Please be patient as the page loads
|
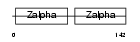
ZBP1_MOUSE
|
||||||
| SwissProt Accessions | Q9QY24, Q8VE02, Q9D8B9 | Gene names | Zbp1, Dlm1 | |||
|
Domain Architecture |
|
|||||
| Description | Z-DNA-binding protein 1 (Tumor stroma and activated macrophage protein DLM-1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
ZBP1_MOUSE
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9QY24, Q8VE02, Q9D8B9 | Gene names | Zbp1, Dlm1 | |||
|
Domain Architecture |

|
|||||
| Description | Z-DNA-binding protein 1 (Tumor stroma and activated macrophage protein DLM-1). | |||||
|
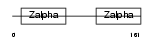
ZBP1_HUMAN
|
||||||
| θ value | 1.00358e-83 (rank : 2) | NC score | 0.918991 (rank : 2) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H171, Q5JY39, Q9BYW4 | Gene names | ZBP1, C20orf183, DLM1 | |||
|
Domain Architecture |
|
|||||
| Description | Z-DNA-binding protein 1 (Tumor stroma and activated macrophage protein DLM-1). | |||||
|
DSRAD_MOUSE
|
||||||
| θ value | 2.21117e-06 (rank : 3) | NC score | 0.202321 (rank : 3) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99MU3, O70375, Q80UZ6, Q8C222, Q99MU2, Q99MU4, Q99MU7 | Gene names | Adar | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific adenosine deaminase (EC 3.5.4.-) (DRADA) (RNA adenosine deaminase 1). | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 4) | NC score | 0.049243 (rank : 12) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
MUCDL_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 5) | NC score | 0.054744 (rank : 8) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9HBB8, Q9H746, Q9HAU3, Q9HBB5, Q9HBB6, Q9HBB7, Q9NX86, Q9NXI9 | Gene names | MUCDHL | |||
|
Domain Architecture |

|
|||||
| Description | Mucin and cadherin-like protein precursor (Mu-protocadherin). | |||||
|
DSRAD_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 6) | NC score | 0.144528 (rank : 4) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P55265, O15223, O43859, O43860, Q9BYM3, Q9BYM4 | Gene names | ADAR, ADAR1, DSRAD, IFI4 | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific adenosine deaminase (EC 3.5.4.-) (DRADA) (136 kDa double-stranded RNA-binding protein) (P136) (K88DSRBP) (Interferon-inducible protein 4) (IFI-4 protein). | |||||
|
MUC1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 7) | NC score | 0.054755 (rank : 7) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q02496 | Gene names | Muc1, Muc-1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (Polymorphic epithelial mucin) (PEMT) (Episialin). | |||||
|
OSTP_MOUSE
|
||||||
| θ value | 0.47712 (rank : 8) | NC score | 0.070331 (rank : 5) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P10923, P19008 | Gene names | Spp1, Eta-1, Op, Spp-1 | |||
|
Domain Architecture |

|
|||||
| Description | Osteopontin precursor (Bone sialoprotein-1) (Secreted phosphoprotein 1) (SPP-1) (Minopontin) (Early T-lymphocyte activation 1 protein) (2AR) (Calcium oxalate crystal growth inhibitor protein). | |||||
|
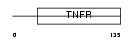
TNR19_MOUSE
|
||||||
| θ value | 0.62314 (rank : 9) | NC score | 0.046664 (rank : 13) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JLL3, Q812G3, Q9JHF1, Q9JJH6, Q9JLL2, Q9QXW7 | Gene names | Tnfrsf19, Taj, Troy | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 19 precursor (Toxicity and JNK inducer) (TRADE). | |||||
|
ABL2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 10) | NC score | 0.006331 (rank : 28) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1045 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P42684, Q5T0X6, Q6NZY6 | Gene names | ABL2, ABLL, ARG | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein kinase ABL2 (EC 2.7.10.2) (Abelson murine leukemia viral oncogene homolog 2) (Tyrosine kinase ARG). | |||||
|
TNKS1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 11) | NC score | 0.016818 (rank : 21) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 477 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O95271, O95272 | Gene names | TNKS, PARP5A, PARPL, TIN1, TINF1, TNKS1 | |||
|
Domain Architecture |

|
|||||
| Description | Tankyrase-1 (EC 2.4.2.30) (TANK1) (Tankyrase I) (TNKS-1) (TRF1- interacting ankyrin-related ADP-ribose polymerase). | |||||
|
OR5C1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 12) | NC score | 0.003165 (rank : 30) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 660 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NGR4, Q96RC4 | Gene names | OR5C1 | |||
|
Domain Architecture |

|
|||||
| Description | Olfactory receptor 5C1 (Olfactory receptor 9-F) (OR9-F). | |||||
|
TPR_HUMAN
|
||||||
| θ value | 2.36792 (rank : 13) | NC score | 0.010992 (rank : 26) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
MDC1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 14) | NC score | 0.027970 (rank : 15) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
NBR1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 15) | NC score | 0.037601 (rank : 14) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q14596, Q13173, Q15026, Q96GB6, Q9NRF7 | Gene names | NBR1, 1A13B, KIAA0049, M17S2 | |||
|
Domain Architecture |

|
|||||
| Description | Next to BRCA1 gene 1 protein (Neighbor of BRCA1 gene 1 protein) (Membrane component, chromosome 17, surface marker 2) (1A1-3B). | |||||
|
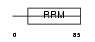
RALY_HUMAN
|
||||||
| θ value | 3.0926 (rank : 16) | NC score | 0.025557 (rank : 18) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UKM9, Q14621, Q5QPL8, Q9BQX6, Q9UJE3 | Gene names | RALY, P542 | |||
|
Domain Architecture |
|
|||||
| Description | RNA-binding protein Raly (hnRNP associated with lethal yellow homolog) (Autoantigen p542). | |||||
|
SYNE1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 17) | NC score | 0.013372 (rank : 24) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1484 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8NF91, O94890, Q5JV19, Q5JV22, Q8N9P7, Q8TCP1, Q8WWW6, Q8WWW7, Q8WXF6, Q96N17, Q9C0A7, Q9H525, Q9H526, Q9NS36, Q9NU50, Q9UJ06, Q9UJ07, Q9ULF8 | Gene names | SYNE1, KIAA0796, KIAA1262, KIAA1756, MYNE1 | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-1 (Nuclear envelope spectrin repeat protein 1) (Synaptic nuclear envelope protein 1) (Syne-1) (Myocyte nuclear envelope protein 1) (Myne-1) (Enaptin). | |||||
|
RPGF2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 18) | NC score | 0.018972 (rank : 20) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y4G8 | Gene names | RAPGEF2, KIAA0313, NRAPGEP, PDZGEF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rap guanine nucleotide exchange factor 2 (Neural RAP guanine nucleotide exchange protein) (nRap GEP) (PDZ domain-containing guanine nucleotide exchange factor 1) (PDZ-GEF1) (RA-GEF). | |||||
|
STK10_HUMAN
|
||||||
| θ value | 5.27518 (rank : 19) | NC score | 0.003823 (rank : 29) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O94804, Q9UIW4 | Gene names | STK10, LOK | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase 10 (EC 2.7.11.1) (Lymphocyte-oriented kinase). | |||||
|
OXR1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 20) | NC score | 0.020621 (rank : 19) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q4KMM3, Q5FWW1, Q99L06, Q99MK1, Q99MP4 | Gene names | Oxr1, C7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oxidation resistance protein 1 (Protein C7). | |||||
|
PDZK4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 21) | NC score | 0.015248 (rank : 22) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q76G19, Q8NB75, Q9BUH9, Q9P284 | Gene names | PDZK4, KIAA1444, PDZRN4L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 4 (PDZ domain-containing RING finger protein 4-like protein). | |||||
|
UBP8_HUMAN
|
||||||
| θ value | 6.88961 (rank : 22) | NC score | 0.010408 (rank : 27) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 751 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P40818, Q7Z3U2, Q86VA0, Q8IWI7 | Gene names | USP8, KIAA0055, UBPY | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 8 (EC 3.1.2.15) (Ubiquitin thioesterase 8) (Ubiquitin-specific-processing protease 8) (Deubiquitinating enzyme 8) (hUBPy). | |||||
|
ANK2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 23) | NC score | 0.012692 (rank : 25) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
MINT_HUMAN
|
||||||
| θ value | 8.99809 (rank : 24) | NC score | 0.013569 (rank : 23) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1291 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96T58, Q9H9A8, Q9NWH5, Q9UQ01, Q9Y556 | Gene names | SPEN, KIAA0929, MINT, SHARP | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
MLF2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 25) | NC score | 0.025725 (rank : 16) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q15773 | Gene names | MLF2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myeloid leukemia factor 2 (Myelodysplasia-myeloid leukemia factor 2). | |||||
|
MLF2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 26) | NC score | 0.025567 (rank : 17) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99KX1 | Gene names | Mlf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myeloid leukemia factor 2 (Myelodysplasia-myeloid leukemia factor 2). | |||||
|
PGBM_MOUSE
|
||||||
| θ value | 8.99809 (rank : 27) | NC score | 0.002827 (rank : 31) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1073 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q05793 | Gene names | Hspg2 | |||
|
Domain Architecture |

|
|||||
| Description | Basement membrane-specific heparan sulfate proteoglycan core protein precursor (HSPG) (Perlecan) (PLC). | |||||
|
RED1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 28) | NC score | 0.053740 (rank : 11) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P78563, O00395, O00465, O00691, O00692, P78555, Q8NFD1 | Gene names | ADARB1, ADAR2, DRADA2, RED1 | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific editase 1 (EC 3.5.-.-) (dsRNA adenosine deaminase) (RNA-editing deaminase 1) (RNA-editing enzyme 1). | |||||
|
RED1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 29) | NC score | 0.054357 (rank : 10) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91ZS8, Q8K3X1, Q91ZS6, Q91ZS7, Q91ZS9, Q99MU8 | Gene names | Adarb1, Adar2, Red1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double-stranded RNA-specific editase 1 (EC 3.5.-.-) (dsRNA adenosine deaminase) (RNA-editing deaminase 1) (RNA-editing enzyme 1). | |||||
|
RED2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 30) | NC score | 0.055323 (rank : 6) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NS39 | Gene names | ADARB2, ADAR3, RED2 | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific editase B2 (EC 3.5.-.-) (dsRNA adenosine deaminase B2) (RNA-dependent adenosine deaminase 3) (RNA-editing deaminase 2) (RNA-editing enzyme 2). | |||||
|
RED2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 31) | NC score | 0.054446 (rank : 9) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JI20, Q3UTP1, Q8CC51, Q9JIE5 | Gene names | Adarb2, Adar3, Red2 | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific editase B2 (EC 3.5.-.-) (dsRNA adenosine deaminase B2) (RNA-dependent adenosine deaminase 3) (RNA-editing deaminase 2) (RNA-editing enzyme 2). | |||||
|
ZBP1_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9QY24, Q8VE02, Q9D8B9 | Gene names | Zbp1, Dlm1 | |||
|
Domain Architecture |

|
|||||
| Description | Z-DNA-binding protein 1 (Tumor stroma and activated macrophage protein DLM-1). | |||||
|
ZBP1_HUMAN
|
||||||
| NC score | 0.918991 (rank : 2) | θ value | 1.00358e-83 (rank : 2) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H171, Q5JY39, Q9BYW4 | Gene names | ZBP1, C20orf183, DLM1 | |||
|
Domain Architecture |

|
|||||
| Description | Z-DNA-binding protein 1 (Tumor stroma and activated macrophage protein DLM-1). | |||||
|
DSRAD_MOUSE
|
||||||
| NC score | 0.202321 (rank : 3) | θ value | 2.21117e-06 (rank : 3) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99MU3, O70375, Q80UZ6, Q8C222, Q99MU2, Q99MU4, Q99MU7 | Gene names | Adar | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific adenosine deaminase (EC 3.5.4.-) (DRADA) (RNA adenosine deaminase 1). | |||||
|
DSRAD_HUMAN
|
||||||
| NC score | 0.144528 (rank : 4) | θ value | 0.0961366 (rank : 6) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P55265, O15223, O43859, O43860, Q9BYM3, Q9BYM4 | Gene names | ADAR, ADAR1, DSRAD, IFI4 | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific adenosine deaminase (EC 3.5.4.-) (DRADA) (136 kDa double-stranded RNA-binding protein) (P136) (K88DSRBP) (Interferon-inducible protein 4) (IFI-4 protein). | |||||
|
OSTP_MOUSE
|
||||||
| NC score | 0.070331 (rank : 5) | θ value | 0.47712 (rank : 8) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P10923, P19008 | Gene names | Spp1, Eta-1, Op, Spp-1 | |||
|
Domain Architecture |

|
|||||
| Description | Osteopontin precursor (Bone sialoprotein-1) (Secreted phosphoprotein 1) (SPP-1) (Minopontin) (Early T-lymphocyte activation 1 protein) (2AR) (Calcium oxalate crystal growth inhibitor protein). | |||||
|
RED2_HUMAN
|
||||||
| NC score | 0.055323 (rank : 6) | θ value | θ > 10 (rank : 30) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NS39 | Gene names | ADARB2, ADAR3, RED2 | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific editase B2 (EC 3.5.-.-) (dsRNA adenosine deaminase B2) (RNA-dependent adenosine deaminase 3) (RNA-editing deaminase 2) (RNA-editing enzyme 2). | |||||
|
MUC1_MOUSE
|
||||||
| NC score | 0.054755 (rank : 7) | θ value | 0.47712 (rank : 7) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q02496 | Gene names | Muc1, Muc-1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (Polymorphic epithelial mucin) (PEMT) (Episialin). | |||||
|
MUCDL_HUMAN
|
||||||
| NC score | 0.054744 (rank : 8) | θ value | 0.0431538 (rank : 5) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9HBB8, Q9H746, Q9HAU3, Q9HBB5, Q9HBB6, Q9HBB7, Q9NX86, Q9NXI9 | Gene names | MUCDHL | |||
|
Domain Architecture |

|
|||||
| Description | Mucin and cadherin-like protein precursor (Mu-protocadherin). | |||||
|
RED2_MOUSE
|
||||||
| NC score | 0.054446 (rank : 9) | θ value | θ > 10 (rank : 31) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JI20, Q3UTP1, Q8CC51, Q9JIE5 | Gene names | Adarb2, Adar3, Red2 | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific editase B2 (EC 3.5.-.-) (dsRNA adenosine deaminase B2) (RNA-dependent adenosine deaminase 3) (RNA-editing deaminase 2) (RNA-editing enzyme 2). | |||||
|
RED1_MOUSE
|
||||||
| NC score | 0.054357 (rank : 10) | θ value | θ > 10 (rank : 29) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91ZS8, Q8K3X1, Q91ZS6, Q91ZS7, Q91ZS9, Q99MU8 | Gene names | Adarb1, Adar2, Red1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double-stranded RNA-specific editase 1 (EC 3.5.-.-) (dsRNA adenosine deaminase) (RNA-editing deaminase 1) (RNA-editing enzyme 1). | |||||
|
RED1_HUMAN
|
||||||
| NC score | 0.053740 (rank : 11) | θ value | θ > 10 (rank : 28) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P78563, O00395, O00465, O00691, O00692, P78555, Q8NFD1 | Gene names | ADARB1, ADAR2, DRADA2, RED1 | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific editase 1 (EC 3.5.-.-) (dsRNA adenosine deaminase) (RNA-editing deaminase 1) (RNA-editing enzyme 1). | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.049243 (rank : 12) | θ value | 0.0193708 (rank : 4) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
TNR19_MOUSE
|
||||||
| NC score | 0.046664 (rank : 13) | θ value | 0.62314 (rank : 9) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JLL3, Q812G3, Q9JHF1, Q9JJH6, Q9JLL2, Q9QXW7 | Gene names | Tnfrsf19, Taj, Troy | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 19 precursor (Toxicity and JNK inducer) (TRADE). | |||||
|
NBR1_HUMAN
|
||||||
| NC score | 0.037601 (rank : 14) | θ value | 3.0926 (rank : 15) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q14596, Q13173, Q15026, Q96GB6, Q9NRF7 | Gene names | NBR1, 1A13B, KIAA0049, M17S2 | |||
|
Domain Architecture |

|
|||||
| Description | Next to BRCA1 gene 1 protein (Neighbor of BRCA1 gene 1 protein) (Membrane component, chromosome 17, surface marker 2) (1A1-3B). | |||||
|
MDC1_HUMAN
|
||||||
| NC score | 0.027970 (rank : 15) | θ value | 3.0926 (rank : 14) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
MLF2_HUMAN
|
||||||
| NC score | 0.025725 (rank : 16) | θ value | 8.99809 (rank : 25) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q15773 | Gene names | MLF2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myeloid leukemia factor 2 (Myelodysplasia-myeloid leukemia factor 2). | |||||
|
MLF2_MOUSE
|
||||||
| NC score | 0.025567 (rank : 17) | θ value | 8.99809 (rank : 26) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99KX1 | Gene names | Mlf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myeloid leukemia factor 2 (Myelodysplasia-myeloid leukemia factor 2). | |||||
|
RALY_HUMAN
|
||||||
| NC score | 0.025557 (rank : 18) | θ value | 3.0926 (rank : 16) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UKM9, Q14621, Q5QPL8, Q9BQX6, Q9UJE3 | Gene names | RALY, P542 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein Raly (hnRNP associated with lethal yellow homolog) (Autoantigen p542). | |||||
|
OXR1_MOUSE
|
||||||
| NC score | 0.020621 (rank : 19) | θ value | 6.88961 (rank : 20) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q4KMM3, Q5FWW1, Q99L06, Q99MK1, Q99MP4 | Gene names | Oxr1, C7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oxidation resistance protein 1 (Protein C7). | |||||
|
RPGF2_HUMAN
|
||||||
| NC score | 0.018972 (rank : 20) | θ value | 4.03905 (rank : 18) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y4G8 | Gene names | RAPGEF2, KIAA0313, NRAPGEP, PDZGEF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rap guanine nucleotide exchange factor 2 (Neural RAP guanine nucleotide exchange protein) (nRap GEP) (PDZ domain-containing guanine nucleotide exchange factor 1) (PDZ-GEF1) (RA-GEF). | |||||
|
TNKS1_HUMAN
|
||||||
| NC score | 0.016818 (rank : 21) | θ value | 1.06291 (rank : 11) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 477 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O95271, O95272 | Gene names | TNKS, PARP5A, PARPL, TIN1, TINF1, TNKS1 | |||
|
Domain Architecture |

|
|||||
| Description | Tankyrase-1 (EC 2.4.2.30) (TANK1) (Tankyrase I) (TNKS-1) (TRF1- interacting ankyrin-related ADP-ribose polymerase). | |||||
|
PDZK4_HUMAN
|
||||||
| NC score | 0.015248 (rank : 22) | θ value | 6.88961 (rank : 21) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q76G19, Q8NB75, Q9BUH9, Q9P284 | Gene names | PDZK4, KIAA1444, PDZRN4L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 4 (PDZ domain-containing RING finger protein 4-like protein). | |||||
|
MINT_HUMAN
|
||||||
| NC score | 0.013569 (rank : 23) | θ value | 8.99809 (rank : 24) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1291 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96T58, Q9H9A8, Q9NWH5, Q9UQ01, Q9Y556 | Gene names | SPEN, KIAA0929, MINT, SHARP | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
SYNE1_HUMAN
|
||||||
| NC score | 0.013372 (rank : 24) | θ value | 3.0926 (rank : 17) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1484 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8NF91, O94890, Q5JV19, Q5JV22, Q8N9P7, Q8TCP1, Q8WWW6, Q8WWW7, Q8WXF6, Q96N17, Q9C0A7, Q9H525, Q9H526, Q9NS36, Q9NU50, Q9UJ06, Q9UJ07, Q9ULF8 | Gene names | SYNE1, KIAA0796, KIAA1262, KIAA1756, MYNE1 | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-1 (Nuclear envelope spectrin repeat protein 1) (Synaptic nuclear envelope protein 1) (Syne-1) (Myocyte nuclear envelope protein 1) (Myne-1) (Enaptin). | |||||
|
ANK2_HUMAN
|
||||||
| NC score | 0.012692 (rank : 25) | θ value | 8.99809 (rank : 23) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
TPR_HUMAN
|
||||||
| NC score | 0.010992 (rank : 26) | θ value | 2.36792 (rank : 13) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
UBP8_HUMAN
|
||||||
| NC score | 0.010408 (rank : 27) | θ value | 6.88961 (rank : 22) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 751 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P40818, Q7Z3U2, Q86VA0, Q8IWI7 | Gene names | USP8, KIAA0055, UBPY | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 8 (EC 3.1.2.15) (Ubiquitin thioesterase 8) (Ubiquitin-specific-processing protease 8) (Deubiquitinating enzyme 8) (hUBPy). | |||||
|
ABL2_HUMAN
|
||||||
| NC score | 0.006331 (rank : 28) | θ value | 1.06291 (rank : 10) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1045 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P42684, Q5T0X6, Q6NZY6 | Gene names | ABL2, ABLL, ARG | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein kinase ABL2 (EC 2.7.10.2) (Abelson murine leukemia viral oncogene homolog 2) (Tyrosine kinase ARG). | |||||
|
STK10_HUMAN
|
||||||
| NC score | 0.003823 (rank : 29) | θ value | 5.27518 (rank : 19) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O94804, Q9UIW4 | Gene names | STK10, LOK | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase 10 (EC 2.7.11.1) (Lymphocyte-oriented kinase). | |||||
|
OR5C1_HUMAN
|
||||||
| NC score | 0.003165 (rank : 30) | θ value | 1.38821 (rank : 12) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 660 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NGR4, Q96RC4 | Gene names | OR5C1 | |||
|
Domain Architecture |

|
|||||
| Description | Olfactory receptor 5C1 (Olfactory receptor 9-F) (OR9-F). | |||||
|
PGBM_MOUSE
|
||||||
| NC score | 0.002827 (rank : 31) | θ value | 8.99809 (rank : 27) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 1073 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q05793 | Gene names | Hspg2 | |||
|
Domain Architecture |

|
|||||
| Description | Basement membrane-specific heparan sulfate proteoglycan core protein precursor (HSPG) (Perlecan) (PLC). | |||||