferase 1) (Myristoyl-CoA:protein N-myristoyltransferase 1) (NMT 1) (Type I N-myristoyltransferase). | |||||
|
NMT1_HUMAN
|
||||||
| NC score | 0.045291 (rank : 12) | θ value | 5.27518 (rank : 15) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P30419, Q9UE09 | Gene names | NMT1, NMT | |||
|
Domain Architecture |

|
|||||
| Description | Glycylpeptide N-tetradecanoyltransferase 1 (EC 2.3.1.97) (Peptide N- myristoyltransferase 1) (Myristoyl-CoA:protein N-myristoyltransferase 1) (NMT 1) (Type I N-myristoyltransferase). | |||||
|
TCRG1_MOUSE
|
||||||
| NC score | 0.040754 (rank : 13) | θ value | 5.27518 (rank : 17) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8CGF7, Q61051, Q8C490, Q8CHT8, Q9R0R5 | Gene names | Tcerg1, Taf2s | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150) (p144) (Formin- binding protein 28) (FBP 28). | |||||
|
TCRG1_HUMAN
|
||||||
| NC score | 0.040733 (rank : 14) | θ value | 5.27518 (rank : 16) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 929 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O14776 | Gene names | TCERG1, CA150, TAF2S | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150). | |||||
|
AFF2_HUMAN
|
||||||
| NC score | 0.027534 (rank : 15) | θ value | 5.27518 (rank : 14) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P51816, O43786, O60215, P78407, Q13521, Q14323, Q9UNA5 | Gene names | AFF2, FMR2, OX19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation 2 protein) (Protein FMR-2) (FMR2P) (Protein Ox19) (Fragile X E mental retardation syndrome protein). | |||||
|
CD008_HUMAN
|
||||||
| NC score | 0.025334 (rank : 16) | θ value | 8.99809 (rank : 18) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P78312, O43607, P78311, P78313, Q9UEG8 | Gene names | C4orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C4orf8 (Protein IT14). | |||||
|
PTN23_HUMAN
|
||||||
| NC score | 0.024717 (rank : 17) | θ value | 2.36792 (rank : 12) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 646 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9H3S7, Q7KZF8, Q8N6Z5, Q9BSR5, Q9P257, Q9UG03, Q9UMZ4 | Gene names | PTPN23, KIAA1471 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 23 (EC 3.1.3.48) (His- domain-containing protein tyrosine phosphatase) (HD-PTP) (Protein tyrosine phosphatase TD14) (PTP-TD14). | |||||
|
DCC_MOUSE
|
||||||
| NC score | 0.019015 (rank : 18) | θ value | 1.81305 (rank : 10) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P70211 | Gene names | Dcc | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC). | |||||
|
FBN1_HUMAN
|
||||||
| NC score | 0.005024 (rank : 19) | θ value | 8.99809 (rank : 19) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 557 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P35555, Q15972, Q75N87 | Gene names | FBN1, FBN | |||
|
Domain Architecture |

|
|||||
| Description | Fibrillin-1 precursor. | |||||