Please be patient as the page loads
|
RM40_HUMAN
|
||||||
| SwissProt Accessions | Q9NQ50, O95134 | Gene names | MRPL40, NLVCF, URIM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 39S ribosomal protein L40, mitochondrial precursor (L40mt) (MRP-40) (Nuclear localization signal-containing protein deleted in velocardiofacial syndrome) (Up-regulated in metastasis). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
RM40_HUMAN
|
||||||
| θ value | 5.14231e-88 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9NQ50, O95134 | Gene names | MRPL40, NLVCF, URIM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 39S ribosomal protein L40, mitochondrial precursor (L40mt) (MRP-40) (Nuclear localization signal-containing protein deleted in velocardiofacial syndrome) (Up-regulated in metastasis). | |||||
|
RM40_MOUSE
|
||||||
| θ value | 4.22625e-74 (rank : 2) | NC score | 0.963516 (rank : 2) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Z2Q5, Q9CS52 | Gene names | Mrpl40, Nlvcf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 39S ribosomal protein L40, mitochondrial precursor (L40mt) (MRP-40) (Nuclear localization signal-containing protein deleted in velocardiofacial syndrome homolog). | |||||
|
SYCP1_MOUSE
|
||||||
| θ value | 0.279714 (rank : 3) | NC score | 0.047349 (rank : 7) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1456 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q62209, O09205, P70192, Q62329 | Gene names | Sycp1, Scp1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptonemal complex protein 1 (SCP-1). | |||||
|
SMC1A_HUMAN
|
||||||
| θ value | 0.365318 (rank : 4) | NC score | 0.052881 (rank : 4) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1346 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q14683, O14995, Q16351 | Gene names | SMC1L1, DXS423E, KIAA0178, SB1.8, SMC1 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Sb1.8). | |||||
|
SMC1A_MOUSE
|
||||||
| θ value | 0.365318 (rank : 5) | NC score | 0.052079 (rank : 5) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1384 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9CU62, Q3V480, Q9CUX9, Q9D959, Q9WTU1 | Gene names | Smc1l1, Sb1.8, Smc1, Smcb | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Chromosome segregation protein SmcB) (Sb1.8). | |||||
|
MRCKB_MOUSE
|
||||||
| θ value | 0.47712 (rank : 6) | NC score | 0.018504 (rank : 27) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 2180 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q7TT50, Q69ZR5, Q80W33 | Gene names | Cdc42bpb, Kiaa1124 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK beta (EC 2.7.11.1) (CDC42-binding protein kinase beta) (Myotonic dystrophy kinase-related CDC42-binding kinase beta) (Myotonic dystrophy protein kinase-like beta) (MRCK beta) (DMPK-like beta). | |||||
|
CF060_HUMAN
|
||||||
| θ value | 1.38821 (rank : 7) | NC score | 0.035860 (rank : 12) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1270 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8NB25, Q96GY8, Q9H851 | Gene names | C6orf60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C6orf60. | |||||
|
CP250_HUMAN
|
||||||
| θ value | 1.38821 (rank : 8) | NC score | 0.035893 (rank : 11) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1805 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9BV73, O14812, O60588, Q9H450 | Gene names | CEP250, CEP2, CNAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1). | |||||
|
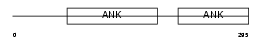
PP16B_HUMAN
|
||||||
| θ value | 1.81305 (rank : 9) | NC score | 0.024617 (rank : 23) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96T49, O94912, Q9NQG4 | Gene names | PPP1R16B, ANKRD4, KIAA0823 | |||
|
Domain Architecture |
|
|||||
| Description | Protein phosphatase 1 regulatory inhibitor subunit 16B (TGF-beta- inhibited membrane-associated protein) (hTIMAP) (CAAX box protein TIMAP) (Ankyrin repeat domain protein 4). | |||||
|
RPB1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 10) | NC score | 0.047355 (rank : 6) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P24928 | Gene names | POLR2A | |||
|
Domain Architecture |

|
|||||
| Description | DNA-directed RNA polymerase II largest subunit (EC 2.7.7.6) (RPB1). | |||||
|
RPB1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 11) | NC score | 0.047308 (rank : 8) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 363 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P08775 | Gene names | Polr2a, Rpii215, Rpo2-1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-directed RNA polymerase II largest subunit (EC 2.7.7.6) (RPB1). | |||||
|
CN135_HUMAN
|
||||||
| θ value | 2.36792 (rank : 12) | NC score | 0.054633 (rank : 3) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q63HM2, Q9BQG8, Q9H9F2 | Gene names | C14orf135, FBP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf135 precursor (Hepatitis C virus F protein-binding protein 2) (HCV F protein-binding protein 2). | |||||
|
CP250_MOUSE
|
||||||
| θ value | 2.36792 (rank : 13) | NC score | 0.036958 (rank : 10) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1583 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q60952, Q2I8G3, Q3UTR4, Q6PFF6, Q8BLC6 | Gene names | Cep250, Cep2, Inmp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1) (Intranuclear matrix protein). | |||||
|
MINT_MOUSE
|
||||||
| θ value | 2.36792 (rank : 14) | NC score | 0.026709 (rank : 21) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
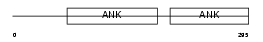
PP16B_MOUSE
|
||||||
| θ value | 2.36792 (rank : 15) | NC score | 0.024104 (rank : 24) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VHQ3 | Gene names | Ppp1r16b | |||
|
Domain Architecture |
|
|||||
| Description | Protein phosphatase 1 regulatory inhibitor subunit 16B (TGF-beta- inhibited membrane-associated protein) (CAAX box protein TIMAP). | |||||
|
SYCP1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 16) | NC score | 0.043093 (rank : 9) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1313 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q15431, O14963 | Gene names | SYCP1, SCP1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptonemal complex protein 1 (SCP-1). | |||||
|
SON_HUMAN
|
||||||
| θ value | 4.03905 (rank : 17) | NC score | 0.023435 (rank : 25) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
SON_MOUSE
|
||||||
| θ value | 4.03905 (rank : 18) | NC score | 0.032954 (rank : 14) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 957 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9QX47, Q9CQ12, Q9CQK6, Q9QXP5 | Gene names | Son | |||
|
Domain Architecture |

|
|||||
| Description | SON protein. | |||||
|
GBP2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 19) | NC score | 0.028353 (rank : 19) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P32456, Q86TB0 | Gene names | GBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon-induced guanylate-binding protein 2 (GTP-binding protein 2) (Guanine nucleotide-binding protein 2) (GBP-2) (HuGBP-2). | |||||
|
LIPA1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 20) | NC score | 0.027368 (rank : 20) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1294 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q13136, Q13135, Q14567, Q8N4I2 | Gene names | PPFIA1, LIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-1 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-1) (PTPRF-interacting protein alpha-1) (LAR-interacting protein 1) (LIP.1). | |||||
|
CK5P2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 21) | NC score | 0.031592 (rank : 16) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1151 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8K389, Q6PCN1, Q6ZPL0 | Gene names | Cdk5rap2, Kiaa1633 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CDK5 regulatory subunit-associated protein 2 (CDK5 activator-binding protein C48). | |||||
|
GOGA4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 22) | NC score | 0.029004 (rank : 18) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1939 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q91VW5, O70365, Q8CGH6 | Gene names | Golga4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (tGolgin-1). | |||||
|
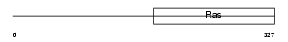
IF2M_MOUSE
|
||||||
| θ value | 6.88961 (rank : 23) | NC score | 0.022534 (rank : 26) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q91YJ5 | Gene names | Mtif2 | |||
|
Domain Architecture |
|
|||||
| Description | Translation initiation factor IF-2, mitochondrial precursor (IF-2Mt) (IF-2(Mt)). | |||||
|
YLPM1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 24) | NC score | 0.030801 (rank : 17) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P49750, P49752, Q96I64, Q9P1V7 | Gene names | YLPM1, C14orf170, ZAP3 | |||
|
Domain Architecture |

|
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3) (ZAP113). | |||||
|
YLPM1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 25) | NC score | 0.032627 (rank : 15) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
NUF2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 26) | NC score | 0.034879 (rank : 13) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 661 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9BZD4, Q8WU69, Q96HJ4, Q96Q78 | Gene names | CDCA1, NUF2, NUF2R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinetochore protein Nuf2 (hNuf2) (hNuf2R) (hsNuf2) (Cell division cycle-associated protein 1). | |||||
|
SYNE2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 27) | NC score | 0.025969 (rank : 22) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1323 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8WXH0, Q8N1S3, Q8NF49, Q8TER7, Q8WWW3, Q8WWW4, Q8WWW5, Q8WXH1, Q9NU50, Q9UFQ4, Q9Y2L4, Q9Y4R1 | Gene names | SYNE2, KIAA1011, NUA | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-2 (Nuclear envelope spectrin repeat protein 2) (Syne-2) (Synaptic nuclear envelope protein 2) (Nucleus and actin connecting element protein) (Protein NUANCE). | |||||
|
US6NL_MOUSE
|
||||||
| θ value | 8.99809 (rank : 28) | NC score | 0.016119 (rank : 28) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80XC3, Q6ZQK8 | Gene names | Usp6nl, Kiaa0019 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USP6 N-terminal-like protein. | |||||
|
RM40_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 5.14231e-88 (rank : 1) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9NQ50, O95134 | Gene names | MRPL40, NLVCF, URIM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 39S ribosomal protein L40, mitochondrial precursor (L40mt) (MRP-40) (Nuclear localization signal-containing protein deleted in velocardiofacial syndrome) (Up-regulated in metastasis). | |||||
|
RM40_MOUSE
|
||||||
| NC score | 0.963516 (rank : 2) | θ value | 4.22625e-74 (rank : 2) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Z2Q5, Q9CS52 | Gene names | Mrpl40, Nlvcf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 39S ribosomal protein L40, mitochondrial precursor (L40mt) (MRP-40) (Nuclear localization signal-containing protein deleted in velocardiofacial syndrome homolog). | |||||
|
CN135_HUMAN
|
||||||
| NC score | 0.054633 (rank : 3) | θ value | 2.36792 (rank : 12) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q63HM2, Q9BQG8, Q9H9F2 | Gene names | C14orf135, FBP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf135 precursor (Hepatitis C virus F protein-binding protein 2) (HCV F protein-binding protein 2). | |||||
|
SMC1A_HUMAN
|
||||||
| NC score | 0.052881 (rank : 4) | θ value | 0.365318 (rank : 4) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1346 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q14683, O14995, Q16351 | Gene names | SMC1L1, DXS423E, KIAA0178, SB1.8, SMC1 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Sb1.8). | |||||
|
SMC1A_MOUSE
|
||||||
| NC score | 0.052079 (rank : 5) | θ value | 0.365318 (rank : 5) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1384 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9CU62, Q3V480, Q9CUX9, Q9D959, Q9WTU1 | Gene names | Smc1l1, Sb1.8, Smc1, Smcb | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Chromosome segregation protein SmcB) (Sb1.8). | |||||
|
RPB1_HUMAN
|
||||||
| NC score | 0.047355 (rank : 6) | θ value | 1.81305 (rank : 10) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P24928 | Gene names | POLR2A | |||
|
Domain Architecture |

|
|||||
| Description | DNA-directed RNA polymerase II largest subunit (EC 2.7.7.6) (RPB1). | |||||
|
SYCP1_MOUSE
|
||||||
| NC score | 0.047349 (rank : 7) | θ value | 0.279714 (rank : 3) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1456 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q62209, O09205, P70192, Q62329 | Gene names | Sycp1, Scp1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptonemal complex protein 1 (SCP-1). | |||||
|
RPB1_MOUSE
|
||||||
| NC score | 0.047308 (rank : 8) | θ value | 1.81305 (rank : 11) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 363 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P08775 | Gene names | Polr2a, Rpii215, Rpo2-1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-directed RNA polymerase II largest subunit (EC 2.7.7.6) (RPB1). | |||||
|
SYCP1_HUMAN
|
||||||
| NC score | 0.043093 (rank : 9) | θ value | 2.36792 (rank : 16) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1313 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q15431, O14963 | Gene names | SYCP1, SCP1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptonemal complex protein 1 (SCP-1). | |||||
|
CP250_MOUSE
|
||||||
| NC score | 0.036958 (rank : 10) | θ value | 2.36792 (rank : 13) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1583 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q60952, Q2I8G3, Q3UTR4, Q6PFF6, Q8BLC6 | Gene names | Cep250, Cep2, Inmp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1) (Intranuclear matrix protein). | |||||
|
CP250_HUMAN
|
||||||
| NC score | 0.035893 (rank : 11) | θ value | 1.38821 (rank : 8) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1805 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9BV73, O14812, O60588, Q9H450 | Gene names | CEP250, CEP2, CNAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1). | |||||
|
CF060_HUMAN
|
||||||
| NC score | 0.035860 (rank : 12) | θ value | 1.38821 (rank : 7) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1270 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8NB25, Q96GY8, Q9H851 | Gene names | C6orf60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C6orf60. | |||||
|
NUF2_HUMAN
|
||||||
| NC score | 0.034879 (rank : 13) | θ value | 8.99809 (rank : 26) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 661 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9BZD4, Q8WU69, Q96HJ4, Q96Q78 | Gene names | CDCA1, NUF2, NUF2R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinetochore protein Nuf2 (hNuf2) (hNuf2R) (hsNuf2) (Cell division cycle-associated protein 1). | |||||
|
SON_MOUSE
|
||||||
| NC score | 0.032954 (rank : 14) | θ value | 4.03905 (rank : 18) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 957 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9QX47, Q9CQ12, Q9CQK6, Q9QXP5 | Gene names | Son | |||
|
Domain Architecture |

|
|||||
| Description | SON protein. | |||||
|
YLPM1_MOUSE
|
||||||
| NC score | 0.032627 (rank : 15) | θ value | 6.88961 (rank : 25) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
CK5P2_MOUSE
|
||||||
| NC score | 0.031592 (rank : 16) | θ value | 6.88961 (rank : 21) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1151 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8K389, Q6PCN1, Q6ZPL0 | Gene names | Cdk5rap2, Kiaa1633 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CDK5 regulatory subunit-associated protein 2 (CDK5 activator-binding protein C48). | |||||
|
YLPM1_HUMAN
|
||||||
| NC score | 0.030801 (rank : 17) | θ value | 6.88961 (rank : 24) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P49750, P49752, Q96I64, Q9P1V7 | Gene names | YLPM1, C14orf170, ZAP3 | |||
|
Domain Architecture |

|
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3) (ZAP113). | |||||
|
GOGA4_MOUSE
|
||||||
| NC score | 0.029004 (rank : 18) | θ value | 6.88961 (rank : 22) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1939 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q91VW5, O70365, Q8CGH6 | Gene names | Golga4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (tGolgin-1). | |||||
|
GBP2_HUMAN
|
||||||
| NC score | 0.028353 (rank : 19) | θ value | 5.27518 (rank : 19) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P32456, Q86TB0 | Gene names | GBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon-induced guanylate-binding protein 2 (GTP-binding protein 2) (Guanine nucleotide-binding protein 2) (GBP-2) (HuGBP-2). | |||||
|
LIPA1_HUMAN
|
||||||
| NC score | 0.027368 (rank : 20) | θ value | 5.27518 (rank : 20) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1294 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q13136, Q13135, Q14567, Q8N4I2 | Gene names | PPFIA1, LIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-1 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-1) (PTPRF-interacting protein alpha-1) (LAR-interacting protein 1) (LIP.1). | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.026709 (rank : 21) | θ value | 2.36792 (rank : 14) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
SYNE2_HUMAN
|
||||||
| NC score | 0.025969 (rank : 22) | θ value | 8.99809 (rank : 27) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1323 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8WXH0, Q8N1S3, Q8NF49, Q8TER7, Q8WWW3, Q8WWW4, Q8WWW5, Q8WXH1, Q9NU50, Q9UFQ4, Q9Y2L4, Q9Y4R1 | Gene names | SYNE2, KIAA1011, NUA | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-2 (Nuclear envelope spectrin repeat protein 2) (Syne-2) (Synaptic nuclear envelope protein 2) (Nucleus and actin connecting element protein) (Protein NUANCE). | |||||
|
PP16B_HUMAN
|
||||||
| NC score | 0.024617 (rank : 23) | θ value | 1.81305 (rank : 9) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96T49, O94912, Q9NQG4 | Gene names | PPP1R16B, ANKRD4, KIAA0823 | |||
|
Domain Architecture |

|
|||||
| Description | Protein phosphatase 1 regulatory inhibitor subunit 16B (TGF-beta- inhibited membrane-associated protein) (hTIMAP) (CAAX box protein TIMAP) (Ankyrin repeat domain protein 4). | |||||
|
PP16B_MOUSE
|
||||||
| NC score | 0.024104 (rank : 24) | θ value | 2.36792 (rank : 15) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VHQ3 | Gene names | Ppp1r16b | |||
|
Domain Architecture |

|
|||||
| Description | Protein phosphatase 1 regulatory inhibitor subunit 16B (TGF-beta- inhibited membrane-associated protein) (CAAX box protein TIMAP). | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.023435 (rank : 25) | θ value | 4.03905 (rank : 17) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
IF2M_MOUSE
|
||||||
| NC score | 0.022534 (rank : 26) | θ value | 6.88961 (rank : 23) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q91YJ5 | Gene names | Mtif2 | |||
|
Domain Architecture |

|
|||||
| Description | Translation initiation factor IF-2, mitochondrial precursor (IF-2Mt) (IF-2(Mt)). | |||||
|
MRCKB_MOUSE
|
||||||
| NC score | 0.018504 (rank : 27) | θ value | 0.47712 (rank : 6) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 2180 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q7TT50, Q69ZR5, Q80W33 | Gene names | Cdc42bpb, Kiaa1124 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK beta (EC 2.7.11.1) (CDC42-binding protein kinase beta) (Myotonic dystrophy kinase-related CDC42-binding kinase beta) (Myotonic dystrophy protein kinase-like beta) (MRCK beta) (DMPK-like beta). | |||||
|
US6NL_MOUSE
|
||||||
| NC score | 0.016119 (rank : 28) | θ value | 8.99809 (rank : 28) | |||
| Query Neighborhood Hits | 28 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80XC3, Q6ZQK8 | Gene names | Usp6nl, Kiaa0019 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USP6 N-terminal-like protein. | |||||